Abstract
In patients with advanced-stage cancer, cancer-associated anorexia affects treatment success and patient survival. However, the underlying mechanism is poorly understood. Here, we show that Dilp8, a Drosophila homologue of mammalian insulin-like 3 peptide (INSL3), is secreted from tumour tissues and induces anorexia through the Lgr3 receptor in the brain. Activated Dilp8-Lgr3 signalling upregulated anorexigenic nucleobinding 1 (NUCB1) and downregulated orexigenic short neuropeptide F (sNPF) and NPF expression in the brain. In the cancer condition, the protein expression of Lgr3 and NUCB1 was significantly upregulated in neurons expressing sNPF and NPF. INSL3 levels were increased in tumour-implanted mice and INSL3-treated mouse hypothalamic cells showed Nucb2 upregulation and Npy downregulation. Food consumption was significantly reduced in intracerebrospinal INSL3-injected mice. In patients with pancreatic cancer, higher serum INSL3 levels increased anorexia. These results indicate that tumour-derived Dilp8/INSL3 induces cancer anorexia by regulating feeding hormones through the Lgr3/Lgr8 receptor in Drosophila and mammals.
Similar content being viewed by others
Main
Anorexia, which is common in patients with advanced-stage cancer, leads to progressive weight loss and contributes to cancer-related mortality. It is closely linked to cancer-associated cachexia1,2,3,4,5, which severely affects the quality of life of patients with cancer6,7. In a case study of 66 cancer patients near the end of life, more than 60% of patients showed anorexia despite not receiving chemotherapy8,9. This suggests that the development of anorexia in patients with cancer is independent from the treatment. However, the molecular mechanism underlying cancer anorexia is largely unknown.
In the cancer anorexia-cachexia syndrome, secreted tumour-derived humoral cytokines play an essential role in inter-organ/tissue communication10,11,12,13. Systemic coordination and integration among organs and tissues by inter-organ communication is important in normal conditions and in pathologies such as cancer and metabolic syndrome14. Numerous evidence suggest that tumours elicit inflammatory responses15,16 and metabolic dysfunction in muscle and fat through the activation of pro-inflammatory cytokines17,18,19,20. Systemic inflammatory cytokines such as tumour necrosis factor-α, interleukin-1α (IL-1α), IL-1β, IL-6, transforming growth factor-β and parathyroid-hormone-related protein (PTHrP) function as tumour-derived hormones that mediate muscle wasting in mouse cancer models and human cancer patients21,22,23. Recently, growth differentiation factor (GDF15) has also been implicated in cancer cachexia due to the correlation of GDF15 with food intake and body weight loss in numerous chronic human diseases24,25,26. Recent studies have shown that ImpL2, a Drosophila homologue of mammalian insulin-like growth factor binding protein (IGFBP), is secreted from tumour tissue and impairs muscle function and tissue wasting by inhibiting IGF-like signalling in Drosophila27,28. These Drosophila cancer models reveal how tumour-secreted cytokines cause cachexia-like tissue wasting, demonstrating that Drosophila is a good model system to elucidate the mechanisms of the tumour–host interaction.
In this Article, we establish a Drosophila cancer anorexia model by overexpressing the known oncogene yorkie and demonstrate that tumour-derived Dilp8 systemically regulates the expression of feeding neuropeptides through its receptor Lgr3 in the brain. We also show that this cancer anorexia pathway is conserved in mammals. Tumour-derived insulin-like 3 peptide (INSL3), a mammalian homologue of Dilp8, regulates anorexigenic Nucb2 and orexigenic Npy through its receptor Lgr8 in mouse hypothalamic cells and pancreatic-tumour-cell-implanted mice. In addition, we show that the level of INSL3 is substantially increased in the serum of patients with pancreatic cancer.
Results
Cancer anorexia precedes organ wasting in the Drosophila eye tumour model
To investigate the molecular mechanism of cancer-induced anorexia in Drosophila, we established a Drosophila cancer model by overexpressing yorkie (yki), a transcriptional coactivator of the Hippo pathway29,30,31, in adult eyes using the GMR-Gal4 driver. Mutations in the Hippo pathway lead to malignant tumour formation in Drosophila and mammals, and are often detected in human cancers32,33,34. Overexpression of wild-type yki in the eyes (GMR>ykiWT) caused a mild rough eye phenotype, whereas the constitutively active form GMR>ykiS168A (hereafter referred to as GMR>ykiSA) resulted in a dramatic protrusion of the eyes caused by massive cell proliferation (Extended Data Fig. 1a and Supplementary Fig. 1a). We examined whether the survival rate was lower in the fly tumour model than in normal flies, similar to the high mortality of patients with cancer. The results showed that GMR>ykiSA flies had a significantly shorter life span (Fig. 1a). Next, cancer anorexia in GMR>ykiSA flies was examined by measuring the relative food intake using blue dye35 and capillary feeder (CAFE) assays36,37. Food intake was ~30% lower in 10-day-old GMR>ykiSA adult flies than in the controls of both assays (Fig. 1b and Extended Data Fig. 1b,c), suggesting that the fly cancer model mimics human cancer anorexia.
a, The life span of the constitutively active form of yki flies (GMR>ykiSA) was shorter than that of control flies (GMR-Gal4). b, The 10-day-old yki overexpressing flies (GMR>ykiWT and GMR>ykiSA) showed decreased food intake compared with that of the control in the blue dye feeding assay. n = 5 biologically independent experiments; GMR>ykiWT *P = 0.013, GMR>ykiSA **P = 0.006. c, The ovaries of GMR>ykiSA flies showed a severe wasting phenotype after 15 days. Scale bar, 200 μm. d, The triglyceride levels of GMR>ykiSA flies showed reduced lipid levels after 15 days. n = 3 biologically independent experiments; day 15 *P = 0.044, day 20 *P = 0.019. e, Reduced food intake was observed in five-day-old GMR>ykiSA flies. n = 3 biologically independent experiments; day 5 *P = 0.041, day 10 *P = 0.015, day 15 *P = 0.016, day 20 *P = 0.039. Data are presented as mean ± s.e.m. Statistical significance was determined with a two-tailed Student’s t-test; *P < 0.05, **P < 0.01.
ImpL2 induces cachexia-like wasting in intestinal malignant tumours and in a tumour allograft model27,28. We avoided tumour formation in tissues involved in the brain–gut axis to minimize the effects on feeding behaviour. In our fly eye cancer model (GMR>ykiSA), a mild wasting phenotype in the ovary and reduced triglyceride levels in the fat tissues were observed up to 10 days, whereas a severe wasting phenotype developed after 15–20 days (Fig. 1c,d and Extended Data Fig. 1d,e). The thoracic muscle of 15-day-old GMR>ykiSA adult flies also showed an abnormal mitochondrial structure with irregular packing of mitochondria between the muscle fibres, which indicates the muscle wasting phenotype (Extended Data Fig. 1f and Supplementary Fig. 1b). GMR>ykiSA flies also showed reduced food intake after five days (Fig. 1e). These results indicated that cancer anorexia seems to precede cachexia-like organ wasting in GMR>ykiSA flies.
Tumour-derived Dilp8 suppresses food intake through its receptor Lgr3
Recent studies show that Dilp8, ImpL2 and Upd2 are tumour-secreted factors27,28. RNA-sequencing (RNA-seq) analysis confirmed that the mRNA levels of these three genes were highly increased in the heads of GMR>ykiSA flies (Extended Data Fig. 2a). Dilp8 and Upd2 were highly expressed in five-day-old flies, whereas ImpL2 mRNA expression was peaked in the 15-day-old fly cancer model (Fig. 2a–c). Because cancer anorexia precedes organ wasting, Dilp8 or Upd2 may be involved in cancer anorexia rather than organ wasting. Next, relative food intake was measured after inhibiting Dilp8, ImpL2 and Upd2 in the GMR>ykiSA cancer condition. Dilp8 or ImpL2 inhibition (Dilp8-RNAi or ImpL2-RNAi) significantly rescued the reduced food intake in the GMR>ykiSA cancer model compared with the effect of inhibition of Upd2 (Fig. 2d). To further test our findings in the GMR>ykiSA tumour model, we generated a second fly tumour model in which Rasv12 was overexpressed in the eye using the sevenless-Gal4 driver. We confirmed that sev>Rasv12 flies also display anorexia and that this phenotype was effectively rescued by Dilp8-RNAi (Extended Data Fig. 2b). These findings indicate that Dilp8 is probably a key secreted factor for inducing cancer anorexia. Consistently, Dilp8 mRNA level was increased ~2.5-fold in tumour tissue (eye disc) but not in non-tumour tissue (brain) (Fig. 2e). To determine whether Dilp8 is secreted from GMR>ykiSA tumour tissues, we generated an antibody against Dilp8 and confirmed the specificity in the Dilp8M100727 hypomorphic mutant38,39 (Extended Data Fig. 2c). Western blot analysis detected Dilp8 in the adult haemolymph of GMR>ykiSA flies (Fig. 2f), suggesting that tumour-derived Dilp8 is released into the circulating haemolymph to relay the signal to the target tissues. The elevated Dilp8 protein level in GMR>ykiSA fly heads was rescued in GMR>ykiSA, Dilp8-RNAi fly heads (Extended Data Fig. 2d,e).
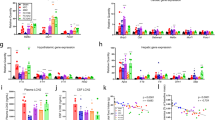
a–c, Dilp8 mRNA expression levels peaked in 5-day-old GMR>ykiSA flies and gradually decreased (a), while Upd2 (b) and ImpL2 (c) mRNA expression levels peaked in 5-day-old and 15-day-old GMR>ykiSA flies, respectively. n = 4 biologically independent experiments: day 5 **P = 0.003, day 10 *P = 0.025, day 15 **P = 0.001, day 20 **P = 0.002 (a); day 5 **P = 0.002, day 10 ***P = 0.0004, day 15 ***P = 0.0002, day 20 **P = 0.003 (b); day 5 **P = 0.0076, day 10 ***P = 0.00006, day 15 ***P = 0.00058, day 20 ***P = 0.0009 (c). d, Among the tumour-derived cytokines Dilp8, ImpL2 and Upd2, Dilp8-RNAi significantly rescued the reduced food intake by GMR>ykiSA in the feeding assay. n = 3 biologically independent experiments; GMR>ykiSA *P = 0.043, GMR>ykiSADilp8-RNAi *P = 0.018, GMR>ykiSAImpL2-RNAi *P = 0.041, GMR>ykiSAUpd2-RNAi NS = 0.792. e, Dilp8 mRNA level was increased ~2.5-fold only in tumour tissues (eye disc). n = 3 biologically independent experiments; brain NS = 0.103, eye disc *P = 0.047. f, Circulating Dilp8 was detected in the adult haemolymph of ykiSA overexpressing flies. An asterisk indicates a non-specific band. n = 3 biologically independent experiments. g,h, The expression levels of Dilp8 (g) and its receptor Lgr3 (h) gradually decreased in response to starvation and rapidly increased after refeeding. n = 3 biologically independent experiments: 12 h *P = 0.056, 24 h *P = 0.001, refed **P = 0.005 (g); 12 h *P = 0.029, 24 h *P = 0.039, refed **P = 0.003 (h). i,j, Dilp8M100727 and Lgr3Δ50 mutants showed increased food intake compared to the w− control flies in the blue dye feeding assay (i) and CAFE assay (j): n = 6 biologically independent experiments, Dilp8M100727 **P = 0.0067, Lgr3Δ50 **P = 0.007 (i); n = 3 biologically independent experiments (j). Data are presented as mean ± s.e.m. Statistical significance was determined with a two-tailed Student’s t-test; *P < 0.05, **P < 0.01, ***P < 0.001.
Recent studies show that the Lgr3 relaxin protein is a Dilp8 receptor, which functions in the homeostatic control of organ size during development38,40,41. To determine whether Dilp8 and Lgr3 have any function in the regulation of feeding behaviour, we examined the mRNA expression of these molecules in response to food deprivation and performed a feeding assay with Dilp8 and Lgr3 mutants. Dilp8 and Lgr3 expression levels gradually decreased in response to starvation and rapidly increased after refeeding (Fig. 2g,h). To test food intake in Dilp8 and Lgr3 mutants, the Lgr3Δ50 deletion mutant was generated by CRISPR-Cas9-mediated mutagenesis. Lgr3Δ50 was a strong hypomorphic mutant (Extended Data Fig. 2f,g). Dilp8M100727 and Lgr3Δ50 mutant flies showed significantly increased food intake in blue dye and CAFE assays (Fig. 2i,j). GMR>ykiSA,Dilp8MI00727 and GMR>ykiSA,Lgr3Δ50 flies reversed the cancer anorexia phenotype observed in GMR>ykiSA flies (Extended Data Fig. 2h). Conversely, Dilp8 overexpression flies (GMR>Dilp8) showed reduced food intake (Extended Data Fig. 2i). These results reveal a metabolic function of Dilp8-Lgr3 signalling in the regulation of food intake.
Lgr3 is predominantly expressed in the adult head, and its expression in the head is dramatically increased in the cancer condition (Extended Data Fig. 2j), indicating that Lgr3 can receive the ligand signal in the adult head. We found that the Dilp8 and Lgr3 mutations could not reverse the tumour phenotype caused by yki overexpression (Extended Data Fig. 2k and Supplementary Fig. 2a), implying that the Dilp8-Lgr3 axis is not involved in autonomous cancer growth.
Dilp8-Lgr3 signalling regulates anorexigenic NUCB1 and orexigenic sNPF and NPF expression in the Drosophila brain
To identify the feeding regulators that suppress food intake in response to Dilp8 secretion, we examined the expression levels of Drosophila neuropeptides involved in feeding behaviour by RNA-seq analysis in GMR>ykiSA flies (Supplementary Table 1). The results showed alterations in the expression of NUCB1, a homologue of the mammalian anorexigenic factor NUCB2, and the orexigenic neuropeptides sNPF and NPF in cancer. Consistent with RNA-seq data, the mRNA level of NUCB1 was increased and sNPF and NPF levels were decreased in GMR>ykiSA flies (Fig. 3a–c).
a–c, The relative mRNA level of NUCB1 (a) was increased (**P = 0.0075), and sNPF (b) (**P = 0.0097) and NPF (c) mRNA levels (*P = 0.010) were decreased in GMR>ykiSA. n = 3 biologically independent experiments. d, NUCB1 mRNA levels were decreased, whereas sNPF and NPF mRNA levels were increased in Dilp8MI00727 and Lgr3Δ50 mutants. n = 3 biologically independent experiments; NUCB1, Dilp8MI00727 *P = 0.012, Lgr3Δ50 *P = 0.015; sNPF, Dilp8MI00727 *P = 0.016 and Lgr3Δ50 **P = 0.005; NPF, Dilp8MI00727 **P = 0.004, Lgr3Δ50 *P = 0.023. e,f, NUCB1 mRNA levels were increased, whereas sNPF and NPF mRNA levels were decreased, in Dilp8-overexpressing (GMR>Dilp8) (e) and Lgr3-overexpressing (Lgr3>Lgr3) (f) flies. n = 3 biologically independent experiments: NUCB1 **P = 0.0026, sNPF *P = 0.0138, NPF *P = 0.014 (e); NUCB1 **P = 0.006, sNPF *P = 0.017, NPF *P = 0.011 (f). g, Drosophila NUCB1 mRNA expression was decreased under the starvation condition and was increased by refeeding. n = 3 biologically independent experiments; 6 h *P = 0.002, 12 h *P = 0.007, 24 h *P = 0.007, refed *P = 0.011. h, The relative food intake was reduced when nesfatin-1 (a processed form of NUCB1) was overexpressed (MJ94>Nesfatin1) and increased when NUCB1 was inhibited in neurons (MJ94>NUCB1-RNAi). n = 4 biologically independent experiments; MJ94>Nesfatin1 *P = 0.035, MJ94>NUCB1-RNAi *P = 0.022. Data are presented as the mean ± s.e.m. Statistical significance was determined with two-tailed Student’s t-test; *P < 0.05, **P < 0.01.
To determine whether Dilp8-Lgr3 signalling regulates the mRNA levels of NUCB1, sNPF and NPF, the mRNA expression of NUCB1, sNPF and NPF was examined in the heads of Dilp8 and Lgr3 mutant and overexpressing flies. NUCB1 mRNA levels were decreased in the Dilp8 mutant (Dilp8MI00727) and increased when Dilp8 was overexpressed. By contrast, sNPF and NPF mRNA levels were increased in the Dilp8 mutant and reduced in response to Dilp8 overexpression (Fig. 3d,e). Consistent with these results, NUCB1, sNPF and NPF mRNA expression levels showed the same expression pattern in Lgr3Δ50 mutant and Lgr3-overexpressing flies (Fig. 3d,f). To show Lgr3 can regulate sNPF and NPF mRNA expressions cell-autonomously because Lgr3-Gal4 expression is overlapped with a small subset of sNPF neurons (Extended Data Fig. 2l and Supplementary Fig. 2b), we crossed UAS-sNPF RNAi and UAS-NPF RNAi flies with the Lgr3-Gal4 line. Both Lgr3>sNPF RNAi and Lgr3>NPF RNAi showed significant knockdown of mRNA levels (Extended Data Fig. 2m,n). Taken together, these results indicate that the anorexigenic peptide NUCB1 and the orexigenic neuropeptides sNPF and NPF are inversely regulated by tumour-derived Dilp8 and its receptor Lgr3 to induce cancer anorexia.
Drosophila NUCB1 is a homologue of mammalian NUCB2, which is described as a satiety molecule42. However, the biological function of Drosophila NUCB1 has not been studied. NUCB2 and Drosophila NUCB1 show 62% amino-acid similarity. The signal peptide and dibasic cleavage sites are highly conserved (Extended Data Fig. 3), suggesting that nesfatin-1, the 82-amino-acid anorexigenic peptide processed from NUCB2, can also be produced via NUCB1 processing. We tested whether the role of NUCB1 is also involved in feeding suppression. Consistent with mammalian NUCB242, Drosophila NUCB1 mRNA expression in the adult head was decreased under the starvation condition and increased by refeeding (Fig. 3g). Nesfatin-1 (a processed form of NUCB1) overexpression in neurons (MJ94>Nesfatin1) inhibited food intake; conversely, NUCB1 inhibition in neurons (MJ94>NUCB1-RNAi) increased food intake (Fig. 3h). These results demonstrate that NUCB1 has an evolutionarily conserved role as an anorexigenic factor both in Drosophila and mammals.
NUCB1- and Lgr3-expressing neurons are markedly increased and overlapped in the cancer condition
To determine whether secreted Dilp8 and its receptor Lgr3 systemically regulate NUCB1 in the target organ, the expression pattern of NUCB1 in neurons was examined in the adult brain of GMR>ykiSA. A Drosophila NUCB1 antibody was generated for immunostaining and its specificity was assessed by western blot (Extended Data Fig. 4a). In addition, the co-localization of NUCB1 and Lgr3 was assessed to determine whether NUCB1-expressing neurons directly receive tumour-derived Dilp8 signals through the Lgr3 receptor. In the normal non-tumour condition, endogenous NUCB1 was expressed at a low level in few neuronal cells (Fig. 4a,b, arrowheads, Extended Data Fig. 4b,c and Supplementary Fig. 3a). Lgr3 expression in the adult brain was also weak, and the neuronal cells were difficult to distinguish at the endogenous level (Fig. 4c, arrowheads, and Extended Data Fig. 4c). The signal intensity of both NUCB1 and Lgr3 neurons was significantly increased in the cancer condition (GMR>ykiSA) and the signals showed perfect overlap (Fig. 4e–h, boxes, 4i–k, arrows, Extended Data Fig. 4d and Supplementary Fig. 3b), suggesting that Lgr3 and NUCB1 double-positive neurons are anorexigenic neurons in the cancer condition. These findings were supported by the almost undetectable expression of Lgr3 and low level of NUCB1 in the Lgr3 mutant (Lgr3Δ50) in the cancer condition, which were similar to the levels in the control (Fig. 4l–o, arrowheads, Extended Data Fig. 4e and Supplementary Fig. 3c). Additionally, NUCB1-expressing neurons in the cancer condition overlapped with a subset of sNPF and NPF neurons (Extended Data Fig. 4f,g, arrows, and Supplementary Fig. 4a,b). These results demonstrated that tumour-derived Dilp8 effectively regulates NUCB1, sNPF and NPF expression through the Lgr3 receptor in NUCB1- and Lgr3-expressing neurons.
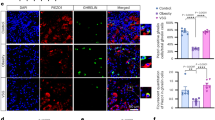
a–d, Immunostaining with antibodies against NUCB1 and Lgr3 showed that a small number of neuronal cells expressed low levels of endogenous NUCB1 and Lgr3 in the adult brain in normal non-tumour condition (arrowheads): nc82 (a); NUCB1 (b); Lgr3 (c); NUCB1 and Lcr3 (d). e–h, In the cancer condition caused by GMR>ykiSA, NUCB1 and Lgr3 immunostaining was significantly increased in the same neuronal cells (arrows): nc82 (e); NUCB1 (f); Lgr3 (g); NUCB1 and Lcr3 (h). i–k, Enlarged image of the white dashed boxes in f–h, showing increased intensity and overlapped NUCB1- and Lgr3-expressing neurons (arrows). l–o, The Lgr3 mutant in the yki overexpression condition (GMR>ykiSA, Lgr3Δ50) showed almost undetectable expression of Lgr3 and a low level of NUCB1 (arrowheads): nc82 (l); NUCB1 (m); Lgr3 (n); NUCB1 and Lcr3 (o). Scale bars, 100 μm. p, Quantitative PCR of adult heads showed that NUCB1 mRNA levels were increased by approximately twofold in GMR>ykiSA (*P = 0.021), reduced in the Lgr3Δ50 mutant (*P = 0.019) and restored to the control level in GMR>ykiSA, Lgr3Δ50 (*P = 0.049). n = 3 biologically independent experiments. q,r, Quantitative PCR of adult heads showed that sNPF (q) and NPF (r) mRNA levels were reduced to less than half of the control (GMR-Gal4) in GMR>ykiSA (**P = 0.001 (q), **P = 0.004 (r)), increased in the Lgr3Δ50 mutant (*P = 0.006 (q); *P = 0.045 (r)) and restored in GMR>ykiSA, Lgr3Δ50 (**P = 0.028 (q); *P = 0.027 (r)). n = 3 biologically independent experiments. s–u, Dilp8 treatment increased NUCB1 mRNA levels (s) and decreased sNPF (t) and NPF (u) mRNA levels in the ex vivo culture. n = 3 biologically independent experiments: 0.5 h **P = 0.002, 1 h **P = 0.0097, 2 h **P = 0.012 (s); 0.5 h **P = 0.007, 1 h ***P = 0.0009, 2 h **P = 0.003 (t); 5 h **P = 0.0088, 1 h **P = 0.0023, 2 h ***P = 0.0004 (u). Data are presented as the mean ± s.e.m. Statistical significance was determined with a two-tailed Student’s t-test; *P < 0.05, **P < 0.01, ***P < 0.001.
We further confirmed NUCB1, sNPF and NPF expression in the cancer condition by quantitative polymerase chain reaction with reverse transcription (qRT-PCR). The increased NUCB1 expression in the GMR>ykiSA cancer model was significantly suppressed in the Lgr3 mutant (Fig. 4p). Consistent with the qRT-PCR results, western blot analysis showed that the upregulation of the NUCB1 protein in the cancer condition was restored to the control level in the Lgr3 mutant (Extended Data Fig. 5a,b). Conversely, the downregulation of sNPF and NPF mRNA in the cancer condition was restored by the Lgr3 mutation (Fig. 4q,r).
Because tumour-derived Dilp8 systemically controls the expression of NUCB1, sNPF and NPF in the brain, we investigated whether these neuropeptides are regulated by Dlip8. An ex vivo culture assay was performed by adding synthetic Dilp8 peptide to the culture medium of dissected fly adult brains (Extended Data Fig. 5c). NUCB1 mRNA expression was upregulated at 0.5 h and the levels were maintained for >2 h (Fig. 4s). sNPF and NPF were downregulated in response to the Dilp8 peptide (Fig. 4t,u). Notably, Lgr3 expression was upregulated at early time points (Extended Data Fig. 5d), consistent with the immunostaining results showing that Lgr3 neurons were upregulated in the cancer condition compared with the levels in the non-cancer normal condition. Taken together, these data support our previous results that NUCB1, sNPF and NPF are regulated by Dilp8-Lgr3 signalling to induce cancer anorexia.
INSL3-LGR8 signalling regulates anorexigenic Nucb2 and orexigenic Npy expression in mouse hypothalamic cells
To identify a Dilp8 homologue in mammals, we searched for a mammalian Lgr3 homologue by phylogenetic comparative analysis. Mammalian Lgr7/8 (RXFP2, relaxin/insulin-like family peptide receptor 2) was identified as a homologue of Drosophila Lgr3 (Extended Data Fig. 6a). Lgr8 is a receptor for INSL3 and Lgr7 is a receptor for H2 relaxin43. A separate phylogenetic comparative analysis showed that Dilp8 and INSL3 are closely related peptides (Extended Data Fig. 6b). Comparison of the mature structures of INSL3 and Dilp8 showed that Dilp8 and INSL3 have six conserved invariant cysteine residues for disulfide bond formation (Fig. 5a). These findings indicate that Drosophila Dilp8-Lgr3 signalling may be conserved in the mammalian INSL3-Lgr8 pathway.
a, The structural homology of human INSL3 and fly Dilp8. b, Treatment in the INSL3 peptide increased the Nucb2 mRNA level (1 h **P = 0.0037, 2 h **P = 0.0036) and decreased the Npy mRNA level (1 h *P = 0.029, 2 h *P = 0.018) in mHypoE-N3 cells. n = 3 biologically independent experiments. c, No significant changes in Nucb2 and Npy mRNA levels were detected after INSL3 treatment in mHypoE-N39 cells. n = 3 biologically independent experiments. d,e, Knockdown of Lgr8 in N3 cells showed no significant changes in Nucb2 (d) (control **P = 0.0019, siLgr8 NS = 0.698) and Npy (e) mRNA levels (control **P = 0.0035, siLgr8 NS = 0.431) by INSL3 treatment. n = 3 biologically independent experiments. f,g, Lgr8 overexpression in N39 cells showed increased Nucb2 (f) (*P = 0.0195) and decreased Npy mRNA level (g) (*P = 0.034) with INSL3 treatment. n = 3 biologically independent experiments. Data are presented as the mean ± s.e.m. Statistical significance was determined with two-tailed Student’s t-test; *P < 0.05, **P < 0.01.
The hypothalamus is a feeding centre in mammals and it expresses numerous neuropeptides involved in the regulation of feeding behaviour44,45. Lgr8 protein and mRNA were predominantly expressed in the hypothalamus and hippocampus of the mouse brain (Extended Data Fig. 6c,d and Supplementary Fig. 5). To test whether INSL3 regulates Nucb2 and Npy expression through the Lgr8 receptor, mouse hypothalamic mHypoE-N3 cells, which express Lgr8 (Extended Data Fig. 6e), were treated with INSL3. Consistent with the fly data, INSL3-treated N3 cells showed increased NUCB2 and reduced NPY expression (Fig. 5b). However, in mHypoE-N39 cells, which express low levels of Lgr8 (Extended Data Fig. 6e), INSL3 treatment had no significant effect on Nucb2 and Npy expression (Fig. 5c). In the Lgr8 knockdown N3 cells with Lgr8 siRNA, INSL3 treatment had no significant effect on Nucb2 and Npy expression (Fig. 5d,e). Conversely, in Lgr8-overexpressed N39 cells obtained by the transfection of Lgr8, INSL3 treatment showed increased Nucb2 and reduced Npy expression (Fig. 5f,g). These results indicate that Nucb2 and Npy are regulated by INSL3-Lgr8 signalling in mammalian cells.
INSL3 regulates Nucb2 and Npy, and induces anorexia in the mouse tumour model
To test cancer cachexia-anorexia in the mouse model, we generated tumour-implanted mouse models that were subcutaneously (s.c.) implanted with two different cancer cell lines, C26 (colon carcinoma cells) and LLC (Lewis lung carcinoma cells), and monitored for food intake and body weight changes. After two weeks of tumour inoculation, C26 implanted mice showed body weight loss (Extended Data Fig. 6f), but not LLC implanted mice (Extended Data Fig. 6g), even though the tumour weights of both tumours were similar (Extended Data Fig. 6h). Daily food intake was significantly reduced in C26 implanted mice (Fig. 6a), but not in LLC implanted mice (Fig. 6b). Interestingly, the level of secreted INSL3 in the conditioned media of C26 cells was higher than that of the LLC cells (Fig. 6c). In the C26 implanted mice model, daily food intake and body weight were measured. From day 11, food intake was significantly decreased but body weight was unchanged (Fig. 6d,e). When we analysed the tissue wasting, muscle wasting and its molecular markers (MuRF-1, MAFbx, Fbxo31 and Musa1) were unaffected at day 12, but showed an increase at day 21 (Fig. 6f,i and Extended Data Fig. 6i,j). However, lipolysis in the fat tissue and its molecular markers (Atgl and Hsl) were increased at day 12 and day 21 (Fig. 6f,i and Extended Data Fig. 6i,j). In the hypothalamic tissues of C26 tumour implanted mice, Nucb2 mRNA levels were increased at day 12 and 21 (Fig. 6g,j), but Npy expression was decreased only at day 21 (Fig. 6h,k). However, no such changes were observed in the hypothalamic sample of LLC tumour implanted mice (Extended Data Fig. 6k,l). These results indicate that cancer anorexia precedes the appearance of muscle wasting in the C26 tumour implanted mice model.
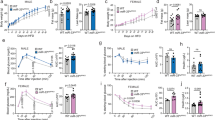
a,b, Food intake was reduced in C26 tumour mice (a) (***P = 2.49 × 10−8) but not in LLC tumour mice (b) (NS = 0.295). n = 7 mice. c, A higher secreted level of INSL3 was seen in C26 culture medium than in LLC medium. n = 3 biologically independent experiments, **P = 0.026. d, Food intake was significantly reduced from day 11 in C26 tumour implanted mice. Control n = 6 mice, C26 n = 8 mice; day 11 *P = 0.013, day 12 NS = 0.078, day 13 ***P = 0.0009. e, Body weight was unchanged in C26 tumour mice until day 13. Control n = 6 mice, C26 n = 8 mice. f, Muscle wasting (MuRF-1, MAFbx, Fbxo31 and Musa1) and fat lipolysis (Atgl and Hsl) molecular markers of C26 tumour mice at day 12 in gastrocnemius (GCM) and epididymal white adipose tissue (eWAT). Control n = 5 mice, C26 n = 6 mice; Atgl **P = 0.0017; Hsl ***P = 0.0003. g,h, Significantly increased Nucb2 (g) (*P = 0.017) and unchanged Npy (h) (NS = 0.164) mRNA levels in C26 tumour mice hypothalamus at day 12. Control n = 5 mice, C26 n = 5 mice. i, Increased muscle wasting and fat lipolysis molecular markers of C26 tumour mice at day 21. Control n = 5 mice, C26 n = 6 mice; MuRF-1 *P = 0.033, MAFbx *P = 0.042, Fbxo31 NS = 0.082, Musa1 NS = 0.368, Atgl *P = 0.022, Hsl **P = 0.005. j,k, Significantly increased Nucb2 (j) (*P = 0.035) and decreased Npy (k) (**P = 0.008) mRNA levels in C26 tumour mice hypothalamus at day 21. Control n = 5 mice, C26 n = 5 mice. l, The synthetic INSL3 peptide i.c.v. injection reduced food intake at both 5 μg μl−1 (*P = 0.011) and 1 μg μl−1 (**P = 0.009). n = 9 mice. m–o, The synthetic INSL3 peptide i.p. injected LLC tumour-bearing mice showed reduced food intake (m) (PBS n = 13 mice, INSL3 n = 13 mice; ***P = 0.0007) and increased Nucb2 mRNA level (n) (**P = 0.002), but the Npy mRNA level was unchanged (o) (PBS n = 3 mice, INSL3 n = 4 mice). Data are presented as the mean ± s.e.m. Statistical significance was determined with two-tailed Student’s t-test; *P < 0.05, **P < 0.01, ***P < 0.001.
To address whether INSL3 regulates food intake, we then measured the food intake of mice after intracerebrospinal (i.c.v.) injection of artificial CSF (aCSF) or INSL3. The food intake was significantly reduced in INSL3-injected mice compared with aCSF-injected mice (Fig. 6l), indicating that INSL3 can reduce food intake by functioning in the brain. We next tested whether peripheral INSL3 regulates food intake by delivering INSL3 peptide intraperitoneally (i.p.) and measuring the food intake. However, i.p. INSL3 did not induce significant changes in food intake or gene expression of Nucb2 and Npy in the hypothalamus (Extended Data Fig. 6m–o). We hypothesized that systemic changes induced by the tumour may be required for CNS actions of peripherally delivered INSL3. To test this notion, we performed i.p. injection of INSL3 to LLC tumour-bearing mice that did not show anorexia. These experiments resulted in significantly reduced food intake and increased Nucb2 mRNA expression in the hypothalamus (Fig. 6m–o). Taken together, these results suggest that cancer-derived INSL3 causes anorexia in mammals, similar to the effect of tumour-derived Dilp8 in Drosophila.
Serum INSL3 levels correlate with anorexia severity in patients with pancreatic cancer
Approximately 80% of pancreatic ductal adenocarcinoma (PDAC) patients present the cancer anorexia-cachexia syndrome46,47. To test cancer anorexia in a mouse pancreatic cancer model, NSG mice were s.c. implanted with two different human-derived pancreatic cancer cell lines, Capan1 or Panc1, and food intake and anorexia-associated body mass were analysed. Tumour volume was gradually increased in both tumour-implanted mice (Extended Data Fig. 7a). Reduced food intake was observed at five weeks in the Capan1-implanted group compared with that in the non-tumour-implanted control or Panc1-implanted group (Fig. 7a). The wasting phenotype was observed in both tumour-implanted mice starting at five weeks after implantation (Extended Data Fig. 7b). The secreted INSL3 level was dramatically higher in Capan1 cancer cell culture media than in HPDE controls and Panc1 cells (Fig. 7b). In Capan1 cells, the INSL3 expression level was significantly high among relaxin-INSL family genes (Extended Data Fig. 7c). These results confirm that INSL3 is an evolutionarily conserved tumour-derived factor for inducing anorexia in cancer.
a, The food intake of Capan1-implanted mice at five weeks was decreased compared with that in control or Panc1-implanted mice. n = 4 mice for each group, **P = 0.009. b, The secreted level of INSL3 in Capan1 culture media was remarkably increased compared with those in the control. n = 4 biologically independent experiments, ***P = 0.0001. c, A highly increased INSL3 level was detected in the serum of patients with pancreatic cancer. One-way analysis of variance (ANOVA) *P = 0.040. d, The protein level of INSL3 was increased in pancreatic tumour tissue. Scale bars, 100 μm. e,f, Unlike the non-PDAC group (e), patients in the PDAC group showed a decrease in food intake with increasing INSL3 levels (f). g, The INSL3 serum level was increased significantly in PDAC patients with various degrees of anorexia than in PDAC patients without anorexia. n = 6 anorexia-absent patients, n = 13 anorexia-present patients, *P = 0.029. h, Our model of the cancer anorexia mechanism. Data are presented as the mean ± s.e.m. Statistical significance in a, b and g was determined with a two-tailed Student’s t-test, and in c, e and f with one-way ANOVA; *P < 0.05, **P < 0.01, ***P < 0.001.
We compared INSL3 levels in the blood of patients with pancreatic cancer (n = 19), patients with pancreatic benign disease (n = 5) and patients with non-pancreatic benign disease (n = 7) (Supplementary Table 2). This revealed that INSL3 serum level was significantly increased in patients with pancreatic cancer compared to other benign disease groups (Fig. 7c). Increased INSL3 expression was also found in pancreatic tumour tissue compared to normal tissue (Fig. 7d and Supplementary Fig. 6). In addition, we checked the correlation of INSL3 serum levels and feeding problems in a human PDAC patient group (n = 19) and a non-PDAC patient group (n = 8) (Supplementary Table 3). There was a negative correlation between serum INSL3 level and calorie intake in PDAC patients (R2 = 0.206), but a significant positive correlation in non-PDAC patients (R2 = 0.0991) (Fig. 7e,f). Also, INSL3 serum level was increased significantly in PDAC patients with various degrees of anorexia compared with PDAC patients without anorexia (Fig. 7g). These results indicate that tumour-derived INSL3 is associated with the induction of cancer anorexia in patients with pancreatic cancer.
Discussion
In this study, we have examined the cancer anorexia mechanism induced by the secretion of Dilp8/INSL3 from tumour tissues, which systemically controls the expression of feeding peptide hormones through the Lgr3/Lgr8 receptor in the brain (Fig. 7h). Dilp8 secretion in our yki tumour model was strongly supported by a recent study showing that Yorkie (yki) and its binding partner Scalloped directly regulate Dilp8 expression48. The present data demonstrate that Dilp8 and its mammalian orthologue INSL3 are critical factors involved in cancer anorexia in Drosophila and mammals.
Two recent studies using fly cancer models reported that the inhibition of insulin signalling induces systemic organ wasting, a characteristic of cancer cachexia27,28. Both papers reported that ImpL2, a mammalian IGFBP that is secreted from hyper-proliferating tissues and tumours, causes systemic organ wasting independently of food consumption. Consistent with previous data, our fly cancer model showed the organ wasting phenotype, and the severity of the wasting phenotype increased time-dependently (Fig. 1c,d). Importantly, the anorexia phenotype was observed at early stages, consistent with the fact that most cancer patients show loss of appetite at the pre-cachexia stage4. Although the inhibition of Dilp8 does not restore the cancer-induced organ-wasting phenotype27,28, Dilp8 inhibition rescued the cancer anorexia phenotype (Fig. 2d), suggesting that Dilp8 and ImpL2 play critical roles in regulating cancer anorexia and cachexia, respectively.
Another Drosophila tumour-secreted factor, Upd2, does not cause organ wasting27,28 or anorexia in flies (Fig. 2d), whereas mouse IL-6 (equivalent to Upd2) induces white adipose tissue browning in the cachexia condition49. A recent report shows that tumour-derived IL-6/Upd induces non-cell-autonomous autophagy around tumour tissues50. This paper suggests that tumour cells can acquire nutrition by eating adjacent cells for growth and proliferation.
We discovered a fly anorexigenic hormone, Drosophila NUCB1, a homologue of mammalian NUCB2. Nesfatin-1, a processed form of NUCB2, was initially discovered as a potential anorexigenic modulator of food intake and body weight independent of leptin signalling42,51,52,53. The NUCB2 precursor protein is expressed in the hypothalamic region, including the supraoptic nucleus, lateral hypothalamic area, arcuate nucleus, paraventricular nucleus42 and parabrachial nucleus (PBN)54. Recent findings indicate that calcitonin gene-related peptide (CGPR)-positive neurons in the PBN region contribute to cancer-induced appetite suppression in mice implanted with LLC cells55. The Drosophila NUCB1/nesfatin-1 described in this study is identified as an evolutionarily conserved anorexigenic molecule expressed in the fly brain. We propose that NUCB1-expressing neurons in the brain are important for anorexigenic regulation in normal and cancer conditions. Despite substantial evidence supporting that nesfatin-1 binding to G protein-coupled receptors increases the cAMP response56,57,58, a receptor specific to NUCB1/nesfatin-1 remains to be identified.
In this Article, we have demonstrated that tumour-secreted Dilp8 induces cancer anorexia through the Lgr3 receptor, and INSL3 and Lgr8 are predicted to be mammalian orthologues of Dilp8 and Lgr3, respectively. We have also showed that the regulation of INSL3 in mice is conserved in cancer anorexia. The secreted INSL3 in cell culture medium and INSL3 serum level were increased in the mouse cancer model and in human cancer patients, respectively. Consistently, food intake was reduced when INSL3 was injected into the brain of mice. Taken together, the fly and mammalian data support our cancer anorexia model (Fig. 7h). These findings also suggest that Dilp8/INSL3-Lgr3 signalling is a potential therapeutic target in cancer anorexia.
Methods
Drosophila culture and stocks
Drosophila melanogaster were maintained at 25 °C on standard media. GMR-Gal4, UAS-ykiWT, UAS-ykiS168A, NUCB1-RNAi, Dilp8MI00727 were obtained from the Bloomington Stock Center and Dilp8-RNAi, ImpL2-RNAi and Upd2-RNAi were obtained from Vienna Drosophila RNAi Center (VDRC). UAS-Dilp8 and UAS-Lgr3 were gifts from A.M. Gontijo (NOVA Medical School) and Lgr3-Gal4 was a gift from P. Leopold (Institute of Biology Valrose). The pUAS-Nesfatin1 transgenic fly was generated by the p-element-mediated germline transformation method59 containing the processed form of NUCB1 cDNA.
CRISPR/Cas9-mediated mutagenesis
The Lgr3 mutant was generated by CRISPR/Cas9-mediated mutagenesis60. The single-guide RNA target sequence was designed and generated by Toolgen (toolgen.com). Mutant screening by T7E1 assay and direct sequencing followed the protocol described in ref. 61. The Lgr3 mutant consists of a 50-base-pair deletion upstream of the target sequences. Capital letters indicate the second exon of the Lgr3 genomic region, bold capital letters indicate target sequences and underlined lower-case letters indicate deleted sequences:
ctaggaaattataaaaggtacaaacaaaagatattaaaatgtttcaaagacacaaatgtttgcttgatggctaacagctctaggaaatatatattttggttaaaatcatcactagattttgttttgatttcccccaaaatctgttgagaaactagttgaactcacatgcaatatgctccgctgtccttgagtgataattatttgggatcgtccttttcccagGACCCTGCCCCGCGGCCAGCTTCGCCTGCGACAACGGCACGCTGTGCGTTCCAAGGCGCCAGATGTGCGACAGTCGGAACGATTGCGCCGACAGCAGCGACGAGAATCCG
RNA preparation and quantitative real-time PCR
Adult heads from 20 flies were collected for RNA preparation. Total RNA was extracted using the easy-BLUE reagent (iNtRON Biotechnology). All RNA samples were treated with RNase-free DNase (Promega) and cDNA was synthesized using a SuperScript III First-Strand synthesis system (Invitrogen). For qRT-PCR analysis, an ABI Prism 7900 sequence detection system (Applied Biosystems) and SyberGreen PCR Core reagents (Applied Biosystems) were used. mRNA levels were expressed as the relative fold change against the normalized rp49 mRNA. The comparative cycle threshold (Ct) method (User Bulletin 2, Applied Biosystems) was used to analyse the data.
RNA-sequencing analysis
To extract total RNAs for RNA-seq experiments, 30 adult heads were dissected and isolated with easy-BLUE reagent (iNtRON Biotechnology). RNA quality control and data analyses were performed by ebiogen (ebiogen.com).
Generation of the NUCB1 and Dilp8 antibody
NUCB1 antibody was generated by the immunization of rabbits with the synthetic peptide (TQNKKDHKEAAESSTPATAD) and the immunization of guinea pigs with the synthetic peptide (EEKHKDHEKLHHPGNKAQ) of the NUCB1 protein (Abfrontier). Dilp8 antibody was generated by the immunization of rats with the same peptide sequence as described in ref. 62.
Immunostaining
For immunostaining, adult brains were dissected in phosphate buffered saline (PBS), fixed in 4% paraformaldehyde (PFA) and blocked in 5% bovine serum albumin (BSA). Primary antibodies were incubated overnight at 4 °C and secondary antibodies were incubated for 2 h at room temperature. The tissues were mounted with Vectashield mounting medium (Vectashield) and fluorescence images were acquired by a FluoView confocal microscope (Olympus). sNPF (1:200), NUCB1 (1:100), Lgr3 (1:200, a gift from M. Dominguez), NPF (1:2,000, a gift from P. Shen), brp (nc82, 1:100, DSHB) primary antibodies and anti-mouse immunoglobulin-G (IgG) Alexa 488, anti-guinea pig IgG Alexa 594 and anti-rabbit Alexa 633 secondary antibodies (1:200, Molecular Probes) were used.
Western blot analysis
Total proteins from adult heads were lysed by PRO-PREP protein extraction buffer (iNtRON biotechnology). Western blot analyses were performed as described previously63. β-actin (1:3,000, DSHB), Lgr8 (1:1,000, Novus), NUCB1 (1:1,000) and Dilp8 (1:1,000) primary antibodies and anti-rabbit IgG (1:5,000, Santa Cruz), anti-rat IgG (1:5,000, Invitrogen) and anti-mouse IgG (1:5,000, Invitrogen) secondary antibodies were used.
Measurement of Drosophila life span
Flies used in life span experiments were collected upon eclosion over a 48-h period, with a total of 20 vials (n ≈ 200) for each genotype. Flies were transferred to fresh food every other day, and dead flies were scored and counted. Lifespan analyses were performed and Log rank test statistics calculated using the online application OASIS264.
Drosophila food intake assay
Drosophila food intake was measured by two different assays. The CAFE assay37 was performed with three-day-old adult male flies. Four flies were placed in the CAFE device37 containing a 5% sucrose solution in calibrated glass micropipettes (VWR) and adapted for 12 h before the assay. At time zero, the micropipettes with 5% sucrose solution were replaced, and the amount of liquid consumed was measured every 12 h. For colorimetric food intake assay, flies were starved in PBS-containing vials for 2 h and fed for 15 min in vials containing 0.05% FD&C Blue dye, 7% sucrose and 5% yeast. The flies were frozen, homogenized in PBS, and centrifuged twice for 25 min each. The supernatant was measured at 625 nm. Each experiment consisted of 20 flies and the assay was repeated three times.
Measurement of total body triglyceride level
Ten adult flies were ground in PBS solution and centrifuged. The supernatant was used for analysis. Total glycerol and triglyceride levels were measured using a serum triglyceride determination kit (Sigma). The protein levels were measured to normalize the triglyceride level.
Drosophila starvation
Twenty w− adult flies were starved overnight and fed for 2 h to achieve physiological synchronization. Then, starvations for the experiment were started until each time point. The heads from starved flies were collected for qPCR analysis.
Ex vivo culture assay
The adult brains were dissected and cultured in 100 μl of Schneider’s medium (Sigma) with or without Drosophila Dilp8 peptide (Phoenix) at room temperature and collected for RNA preparation.
Cell culture and mouse models
The embryonic mouse hypothalamic cell lines mHypoE-N3 and mHypoE-N39, purchased from Cellutions Biosystems, were maintained at 37 °C, 5% CO2 in Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and antibiotics. Cells were treated with 100 nM mouse INSL3 peptide (Phoenix Pharmaceutical) after 1 h of starvation. mHypoE-N3 cells were transfected with siRNA targeting mouse Lgr8 (Bioneer, 140498-1, 140498-2, 140498-3) using Lipofectamine RNAiMAX (Invitrogen, 13778150) for 48 h according to the manufacturer’s protocol and then treated with vehicle or INSL3 for 2 h.
C26 and LLC were obtained from the American Type Culture Collection (ATCC). C26 cells were maintained in RPMI-1640 (Hyclone) containing 10% FBS (Hyclone) and 1% antibiotic–antimycotic solution (AA; Gibco/Thermo Fisher Scientific). LLCs were maintained in DMEM (Hyclone) containing 10% FBS and 1% AA.
Seven-week-old female Balb/c mice or C57BL/6 mice (Narabio Inc.) were housed in a specific pathogen-free animal facility (KAIST, Daejeon, Korea) in a temperature-controlled environment at 23 °C under a 12-h:12-h light–dark cycle. For tumour-implanted mouse models, 5 × 105 C26 cells (into Balb/c) or LLC cells (into C57BL/6) were administrated a s.c. injection in 100 µl of sterilized PBS (tumour-implanted group) or 100 µl of sterilized PBS without cells (control group). To identify the effects of C26 or LLC tumour growth on body weight and food intake, body weight was measured every other day and food intake was weighed daily after two weeks of tumour inoculation when C26 mice displayed symptoms of body weight loss. Mice were euthanized at 21 days after tumour cell injection. Organs were collected and used for further analysis. In each animal experiment, mice were randomly selected to each group. All mouse experiments were approved by the Institutional Animal Care and Use Committee (IACUC) of Korea Advanced Institute of Science and Technology.
Human pancreatic adenocarcinoma Capan1 and Panc1 cell lines were obtained from the ATCC, maintained in the ATCC recommended growth media, and incubated at 37 °C, 5% CO2. Briefly, Capan1 and Panc1 cells were grown in Iscove’s modified Dulbecco’s medium with 10% FBS and antibiotics. For tumour-implanted mouse models, NOD.Cg-Prkdcscid Il2rgtm1Wji/SzJ (NSG) mice (The Jackson Laboratory) were housed in a pathogen-free animal facility at KRIBB and used for tumour implantation studies. Six-week-old male NSG mice were implanted with 5.0 × 105 Capan1 or Panc1 cells s.c. in their abdomens. Weight and food intake were measured weekly. Tumour measurements were performed every week with a caliper, and volumes were calculated using the following formula: V = 1/2(length (mm) × width (mm)2). This study is compliant with all relevant ethical regulations regarding animal research.
RNA fluorescence in situ hybridization
RNA fluorescence in situ hybridization (FISH) was performed using an RNAscope multiplex fluorescence kit (ACDbio, 323133). In brief, mice were anaesthetized with Zoletil (Virbac Korea, Zoletil 50) and transcardially perfused with ice-cold PBS followed by 1% PFA. Brains were extracted, post-fixed overnight in 4% PFA at 4 °C, then 40-μm-thick coronal brain sections were obtained on a vibratome (Leica, VT1200S) and collected in six equal series. Slices were stored at −20 °C in hand-made anti-freezing solution (poly and glycerol mixture) until RNA FISH was performed. The sections were removed from anti-freezing solution and washed six times in Tris-buffered saline, for 10 min. After washing, sections were mounted directly onto Histobond (Marienfeld, 0810001) slides and the sections were dried at 60 °C overnight. The following day, slides were dehydrated with 100% ethanol and dipped in boiling target retrieval solution for 20 min. A hydrophobic barrier was created and slides were treated with protease III for 30 min at 40 °C. The probe against Rxfp2 mRNA (ACDbio, 589261-C3) was added into slides. After 3 h of incubation at 40 °C, hybridize-Amp probes #1-4 were applied on slides sequentially to amplify the signal. For every RNA FISH run, the Escherichia coli gene dapB was used as a negative control, and Mm-Polr2a, Mm-PPIB and Mm-UBC were used as housekeeping genes. All sections were counterstained with DAPI (Invitrogen, 62248). Fluorescent images were captured using a Zeiss LSM 880 confocal scanning microscope (Carl Zeiss, LSM 880). Contrast and brightness were corrected using ZEN blue software.
Enzyme-linked immunosorbent assay
Cultured media for C26, LLC, Capan1, Panc1 and HPDE (human pancreatic duct epithelial) cell lines and patients’ blood were used for INSL3 detection. The process was performed as described in the INSL3 enzyme-linked immunosorbent assay kit datasheet from MyBioSource Inc. All samples were measured at individual levels and the results were from more than three independent experiments.
Stereotaxic surgery and intracerebrospinal injection
A guide cannula (26 G; Plastics One, C315GS-5/SPC) was inserted into the lateral ventricle (1.0 mm lateral; −0.7 mm posterior to bregma; 1.3 mm below the skull surface) of mice using a stereotaxic apparatus (KOPF, model 900LS). Mice were anaesthetized with Zoletil (Virbac Korea, Zoletil 50) during stereotaxic surgery and were allowed to recover after surgery for at least seven days. Food intake and body weight were recorded during the recovery period and mice that did not regain normal weight were excluded from studies. After the recovery period, proper insertion of the cannula was assessed by measuring the drinking response to i.c.v. injection of angiotensin II (2 µg µl−1; Sigma, A9525). For i.c.v. injections to the lateral ventricle, aCSF (Tocris, 3525) or INSL3 peptide (Anygen) dissolved in aCSF at a concentration of 5 µg µl−1 or 1 µg µl−1 was injected over 60 s in a final volume of 1 µl using a 33-G injector cannula (Plastics One, C315I/SPC) extending 0.5 mm beyond the tip of the i.c.v. cannula.
Data collection from patients with pancreatic cancer
The study complied with the Declaration of Helsinki, and the protocol was reviewed and approved by the Institutional Review Board of Asan Medical Center (IRB no. 2017-0305). Written informed consent was obtained from all participants. Patient-, surgery- and oncology-related data were obtained from medical records. Peripheral blood was drawn from patients with pancreatic cancer and other diseases who were diagnosed by pathological examination at Asan Medical Center in South Korea. The plasma was separated and stored at −70 °C until analysis. Anorexia grade was measured by the Common Terminology Criteria for Adverse Events (CTCAE) method version 4.0 (https://www.uptodate.com/contents/common-terminology-criteria-for-adverse-events) and classified as absence (Grade 0) and presence (Grade 1, 2, 3). Calorie intake was measured by the 24-hour dietary recall method65,66. This study is compliant with all relevant ethical regulations regarding research involving human participants.
Statistics and reproducibility
All experiments were performed more than three times with similar results. Statistical analyses were conducted using GraphPad Prism version 8.1 (GraphPad Software) and Microsoft Excel-based application (Microsoft Office Professional Plus 2016) for the Student t-test statistical analysis. To compare two normally distributed groups, unpaired two-tailed t-tests were used. For multiple comparisons between normally distributed groups (Supplementary Tables 1 and 2), one-way ANOVA and Mann–Whitney U-test were used. Lack of an asterisk means non-significant (P > 0.05). Values in this paper are presented as mean ± s.e.m., P values are indicated as follows: *P < 0.05; **P < 0.01; ***P < 0.001.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this Article.
Data availability
The RNA-seq data have been deposited in the GEO database under accession code GSE154404. Data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
References
Evans, W. J. et al. Cachexia: a new definition. Clin. Nutr. 27, 793–799 (2008).
Argiles, J. M., Busquets, S., Stemmler, B. & Lopez-Soriano, F. J. Cancer cachexia: understanding the molecular basis. Nat. Rev. Cancer 14, 754–762 (2014).
Tisdale, M. J. Mechanisms of cancer cachexia. Physiol. Rev. 89, 381–410 (2009).
Fearon, K. C. H., Glass, D. J. & Guttridge, D. C. Cancer cachexia: mediators, signaling and metabolic pathways. Cell Metab. 16, 153–166 (2012).
Wagner, E. F. & Petruzzelli, M. Cancer metabolism: a waste of insulin interference. Nature 521, 430–431 (2015).
Sadeghi, M. et al. Cancer cachexia: diagnosis, assessment and treatment. Crit. Rev. Oncol. Hematol. 127, 91–104 (2018).
Tisdale, M. J. Cancer anorexia and cachexia. Nutrition 17, 438–442 (2001).
von Haehling, S. & Anker, S. D. Treatment of cachexia: an overview of recent developments. Int. J. Cardiol. 184, 736–742 (2015).
Tranmer, J. E. et al. Measuring the symptom experience of seriously ill cancer and noncancer hospitalized patients near the end of life with the memorial symptom assessment scale. J. Pain Symptom Manage. 25, 420–429 (2003).
Bennani-Baiti, N. & Davis, M. P. Cytokines and cancer anorexia cachexia syndrome. Am. J. Hosp. Palliat. Care 25, 407–411 (2008).
Patra, S. K. & Arora, S. Integrative role of neuropeptides and cytokines in cancer anorexia-cachexia syndrome. Clin. Chim. Acta 413, 1025–1034 (2012).
Ramos, E. J. et al. Cancer anorexia-cachexia syndrome: cytokines and neuropeptides. Curr. Opin. Clin. Nutr. Metab. Care 7, 427–434 (2004).
Mendes, M. C., Pimentel, G. D., Costa, F. O. & Carvalheira, J. B. Molecular and neuroendocrine mechanisms of cancer cachexia. J. Endocrinol. 226, R29–R43 (2015).
Droujinine, I. A. & Perrimon, N. Interorgan communication pathways in physiology: focus on Drosophila. Annu. Rev. Genet. 50, 539–570 (2016).
Gonzalez, H., Hagerling, C. & Werb, Z. Roles of the immune system in cancer: from tumor initiation to metastatic progression. Genes Dev. 32, 1267–1284 (2018).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Holmes, D. Metabolism: WAT browning—key feature of cancer-associated cachexia. Nat. Rev. Endocrinol. 10, 578 (2014).
Fearon, K., Arends, J. & Baracos, V. Understanding the mechanisms and treatment options in cancer cachexia. Nat. Rev. Clin. Oncol. 10, 90–99 (2013).
Ezeoke, C. C. & Morley, J. E. Pathophysiology of anorexia in the cancer cachexia syndrome. J. Cachexia Sarcopenia Muscle 6, 287–302 (2015).
Wang, G. et al. Metastatic cancers promote cachexia through ZIP14 upregulation in skeletal muscle. Nat. Med. 24, 770–781 (2018).
Petruzzelli, M. & Wagner, E. F. Mechanisms of metabolic dysfunction in cancer-associated cachexia. Genes Dev. 30, 489–501 (2016).
Kir, S. et al. PTH/PTHrP receptor mediates cachexia in models of kidney failure and cancer. Cell Metab. 23, 315–323 (2016).
Kir, S. et al. Tumour-derived PTH-related protein triggers adipose tissue browning and cancer cachexia. Nature 513, 100–104 (2014).
Tsai, V. W. et al. Anorexia/cachexia of chronic diseases: a role for the TGF-β family cytokine MIC-1/GDF15. J. Cachexia Sarcopenia Muscle 3, 239–243 (2012).
Hsu, J. Y. et al. Non-homeostatic body weight regulation through a brainstem-restricted receptor for GDF15. Nature 550, 255–259 (2017).
Tsai, V. W. W., Husaini, Y., Sainsbury, A., Brown, D. A. & Breit, S. N. The MIC-1/GDF15-GFRAL pathway in energy homeostasis: implications for obesity, cachexia and other associated diseases. Cell Metab. 28, 353–368 (2018).
Kwon, Y. et al. Systemic organ wasting induced by localized expression of the secreted insulin/IGF antagonist ImpL2. Dev. Cell 33, 36–46 (2015).
Figueroa-Clarevega, A. & Bilder, D. Malignant Drosophila tumors interrupt insulin signaling to induce cachexia-like wasting. Dev. Cell 33, 47–55 (2015).
Pan, D. The Hippo signaling pathway in development and cancer. Dev. Cell 19, 491–505 (2010).
Zeng, Q. & Hong, W. The emerging role of the Hippo pathway in cell contact inhibition, organ size control and cancer development in mammals. Cancer Cell 13, 188–192 (2008).
Meng, Z., Moroishi, T. & Guan, K. L. Mechanisms of Hippo pathway regulation. Genes Dev. 30, 1–17 (2016).
Harvey, K. F., Zhang, X. & Thomas, D. M. The Hippo pathway and human cancer. Nat. Rev. Cancer 13, 246–257 (2013).
Halder, G. & Camargo, F. D. The Hippo tumor suppressor network: from organ size control to stem cells and cancer. Cancer Res. 73, 6389–6392 (2013).
Misra, J. R. & Irvine, K. D. The Hippo signaling network and its biological functions. Annu. Rev. Genet. 52, 65–87 (2018).
Shell, B. C. et al. Measurement of solid food intake in Drosophila via consumption–excretion of a dye tracer. Sci. Rep. 8, 11536 (2018).
Diegelmann, S. et al. The CApillary FEeder assay measures food intake in Drosophila melanogaster. J. Vis. Exp. (2017); https://doi.org/10.3791/55024
Ja, W. W. et al. Prandiology of Drosophila and the CAFE assay. Proc. Natl Acad. Sci. USA 104, 8253–8256 (2007).
Colombani, J. et al. Drosophila Lgr3 couples organ growth with maturation and ensures developmental stability. Curr. Biol. 25, 2723–2729 (2015).
Garelli, A., Gontijo, A. M., Miguela, V., Caparros, E. & Dominguez, M. Imaginal discs secrete insulin-like peptide 8 to mediate plasticity of growth and maturation. Science 336, 579–582 (2012).
Garelli, A. et al. Dilp8 requires the neuronal relaxin receptor Lgr3 to couple growth to developmental timing. Nat. Commun. 6, 8732 (2015).
Vallejo, D. M., Juarez-Carreño, S., Bolivar, J., Morante, J. & Dominguez, M. A brain circuit that synchronizes growth and maturation revealed through Dilp8 binding to Lgr3. Science 350, aac6767 (2015).
Oh-I, S. et al. Identification of nesfatin-1 as a satiety molecule in the hypothalamus. Nature 443, 709–712 (2006).
Halls, M. L., Bathgate, R. A. D., Sutton, S. W., Dschietzig, T. B. & Summers, R. J. International union of basic and clinical pharmacology. XCV. Recent advances in the understanding of the pharmacology and biological roles of relaxin family peptide receptors 1–4, the receptors for relaxin family peptides. Pharmacol. Rev. 67, 389–440 (2015).
Kageyama, H., Takenoya, F., Shiba, K. & Shioda, S. Neuronal circuits involving ghrelin in the hypothalamus-mediated regulation of feeding. Neuropeptides 44, 133–138 (2010).
Schwartz, M. W., Woods, S. C., Porte, D., Seeley, R. J. & Baskin, D. G. Central nervous system control of food intake. Nature 404, 661–671 (2000).
Talbert, E. E. et al. Modeling human cancer-induced cachexia. Cell Rep. 28, 1612–1622 (2019).
Ronga, I., Gallucci, F., Riccardi, F. & Uomo, G. Anorexia-cachexia syndrome in pancreatic cancer: recent advances and new pharmacological approach. Adv. Med. Sci. 59, 1–6 (2014).
Boone, E., Colombani, J., Andersen, D. S. & Léopold, P. The Hippo signalling pathway coordinates organ growth and limits developmental variability by controlling dilp8 expression. Nat. Commun. 7, 13505 (2016).
Petruzzelli, M. et al. A switch from white to brown fat increases energy expenditure in cancer-associated cachexia. Cell Metab. 20, 433–447 (2014).
Katheder, N. S. et al. Microenvironmental autophagy promotes tumour growth. Nature 541, 417–420 (2017).
Stengel, A., Mori, M. & Tache, Y. The role of nesfatin-1 in the regulation of food intake and body weight: recent developments and future endeavors. Obes. Rev. 14, 859–870 (2013).
Stengel, A. & Tache, Y. Role of brain NUCB2/nesfatin-1 in the regulation of food intake. Curr. Pharm. Des. 19, 6955–6959 (2013).
Yvette, T. Editorial: New developments on NUCB2/nesfatin-1. Curr. Pharm. Des. 19, 6919–6920 (2013).
Goebel, M., Stengel, A., Lambrecht, N. W. G., Wang, L. & Taché, Y. Nesfatin-1 immunoreactivity in rat brain and spinal cord autonomic nuclei. Neurosci. Lett. 452, 241–246 (2009).
Campos, C. A. et al. Cancer-induced anorexia and malaise are mediated by CGRP neurons in the parabrachial nucleus. Nat. Neurosci. 20, 934–942 (2017).
Ishida, E. et al. Nesfatin-1 induces the phosphorylation levels of cAMP response element-binding protein for intracellular signaling in a neural cell line. PLoS ONE 7, e50918 (2012).
Brailoiu, G. C. et al. Nesfatin-1: distribution and interaction with a G protein-coupled receptor in the rat brain. Endocrinology 148, 5088–5094 (2007).
Aydin, S. Multi-functional peptide hormone NUCB2/nesfatin-1. Endocrine 44, 312–325 (2013).
Rubin, G. M. & Spradling, A. C. Genetic transformation of Drosophila with transposable element vectors. Science 218, 348–353 (1982).
Gaj, T., Gersbach, C. A. & Barbas, C. F. 3rd ZFN, TALEN and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 31, 397–405 (2013).
Lee, J. S. et al. RNA-guided genome editing in Drosophila with the purified Cas9 protein. G3 (Bethesda) 4, 1291–1295 (2014).
Colombani, J., Andersen, D. S. & Léopold, P. Secreted peptide Dilp8 coordinates Drosophila tissue growth with developmental timing. Science 336, 582–585 (2012).
Lee, K.-S., You, K.-H., Choo, J.-K., Han, Y.-M. & Yu, K. Drosophila short neuropeptide F regulates food intake and body size. J. Biol. Chem. 279, 50781–50789 (2004).
Han, S. K. et al. OASIS 2: online application for survival analysis 2 with features for the analysis of maximal lifespan and healthspan in aging research. Oncotarget 7, 56147–56152 (2016).
Conway, J. M., Ingwersen, L. A., Vinyard, B. T. & Moshfegh, A. J. Effectiveness of the US Department of Agriculture 5-step multiple-pass method in assessing food intake in obese and nonobese women. Am. J. Clin. Nutr. 77, 1171–1178 (2003).
Conway, J. M., Ingwersen, L. A. & Moshfegh, A. J. Accuracy of dietary recall using the USDA five-step multiple-pass method in men: an observational validation study. J. Am. Diet. Assoc. 104, 595–603 (2004).
Acknowledgements
We thank A.M. Gontijo, P. Leopold, M. Dominguez and P. Shen for providing reagents, D.-W. Kim and G. Wee for helping with experiments, O.-Y. Kwon for suggesting nesfatin in flies, and J.-S. Lee, K.-W. Choi, Y.-J. Kim and M. Subramanian for comments on the manuscript. Drosophila stocks were obtained from the Bloomington Stock Center (Bloomington) and Vienna Drosophila RNAi Center (VDRC). This work was supported by grants from the KRIBB Research Initiative Program, the National Research Council of Science & Technology (CRC-15-04-KIST), the National Research Foundation of Korea (2015R1A5A1009024, 2017K1A1A2013124, 2018R1A2A3075389, 2019R1A2C2004149 and 2019R1A2C2089484) and the Korean Health Technology R&D Project, Ministry of Health & Welfare, Korea (HI14C2640).
Author information
Authors and Affiliations
Contributions
E.Y., K.-S.L. and K.Y. designed the research. S.C.K. provided key reagents. E.Y., H.S., W.Y., E.J., S.H.H., D.-W.K. and T.H.R. performed experiments. E.Y., S.K., J.M.S., S.C.K., K.-S.L. and K.Y. analysed the data. E.Y., K.-S.L. and K.Y. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks Gregory Longmore, Alessandro Laviano and the other, anonymous, reviewer for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Generation of the Drosophila cancer anorexia model.
a, Drosophila cancer model was established by overexpressing a wild-type (ykiWT) and a constitutively active form of yki (ykiS168A) using GMR-Gal4. Expression of the wild-type of yki (GMR > ykiWT) caused the mild rough eye phenotype, whereas the active form (GMR > ykiSA) showed dramatic protrusion of the eye. Scale bars are 200μm. b, The 10 day-old yki overexpression flies showed decreased food intake compared with that of the control in the blue dye feeding assay. n = 3 biologically independent experiments; 1.5 days *P = 0.022, 2 days **P = 0.003. c, UAS-ykiWT and UAS-ykiSA control flies showed normal feeding. n = 3 biologically independent experiments. d, The triglyceride levels of GMR > ykiSA was analyzed with female flies. n = 3 biologically independent experiments; day 10 *P = 0.038, day 15 *P = 0.035, day 20 *P = 0.012. e, Quantification of ovary size in Fig. 1c. n = 3 biologically independent experiments; day 10 ***P = 0.0006, day 15 **P = 0.0014, day 20 ***P < 0.001. f, The thoracic muscle of 15-day-old GMR > ykiSA showed abnormal mitochondrial structure and irregular packing of mitochondria between muscle fibers. n = 3 biologically independent experiments. Scale bar is 10μm. Data are presented as the mean ± s.e.m. Statistical significance was determined with two-tailed Student’s t-test; *P < 0.05, **P < 0.01, ***P < 0.001. Statistical source data.
Extended Data Fig. 2 RNA-seq analysis, anorexia in sev > Rasv12 flies, Lgr3Δ50 generation, Lgr3 expression in adult tissues, and eye phenotypes in GMR > ykiSA and the mutants of Dilp8 and Lgr3.
a, RNA-seq analysis showed the increased levels of three tumor-secreted factors (Dilp8, ImpL2, and Upd2) in GMR > ykiSA flies. b, sev > Rasv12 flies showed anorexia and this phenotype was effectively rescued by Dilp8-RNAi. n = 3 biologically independent experiments; sev > Rasv12 *P = 0.012, sev > Rasv12 Dilp8-RNAi *P = 0.013, sev > Rasv12 ImpL2-RNAi *P = 0.019. c, Dilp8 antibody test was performed by western blot and showed almost no expression in the Dilp8M100727 mutant. d, Dilp8 protein was increased in GMR > ykiSA fly heads and reduced in GMR > ykiSA, Dilp8-RNAi fly heads by western blot. e, The quantification of d using ImageJ program. n = 3 biologically independent experiments; GMR > ykiSA *P = 0.047, GMR > ykiSA Dilp8-RNAi *P = 0.045. f, The Lgr3Δ50 mutant was obtained using the CRISPR/Cas9-mediated mutagenesis. It consists of 50 base pair deletion in the intron region upstream of target sequences. g, Lgr3 mRNA expression in the Lgr3Δ50 mutant showed that the Lgr3Δ50 mutant is a strong hypomorphic allele. n = 3 biologically independent experiments, **P = 0.007. h, GMR > ykiSA,Dilp8MI00727 and GMR > ykiSA, Lgr3Δ50 flies respectively blunted or reversed the cancer anorexia phenotype observed in GMR > ykiSA flies. n = 4 biologically independent experiments; GMR > ykiSA *P = 0.010, GMR > ykiSADilp8M100727 *P = 0.012, GMR > ykiSALgr3Δ50 *P = 0.027. i, Dilp8 overexpression flies (GMR > Dilp8) showed reduced food intake. n = 6 biologically independent experiments, *P = 0.034. j, In GMR > ykiSA and GMR-Gal4 control flies, the Lgr3 mRNA expression level in the head is very high relative to other tissues. n = 3 biologically independent experiments; head *P = 0.017; thorax **P = 0.006, gut *P = 0.035. k, The eye phenotype in GMR > ykiSA is similar with those of GMR > ykiSA, Dilp8M100727 and GMR > ykiSA, Lgr3Δ50. Scale bar is 200μm. l, Lgr3-Gal4 expression marked by GFP. Lgr3 expression was not detectable in sNPF neurons. Scale bar is 100μm. m-n, sNPF (m) (*P = 0.018) and NPF (n) (*P = 0.048) mRNA levels were reduced by RNAi lines with Lgr3-Gal4. n = 3 biologically independent experiments. Data are presented as the mean ± s.e.m. Statistical significance was determined with two-tailed Student’s t-test; *P < 0.05, **P < 0.01. Statistical source data and unprocessed western blots.
Extended Data Fig. 3 Sequence homology of NUCB1 and NUCB2 in Drosophila and mammals.
The very similar signal peptide and cleavage site among mammalian NUCB2 and Drosophila NUCB1 suggest that the evolutionary conserved Nesfatin1 might be produced.
Extended Data Fig. 4 Intensity quantification of NUCB1 and Lgr3 antibodies immunostaining.
a, NUCB1 antibody specificity was tested by the NUCB1 inhibition using heat-shock Gal4 (Hs > NUCB1-RNAi). b, Staining intensity of neurons was measured with LSM 5 software program by drawing a dotted line as shown. c-e, The staining intensity of neuronal cells in Fig. 4 was quantified using LSM 5 software program and illustrated as intensity graphs. Control (n = 5), GMR > ykiSA (n = 10), GMR > ykiSA, Lgr3Δ50 (n = 5). f-g, A subset of sNPF (f) and NPF (g) expressing neurons in the cancer condition (GMR > ykiSA) were overlapped with NUCB1 expressing neurons. Scale bars are 100 μm. Statistical source data and unprocessed western blots.
Extended Data Fig. 5 NUCB1 protein expression in the cancer condition, ex vivo analysis, and Lgr3 expression by the Dilp8 treatment.
a, In western blot of adult heads with the NUCB1 antibody, NUCB1 protein level is increased in GMR > ykiSA, reduced in the Lgr3Δ50 mutant, and restored to the control level in GMR > ykiSA, Lgr3Δ50. b, The western blot bands of (a) were quantified using ImageJ program. n = 3 biologically independent experiments; GMR > ykiSA **P = 0.003, Lgr3Δ50 *P = 0.038, GMR > ykiSALgr3Δ50 ** P = 0.002. c, The schematic presentation of ex vivo culture assay in which the synthetic Dilp8 peptide was treated to the dissected adult brains in the culture media. d, Lgr3 mRNA level was increased after Dilp8 peptide treatment in the ex vivo culture. n = 3 biologically independent experiments; 0.5 hr *P = 0.020, 1 hr *P = 0.021. Data are presented as the mean ± s.e.m. Statistical significance was determined with two-tailed Student’s t-test; *P < 0.05, **P < 0.01. Statistical source data and unprocessed western blots.
Extended Data Fig. 6 Conservation of INSL3/Lgr8 in mammals, and cachexia-anorexia phenotype of C26 or LLC implanted mice.
a. The phylogenetic comparative analysis of Drosophila Lgr3 and mammalian Lgr family shows that Lgr7/8 is a homologue of Lgr3. b, The phylogenetic comparative analysis of fly Dilp8 and mammalian INSL3 shows that they are classified as close peptides. c, Lgr8 protein is strongly expressed in the hypothalamic region and hippocampus of the mouse brain. d, In the RNA FISH, Lgr8 mRNA is strongly expressed in the hypothalamic region and hippocampus of the mouse brain. e, Lgr8 is expressed in the mouse hypothalamic N3 cells, but not in N39 cells. f-g, Body weight was changed in C26 tumor implanted mice (f) not in LLC tumor implanted mice (g). n = 7 mice for both groups. f, 2w + 2day *P = 0.027, 2w + 4day ***P = 0.0004, 2w + 6day **P = 0.0034. h, Tumor volume and weight changes were measured in C26 and LLC implanted mice. n = 6 mice for both groups. i-j, Muscle and fat weight of C26 tumor implanted mice at day 12 (i) and day 21 (j). Tibialis anterior (TA), gastrocnemius (GCM), inguinal white adipose tissue (iWAT), and epididymal white adipose tissue (eWAT). Control n = 3 mice, C26 day 12 n = 5 mice, C26 day 21 n = 4 mice. i, iWAT *P = 0.024, eWAT *P = 0.094. j, GCM *P = 0.038, iWAT **P = 0.001, eWAT *P = 0.013. k-l, Nucb2 (k) and Npy (l) mRNA levels were not changed in LLC implanted mice. n = 5 mice for both groups. m, INSL3 i.p. injected wild-type mice didn’t show reduction of food intake. n = 7 mice for both groups. n-o, Nucb2 (n) and Npy (o) mRNA levels were not changed in INSL3 i.p. injected wild-type mice hypothalamus. n = 3 mice for both groups. Data are presented as the mean ± s.e.m. Statistical significance was determined with two-tailed Student’s t-test; *P < 0.05, **P < 0.01, ***P < 0.001. Statistical source data and unprocessed western blots.
Extended Data Fig. 7 Anorexia-associated body mass changes of human pancreatic cancer implanted mice.
a, Tumor volumes were increased in Panc1 and Capan1 implanted mice. n = 4 mice for three groups; 3 week *P = 0.049, 4 week *P = 0.049, 5 week *P = 0.033. b, Body mass were reduced in Panc1 and Capan1 implanted mice compared to the control mice. n = 4 mice for each group; 3 week Panc1 **P = 0.0014 and Capan1 *P = 0.049, 5 week Panc1 ***P < 0.0001 and Capan1 ***P < 0.0001. 6 week Panc1 ***P < 0.0001 and Capan1 ***P < 0.0001. c, Relaxin-INSL family mRNA expression levels in Capan1 cells. n = 3 biologically independent experiments; Rxn vs INSL3 **P = 0.003. Data are presented as the mean ± s.e.m. Statistical significance was determined with two-tailed Student’s t-test; *P < 0.05. Statistical source data.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6.
Supplementary Tables 1–3
Supplementary Table 1. Feeding regulatory neuropeptides in Drosophila were identified in the RNA sequencing analysis of GMR>ykiSA. Supplementary Table 2. Clinicopathological characteristics among patients with pancreatic cancer and other diseases. Supplementary Table 3. Clinicopathological characteristics of patients enrolled in food intake analysis.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 2
Unprocessed blots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 5
Unprocessed blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed blots.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Yeom, E., Shin, H., Yoo, W. et al. Tumour-derived Dilp8/INSL3 induces cancer anorexia by regulating feeding neuropeptides via Lgr3/8 in the brain. Nat Cell Biol 23, 172–183 (2021). https://doi.org/10.1038/s41556-020-00628-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-00628-z
This article is cited by
-
Molecular basis of GDF15 induction and suppression by drugs in cardiomyocytes and cancer cells toward precision medicine
Scientific Reports (2023)
-
A novel antidiuretic hormone governs tumour-induced renal dysfunction
Nature (2023)
-
Understanding the molecular basis of anorexia and tissue wasting in cancer cachexia
Experimental & Molecular Medicine (2022)
-
Cholecystokinin/sulfakinin peptide signaling: conserved roles at the intersection between feeding, mating and aggression
Cellular and Molecular Life Sciences (2022)
-
Tumour–host interactions through the lens of Drosophila
Nature Reviews Cancer (2021)