Abstract
Detection of endogenous signals and precise control of genetic circuits in the natural context are essential to understand biological processes. However, the tools to process endogenous information are limited. Here we developed a generalizable endogenous transcription-gated switch that releases single-guide RNAs in the presence of an endogenous promoter. When the endogenous transcription-gated switch is coupled with the highly sensitive CRISPR-activator-associated reporter we developed, we can reliably detect the activity of endogenous genes, including genes with very low expression (<0.001 relative to Gapdh; quantitative-PCR analysis). Notably, we could also monitor the transcriptional activity of typically long non-coding RNAs expressed at low levels in living cells using this approach. Together, our method provides a powerful platform to sense the activity of endogenous genetic elements underlying cellular functions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The previously published RNA sequencing data that were re-analysed here are available under the accession code GSM2573084. All of the raw data associated with the figures are listed in the Source data (statistical source data). All of the raw images for the western blots can be found in the Source data (unprocessed western blots and gels). The sequences of all vectors are provided in Supplementary Fig. 1. All other materials and data are available on request. Source data are provided with this paper.
References
Green, A. A., Silver, P. A., Collins, J. J. & Yin, P. Toehold switches: de-novo-designed regulators of gene expression. Cell 159, 925–939 (2014).
Liu, Y. et al. Directing cellular information flow via CRISPR signal conductors. Nat. Methods 13, 938–944 (2016).
Green, A. A. et al. Complex cellular logic computation using ribocomputing devices. Nature 548, 117–121 (2017).
Hirosawa, M. et al. Cell-type-specific genome editing with a microRNA-responsive CRISPR–Cas9 switch. Nucleic Acids Res. 45, e118 (2017).
Siu, K. H. & Chen, W. Riboregulated toehold-gated gRNA for programmable CRISPR–Cas9 function. Nat. Chem. Biol. 15, 217–220 (2019).
Wang, X. W. et al. A microRNA-inducible CRISPR–Cas9 platform serves as a microRNA sensor and cell-type-specific genome regulation tool. Nat. Cell Biol. 21, 522–530 (2019).
Miki, K. et al. Efficient detection and purification of cell populations using synthetic microRNA switches. Cell Stem Cell 16, 699–711 (2015).
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR–Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Yao, X. et al. Homology-mediated end joining-based targeted integration using CRISPR/Cas9. Cell Res. 27, 801–814 (2017).
Mikuni, T., Nishiyama, J., Sun, Y., Kamasawa, N. & Yasuda, R. High-throughput, high-resolution mapping of protein localization in mammalian brain by in vivo genome editing. Cell 165, 1803–1817 (2016).
Nishiyama, J., Mikuni, T. & Yasuda, R. Virus-mediated genome editing via homology-directed repair in mitotic and postmitotic cells in mammalian brain. Neuron 96, 755–768 (2017).
Guttman, M. & Rinn, J. L. Modular regulatory principles of large non-coding RNAs. Nature 482, 339–346 (2012).
Kretz, M. et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 493, 231–235 (2013).
Sauvageau, M. et al. Multiple knockout mouse models reveal lincRNAs are required for life and brain development. eLife 2, e01749 (2013).
Bester, A. C. et al. An integrated genome-wide CRISPRa approach to functionalize lncRNAs in drug resistance. Cell 173, 649–664 (2018).
Cabili, M. N. et al. Localization and abundance analysis of human lncRNAs at single-cell and single-molecule resolution. Genome Biol. 16, 20 (2015).
Chen, L. et al. Tissue expression difference between mRNAs and lncRNAs. Int. J. Mol. Sci. https://doi.org/10.3390/ijms19113416 (2018).
Azlan, A., Obeidat, S. M., Yunus, M. A. & Azzam, G. Systematic identification and characterization of Aedes aegypti long noncoding RNAs (lncRNAs). Sci. Rep. 9, 12147 (2019).
Zhou, H. et al. In vivo simultaneous transcriptional activation of multiple genes in the brain using CRISPR–dCas9-activator transgenic mice. Nat. Neurosci. 21, 440–446 (2018).
Wang, J. et al. Generation of cell-type-specific gene mutations by expressing the sgRNA of the CRISPR system from the RNA polymerase II promoters. Protein Cell 6, 689–692 (2015).
Black, D. L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003).
Matlin, A. J., Clark, F. & Smith, C. W. Understanding alternative splicing: towards a cellular code. Nat. Rev. Mol. Cell Biol. 6, 386–398 (2005).
Xie, K., Minkenberg, B. & Yang, Y. Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc. Natl Acad. Sci. USA 112, 3570–3575 (2015).
Zhang, D. et al. Perfectly matched 20-nucleotide guide RNA sequences enable robust genome editing using high-fidelity SpCas9 nucleases. Genome Biol. 18, 191 (2017).
Nissim, L., Perli, S. D., Fridkin, A., Perez-Pinera, P. & Lu, T. K. Multiplexed and programmable regulation of gene networks with an integrated RNA and CRISPR/Cas toolkit in human cells. Mol. Cell 54, 698–710 (2014).
Ying, Q. L., Stavridis, M., Griffiths, D., Li, M. & Smith, A. Conversion of embryonic stem cells into neuroectodermal precursors in adherent monoculture. Nat. Biotechnol. 21, 183–186 (2003).
Zhang, Z. H., Lu, Y. Y. & Yue, J. Two pore channel 2 differentially modulates neural differentiation of mouse embryonic stem cells. PLoS ONE 8, e66077 (2013).
Wongpaiboonwattana, W. & Stavridis, M. P. Neural differentiation of mouse embryonic stem cells in serum-free monolayer culture. J. Vis. Exp. https://doi.org/10.3791/52823 (2015).
Nair, G., Abranches, E., Guedes, A. M., Henrique, D. & Raj, A. Heterogeneous lineage marker expression in naive embryonic stem cells is mostly due to spontaneous differentiation. Sci. Rep. 5, 13339 (2015).
Neri, F. et al. TET1 is controlled by pluripotency-associated factors in ESCs and downmodulated by PRC2 in differentiated cells and tissues. Nucleic Acids Res. 43, 6814–6826 (2015).
Guo, F. et al. Single-cell multi-omics sequencing of mouse early embryos and embryonic stem cells. Cell Res. 27, 967–988 (2017).
Ramos, A. D. et al. Integration of genome-wide approaches identifies lncRNAs of adult neural stem cells and their progeny in vivo. Cell Stem Cell 12, 616–628 (2013).
Liu, S. J. et al. Single-cell analysis of long non-coding RNAs in the developing human neocortex. Genome Biol. 17, 67 (2016).
Salviano-Silva, A., Lobo-Alves, S. C., Almeida, R. C., Malheiros, D. & Petzl-Erler, M. L. Besides pathology: long non-coding RNA in cell and tissue homeostasis. Noncoding RNA https://doi.org/10.3390/ncrna4010003 (2018).
Sun, Z. et al. The long noncoding RNA Lncenc1 maintains naive states of mouse ESCs by promoting the glycolysis pathway. Stem Cell Rep. 11, 741–755 (2018).
Liu, Y. et al. CRISPR activation screens systematically identify factors that drive neuronal fate and reprogramming. Cell Stem Cell 23, 758–771 (2018).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Gao, N., Hu, J., Zhou, H. & Yang, H. A protocol to generate SPH-OminiCMV-Ents mESCs. Protoc. Exch. https://doi.org/10.21203/rs.3.pex-1273/v1 (2020).
Acknowledgements
We thank L. Quan, H. Wu and S. Qian from the FACS facility in ION as well as Y. Wang, Y. Zhang, X. Chen, D. Xiang and Q. Hu from the Optical Imaging facility. We thank N. Zhong and Q. Wang for their technical assistance. This work was supported by the Basic Frontier Scientific Research Program of the Chinese Academy of Sciences From 0 to 1 original innovation project (grant no. ZDBS-LY-SM001), R&D Program of China (grant nos 2017YFC1001300 and 2018YFC2000100), CAS Strategic Priority Research Program (grant no. XDB32060000), National Natural Science Foundation of China (grant nos 31871502, 31925016, 91957122 and 31901047), Shanghai Municipal Science and Technology Major Project (grant no. 2018SHZDZX05), Shanghai City Committee of Science and Technology Project (grant nos 18411953700, 18JC1410100 and 19XD1424400) and International Partnership Program of Chinese Academy of Sciences (grant no. 153D31KYSB20170059).
Author information
Authors and Affiliations
Contributions
N.G. designed experiments, constructed vectors, performed transfections, generated cell lines and analysed data. J.Hu performed RNA FISH, immunofluorescence staining and assisted with the vector construction. B.H., J.Huang, Yu Wei, J.P., Yinghui Wei, X.S. and L.S. assisted with the generation of cell lines and analysis. Z.J. constructed vectors and performed qPCR analysis and cell-line genotyping. X.H., Q.X. and H.L. performed the transient transfections and analysis in the cell cultures. N.G. and X.F. performed the western blots. Y.S. and Y.Z. designed synthetic sgRNA sequences and analysed the public RNA sequencing data. C.Z. assisted with the vector construction. H.Z. and H.Y. conceived the project, designed experiments, supervised the project and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks Rory Johnson, Ophir Shalem and the other, anonymous, reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Mean mCherry intensity induced by different sgRNAs and Optimization of the miniCMV promoter.
a, The fluorescence intensity of mCherry induced by different sgRNAs (n = 2 repeats). Note that the intensity was quantified by FACS, 24 hours after transient transfection of SPH, miniCMV and sgRNA in 293T cells. b, Representative images showing mCherry expression in N2a cells induced by different mini-promoters, 24 hours after transient transfection, each experiment was independently repeated 3 times with similar results. Scale bar, 200 μm. c, Mean fluorescence intensity of mCherry, 48 hours after transient transfection, n = 3 repeats per group (miniCMV v.s. Mini-TK, p < 0.0001; miniCMV v.s. Luc2CP, p < 0.0001; miniCMV v.s. TRE3G, p < 0.0001; unpaired two-sided Student’s t test). d, Mean mCherry intensity induced by different TS intervals, 48 hours after transient transfection, n = 3 repeats per group. e, The influence of sgRNA copy number on mCherry expression for different TS intervals, number above the bar indicates the number of repeats per group. f, Mean fluorescence intensity of mCherry induced by SPH-OminiCMV and different promoters, 48 hours after transient transfection. n = 3 repeats per group. g, Representative images showing that SPH-OminiCMV induces higher levels of mCherry than commonly used strong promoters. The tagBFP was co-transfected to control the transfection efficiency. Experiments were independently repeated 2 times per group with similar results. Scale bar: 50 μm. N2a cells were transiently transfected with plasmids for all experiments. All values are presented as mean ± s.e.m.; unpaired two-sided Student’s t test; *p < 0.05, **p < 0.01, ***p < 0.001. Statistical source data are provided in Source Data Extended Data Fig. 1.
Extended Data Fig. 2 SPH-OminiCMV induced higher levels of gene expression than SPH-mediated endogenous activation and CMV-mediated overexpression.
a, Schematic showing SPH-mediated endogenous gene activation, CMV- and SPH-OminiCMV-mediated exogenous gene expression. b, Relative mRNA expression levels of different genes, n = 2 repeats per group. Statistical source data are provided in Source Data Extended Data Fig. 2.
Extended Data Fig. 3 Specificity of SPH-OminiCMV and generation of SPH-OminiCMV transgenic mESCs.
a, Tolerance of improved reporter system to mismatched sgRNAs by transient transfection of SPH, OminiCMV and sgRNA in N2a cells. X-axis: No. of sgRNA TS; Y-axis: No. of sgRNA. Scale bar indicates the mean fluorescence intensity (red, high; white, low). The sgRNA information is provided in Supplementary Table 1, n = 2 repeats per group. b, Schematic illustration of the transgenes. c, Genotyping of SPH-OminiCMV transgenic mESCs via PCR, ‘+’ indicates the positive colony and ‘-’ indicates the negative colony, images are representative of several ‘+’ colonies. d, Images showing transient expression of sgRNA induced mCherry expression in the SPH-OminiCMV positive colony, images are representative of 3 experiments. Scale bar: 200 μm. e, Relative mCherry expression quantified by qPCR, n = 3 repeats per group. All values are presented as mean ± s.e.m.. Statistical source data and unprocessed gels are provided in Source Data Extended Data Fig. 3.
Extended Data Fig. 4 Targeted insertion of tRNA-sgRNA-tRNA into the 3’UTR does not affect the normal protein production of target genes.
a, Schematic showing that tRNA-sgRNA-tRNA was inserted into the 3’UTR or intron of Actb, and mean fluorescence intensity of mCherry. Number above the bar indicates the number of repeats per group (SPH-OminiCMV-Ents-Intron 1 v.s. SPH-OminiCMV-Ents-3’UTR, p = 0.5689; unpaired two-sided Student’s t test). b, Western blots, n = 4 repeats per group. c, Quantification of Western blots data showing that insertion of tRNA-sgRNA-tRNA into the 3’UTR of Actb loci did not influence the production of Actb (SPH-OminiCMV-Ents v.s. SPH-OminiCMV, p = 0.3456; SPH-OminiCMV-Ents v.s. P2A-mCherry, p = 0.6301; unpaired two-sided Student’s t test). Number above the bar indicates the number of repeats per group. d, e, Insertion of the tRNA-sgRNA-tRNA targeting LacZ in SPH-OminiCMV mESCs does not induce mCherry expression, number above the bar indicates the number of repeats per group. All values are presented as mean ± s.e.m.; unpaired two-sided Student’s t test; *p < 0.05, **p < 0.01, ***p < 0.001. Statistical source data and unprocessed western blots are provided in Source Data Extended Data Fig. 4.
Extended Data Fig. 5 SPH-OminiCMV-Ents enables the visualization of low-abundance genes during cell differentiation, and insertion of an sgRNA array into the non-expressed gene does not induce mCherry expression.
a, b, Representative images showing mCherry expression during differentiation and quantification of Esrrb mRNA levels using qPCR in SPH-OminiCMV-Ents-Esrrb mESCs during differentiation. Scale bar, 50 μm; n = 4 repeats per group. c, d, Representative images showing mCherry expression and quantification of Sox2 mRNA levels using qPCR in SPH-OminiCMV-Ents-Sox2 mESCs during differentiation. Scale bar, 50 μm; n = 4 repeats per group. e, f, Representative images showing mCherry expression and quantification of Tet1 mRNA levels using qPCR in SPH-OminiCMV-Ents-Tet1 mESCs during differentiation. Scale bar: 50 μm, n = 4 repeats per group. g, Schematic showing insertion of one sgRNA or an sgRNA array into the 3’UTR of Sema3a locus. h, Quantification of Sema3a expression by qPCR in ESCs, n = 3 repeats. i, Images showing mCherry expression, images are representative of 4 experiments. Scale bar, 50 μm. j, Mean mCherry fluorescence intensity. Number above the bar indicates the number of colonies per group (SPH-OminiCMV-Ents-one sgRNA v.s. SPH-OminiCMV, p = 0.5964; SPH-OminiCMV-Ents-sgRNA array v.s. SPH-OminiCMV, p = 0.0761; unpaired two-sided Student’s t test). All values are presented as mean ± s.e.m.; unpaired two-sided Student’s t test; *p < 0.05, **p < 0.01, ***p < 0.001. Statistical source data are provided in Source Data Extended Data Fig. 5.
Extended Data Fig. 6 Homogeneous expression of mCherry by driving dCas9 and activators under a single promoter.
a, Schematic of the vector. Note that dCas9 and P65-HSF1 were expressed by two CAG promoters respectively. b, Histogram of FACS analysis of a SPH-OminiCMV-Ents-Actb colony. c, Different expression levels of activators between mCherry-high and mCherry-low cells from the same SPH-OminiCMV-Ents-Actb colony, n = 3 repeats (High-5% v.s. Low-5%: dCas9, p = 0.0028; p65-HSF1, p = 0.9161; Actb, p = 0.0743; sgRNA, p = 0.0030; unpaired two-sided Student’s t test). d, Schematic showing SPH (single CAG), note that the expression of dCas9 and p65-HSF1 was driven by a single promoter. e, Histogram of FACS analysis of a SPH (single CAG) -OminiCMV-Ents-Actb colony. All values are presented as mean ± s.e.m.; unpaired two-sided Student’s t test; *p < 0.05, **p < 0.01, ***p < 0.001. Statistical source data are provided in Source Data Extended Data Fig. 6.
Extended Data Fig. 7 FACS analysis of mCherry expression.
a, Analysis of the flow cytometry data showing how R1 was gated (side scatter: SSC; forward scatter: FSC). b, Representative histogram of FACS analysis of SPH (single CAG)-OminiCMV-Ents-one sgRNA, SPH (single CAG)-OminiCMV-Ents-sgRNA array and P2A-mCherry cells for different genes. Each experiment was independently repeated several times with similar results. For the number of repeats, see Fig. 5d. c, Representative histogram of SPH (single CAG)-OminiCMV-Ents-one sgRNA and SPH (single CAG)-OminiCMV-Ents-sgRNA array cells for different lncRNAs. Each experiment was independently repeated several times with similar results. For the number of repeats, see Fig. 5f.
Extended Data Fig. 8 The side-by-side comparison of different strategies and downregulation of mCherry and Nanog at the protein level during differentiation.
a, Analysis of the flow cytometry data showing how R1 was gated (side scatter: SSC; forward scatter: FSC). b, Representative histogram of FACS analysis of SPH (single CAG)-OminiCMV-Ents-one sgRNA, SPH (single CAG)-OminiCMV-Ents-sgRNA array and P2A-mCherry cells for different genes. Each experiment was independently repeated several times with similar results. For the number of repeats, see Fig. 5d. c, Representative histogram of SPH (single CAG)-OminiCMV-Ents-one sgRNA and SPH (single CAG)-OminiCMV-Ents-sgRNA array cells for different lncRNAs. Each experiment was independently repeated several times with similar results. For the number of repeats, see Fig. 5f.
Extended Data Fig. 9 SPH (single CAG)-OminiCMV-Ents-sgRNA array induces the highest expression of mCherry.
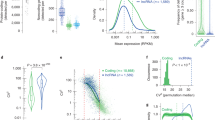
a, Comparison of the fluorescence intensity of SPH-OminiCMV-Ents-one sgRNA, SPH-OminiCMV-Ents-sgRNA array, SPH (single CAG)-OminiCMV-Ents-one sgRNA and SPH (single CAG)-OminiCMV-Ents-sgRNA array systems. Number in the bar indicates the number of colonies per group. Unpaired two-sided Student’s t test. Note that SPH (single CAG)-OminiCMV-Ents-one sgRNA and SPH (single CAG)-OminiCMV-Ents-sgRNA array data are also shown in Fig. 5d; SPH-OminiCMV-Ents-one sgRNA and SPH-OminiCMV-Ents-sgRNA array data are also shown in Figs. 2c, 4e. b, LncRNA expression matrix of embryonic stem cells. Note that the data was downloaded from the public GEO database (GSM2573084) and lncRNAs were ordered according to their expression levels in a decreasing order. LncRNAs marked in red and black indicate those that explored in Fig. 5f. In total, 14432 lncRNAs were detected with a FPKM value higher than 0 and 9640 lncRNAs have a FPKM value higher than that of Pvt1. All values are presented as mean ± s.e.m.; unpaired two-sided Student’s t test; *p < 0.05, **p < 0.01, ***p < 0.001. Statistical source data and p values for a are provided in Source Data Extended Data Fig. 9.
Supplementary information
Supplementary Information
Supplementary sequences.
Supplementary Tables
Supplementary Table 1: sgRNA sequences. Supplementary Table 2: sgRNA sequences for endogenous gene activation. Supplementary Table 3: The sgRNA sequences for inserting the sgRNA precursor. Supplementary Table 4: Primers for identifying the sgRNA insertion. Supplementary Table 5: Relative expression levels. Supplementary Table 6: qPCR primers.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed gels.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 4
Unprocessed western blots.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
About this article
Cite this article
Gao, N., Hu, J., He, B. et al. Endogenous promoter-driven sgRNA for monitoring the expression of low-abundance transcripts and lncRNAs. Nat Cell Biol 23, 99–108 (2021). https://doi.org/10.1038/s41556-020-00610-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-00610-9
This article is cited by
-
Optimal tagging strategies for illuminating expression profiles of genes with different abundance in zebrafish
Communications Biology (2023)
-
Precise tumor immune rewiring via synthetic CRISPRa circuits gated by concurrent gain/loss of transcription factors
Nature Communications (2022)
-
Intron-encoded cistronic transcripts for minimally invasive monitoring of coding and non-coding RNAs
Nature Cell Biology (2022)
-
Long non-coding RNA PAARH promotes hepatocellular carcinoma progression and angiogenesis via upregulating HOTTIP and activating HIF-1α/VEGF signaling
Cell Death & Disease (2022)
-
Monitoring the promoter activity of long noncoding RNAs and stem cell differentiation through knock-in of sgRNA flanked by tRNA in an intron
Cell Discovery (2021)