Abstract
The contribution of ribosome heterogeneity and ribosome-associated proteins to the molecular control of proteomes in health and disease remains unclear. Here, we demonstrate that survival motor neuron (SMN) protein—the loss of which causes the neuromuscular disease spinal muscular atrophy (SMA)—binds to ribosomes and that this interaction is tissue-dependent. SMN-primed ribosomes are preferentially positioned within the first five codons of a set of mRNAs that are enriched for translational enhancer sequences in the 5′ untranslated region (UTR) and rare codons at the beginning of their coding sequence. These SMN-specific mRNAs are associated with neurogenesis, lipid metabolism, ubiquitination, chromatin regulation and translation. Loss of SMN induces ribosome depletion, especially at the beginning of the coding sequence of SMN-specific mRNAs, leading to impairment of proteins that are involved in motor neuron function and stability, including acetylcholinesterase. Thus, SMN plays a crucial role in the regulation of ribosome fluxes along mRNAs encoding proteins that are relevant to SMA pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Ribosome profiling data generated by the current study have been deposited in the Gene Expression Omnibus (GEO) under the accession code GSE154106. Classical and active ribosome profiling data of healthy mouse brains that were reanalysed in the current study were retrieved from the GEO: GSE102318 (Ribo-seq) and GSE102354 (Active Ribo-seq). All other data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
References
Buttgereit, F. & Brand, M. D. A hierarchy of ATP-consuming processes in mammalian cells. Biochem. J. 312, 163–167 (1995).
Russell, J. B. & Cook, G. M. Energetics of bacterial growth: balance of anabolic and catabolic reactions. Microbiol. Rev. 59, 48–62 (1995).
Vogel, C. et al. Sequence signatures and mRNA concentration can explain two-thirds of protein abundance variation in a human cell line. Mol. Syst. Biol. 6, 400 (2010).
Schwanhüusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Tebaldi, T. et al. Widespread uncoupling between transcriptome and translatome variations after a stimulus in mammalian cells. BMC Genom. 13, 220 (2012).
Bhat, M. et al. Targeting the translation machinery in cancer. Nat. Rev. Drug Discov. https://doi.org/10.1038/nrd4505 (2015).
Topisirovic, I. & Sonenberg, N. Translation and cancer. Biochim. Biophys. Acta https://doi.org/10.1016/j.bbagrm.2015.05.004 (2015).
Darnell, J. C. et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 146, 247–261 (2011).
Khalil, B., Morderer, D., Price, P. L., Liu, F. & Rossoll, W. mRNP assembly, axonal transport, and local translation in neurodegenerative diseases. Brain Res. https://doi.org/10.1016/j.brainres.2018.02.018 (2018).
Costa, C. J. & Willis, D. E. To the end of the line: axonal mRNA transport and local translation in health and neurodegenerative disease. Dev. Neurobiol. https://doi.org/10.1002/dneu.22555 (2018).
Bernabò, P. et al. In vivo translatome profiling in spinal muscular atrophy reveals a role for SMN protein in ribosome biology. Cell Rep. 21, 953 (2017).
Simsek, D. & Barna, M. An emerging role for the ribosome as a nexus for post-translational modifications. Curr. Opin. Cell Biol. https://doi.org/10.1016/j.ceb.2017.02.010 (2017).
Mauro, V. P. & Edelman, G. M. The ribosome filter hypothesis. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.192442499 (2002).
Xue, S. & Barna, M. Specialized ribosomes: a new frontier in gene regulation and organismal biology. Nat. Rev. Mol. Cell Biol. 13, 355–369 (2012).
Shi, Z. et al. Heterogeneous ribosomes preferentially translate distinct subpools of mRNAs genome-wide. Mol. Cell https://doi.org/10.1016/j.molcel.2017.05.021 (2017).
Slavov, N., Semrau, S., Airoldi, E., Budnik, B. & van Oudenaarden, A. Differential stoichiometry among core ribosomal proteins. Cell Rep. https://doi.org/10.1016/j.celrep.2015.09.056 (2015).
Kurylo, C. M. et al. Endogenous rRNA sequence variation can regulate stress response gene expression and phenotype. Cell Rep. https://doi.org/10.1016/j.celrep.2018.08.093 (2018).
Parks, M. M. et al. Variant ribosomal RNA alleles are conserved and exhibit tissue-specific expression. Sci. Adv. https://doi.org/10.1126/sciadv.aao0665 (2018).
Fujii, K., Susanto, T. T., Saurabh, S. & Barna, M. Decoding the function of expansion segments in ribosomes. Mol. Cell https://doi.org/10.1016/j.molcel.2018.11.023 (2018).
Kondrashov, N. et al. Ribosome-mediated specificity in Hox mRNA translation and vertebrate tissue patterning. Cell 145, 383–397 (2011).
Lefebvre, S. et al. Identification and characterization of a spinal muscular atrophy-determining gene. Cell 80, 155–165 (1995).
Groen, E. J. N., Talbot, K. & Gillingwater, T. H. Advances in therapy for spinal muscular atrophy: promises and challenges. Nat. Rev. Neurol. https://doi.org/10.1038/nrneurol.2018.4 (2018).
Zhang, Z. et al. SMN Deficiency causes tissue-specific perturbations in the repertoire of snRNAs and widespread defects in splicing. Cell https://doi.org/10.1016/j.cell.2008.03.031 (2008).
Lotti, F. et al. An SMN-dependent U12 splicing event essential for motor circuit function. Cell https://doi.org/10.1016/j.cell.2012.09.012 (2012).
Bäumer, D. et al. Alternative splicing events are a late feature of pathology in a mouse model of spinal muscular atrophy. PLoS Genet. https://doi.org/10.1371/journal.pgen.1000773 (2009).
Simon, C. M. et al. Converging mechanisms of p53 activation drive motor neuron degeneration in spinal muscular atrophy. Cell Rep. https://doi.org/10.1016/j.celrep.2017.12.003 (2017).
Price, P. L., Morderer, D. & Rossoll, W. Assembly defects in spinal muscular atrophy. Adv. Neurobiol. 20, 143–171 (2018).
Sanchez, G. et al. A novel function for the survival motoneuron protein as a translational regulator. Hum. Mol. Genet. 22, 668–684 (2013).
Béchade, C. et al. Subcellular distribution of survival motor neuron (SMN) protein: possible involvement in nucleocytoplasmic and dendritic transport. Eur. J. Neurosci. 11, 293–304 (1999).
Fallini, C., Donlin-Asp, P. G., Rouanet, J. P., Bassell, G. J. & Rossoll, W. Deficiency of the survival of motor neuron protein impairs mRNA localization and local translation in the growth cone of motor neurons. J. Neurosci. 36, 3811–3820 (2016).
Francisco-Velilla, R., Fernandez-Chamorro, J., Ramajo, J. & Martinez-Salas, E. The RNA-binding protein Gemin5 binds directly to the ribosome and regulates global translation. Nucleic Acids Res. https://doi.org/10.1093/nar/gkw702 (2016).
Burghes, A. H. M. & Beattie, C. E. Spinal muscular atrophy: why do low levels of survival motor neuron protein make motor neurons sick? Nat. Rev. Neurosci. https://doi.org/10.1038/nrn2670 (2009).
Pellizzoni, L. Chaperoning ribonucleoprotein biogenesis in health and disease. EMBO Rep. https://doi.org/10.1038/sj.embor.7400941 (2007).
Groen, E. J. N. et al. Temporal and tissue-specific variability of SMN protein levels in mouse models of spinal muscular atrophy. Hum. Mol. Genet. https://doi.org/10.1093/hmg/ddy195 (2018).
Tapia, O. et al. Cellular bases of the RNA metabolism dysfunction in motor neurons of a murine model of spinal muscular atrophy: role of Cajal bodies and the nucleolus. Neurobiol. Dis. https://doi.org/10.1016/j.nbd.2017.08.004 (2017).
Clamer, M. et al. Active ribosome profiling with RiboLace. Cell Rep. https://doi.org/10.1016/j.celrep.2018.09.084 (2018).
Lareau, L. F., Hite, D. H., Hogan, G. J. & Brown, P. O. Distinct stages of the translation elongation cycle revealed by sequencing ribosome-protected mRNA fragments. eLife 3, https://doi.org/10.7554/eLife.01257 (2014).
Hensel, N. & Claus, P. The actin cytoskeleton in SMA and ALS: how does it contribute to motoneuron degeneration? Neuroscientist https://doi.org/10.1177/1073858417705059 (2018).
Tisdale, S. et al. SMN is essential for the biogenesis of U7 small nuclear ribonucleoprotein and 3′-end formation of histone mRNAs. Cell Rep. https://doi.org/10.1016/j.celrep.2013.11.012 (2013).
Singh, R. N., Howell, M. D., Ottesen, E. W. & Singh, N. N. Diverse role of survival motor neuron protein. Biochim. Biophys. Acta https://doi.org/10.1016/j.bbagrm.2016.12.008 (2017).
Nölle, A. et al. The spinal muscular atrophy disease protein SMN is linked to the Rho-kinase pathway via profilin. Hum. Mol. Genet. https://doi.org/10.1093/hmg/ddr425 (2011).
Ramser, J. et al. Rare missense and synonymous variants in UBE1 are associated with X-linked infantile spinal muscular atrophy. Am. J. Hum. Genet. https://doi.org/10.1016/j.ajhg.2007.09.009 (2008).
Aghamaleky Sarvestany, A. et al. Label-free quantitative proteomic profiling identifies disruption of ubiquitin homeostasis as a key driver of Schwann cell defects in spinal muscular atrophy. J. Proteome Res. https://doi.org/10.1021/pr500492j (2014).
Fuller, H. R., Gillingwater, T. H. & Wishart, T. M. Commonality amid diversity: multi-study proteomic identification of conserved disease mechanisms in spinal muscular atrophy. Neuromuscul. Disord. https://doi.org/10.1016/j.nmd.2016.06.004 (2016).
Murray, L. M., Beauvais, A., Gibeault, S., Courtney, N. L. & Kothary, R. Transcriptional profiling of differentially vulnerable motor neurons at pre-symptomatic stage in the Smn2b/- mouse model of spinal muscular atrophy. Acta Neuropathol. Commun. https://doi.org/10.1186/s40478-015-0231-1 (2015).
Saal, L., Briese, M., Kneitz, S., Glinka, M. & Sendtner, M. Subcellular transcriptome alterations in a cell culture model of spinal muscular atrophy point to widespread defects in axonal growth and presynaptic differentiation. RNA https://doi.org/10.1261/rna.047373.114 (2014).
Rossoll, W. et al. Smn, the spinal muscular atrophy-determining gene product, modulates axon growth and localization of β-actin mRNA in growth cones of motoneurons. J. Cell Biol. https://doi.org/10.1083/jcb.200304128 (2003).
Bertrandy, S. et al. The RNA-binding properties of SMN: deletion analysis of the zebrafish orthologue defines domains conserved in evolution. Hum. Mol. Genet. https://doi.org/10.1093/hmg/8.5.775 (1999).
Ottesen, E. W., Singh, N. N., Luo, D. & Singh, R. N. High-affinity RNA targets of the survival motor neuron protein reveal diverse preferences for sequence and structural motifs. Nucleic Acids Res. https://doi.org/10.1093/nar/gky770 (2018).
Weingarten-Gabbay, S. et al. Comparative genetics: systematic discovery of cap-independent translation sequences in human and viral genomes. Science https://doi.org/10.1126/science.aad4939 (2016).
Zhang, H. L. et al. Active transport of the survival motor neuron protein and the role of exon-7 in cytoplasmic localization. J. Neurosci. 23, 6627–6637 (2003).
Jablonka, S. et al. Co-regulation of survival of motor neuron (SMN) protein and its interactor SIP1 during development and in spinal muscular atrophy. Hum. Mol. Genet. https://doi.org/10.1093/hmg/10.5.497 (2001).
Fuller, H. R. & Morris, G. E. SMN complexes of nucleus and cytoplasm: a proteomic study for SMA therapy. Transl. Neurosci. 1, 261–267 (2010).
Han, Y. et al. Ribosome profiling reveals sequence-independent post-initiation pausing as a signature of translation. Cell Res. 24, 842–851 (2014).
Matsuo, Y. et al. Ubiquitination of stalled ribosome triggers ribosome-associated quality control. Nat. Commun. 8, 159 (2017).
Wu, C. C.-C., Zinshteyn, B., Wehner, K. & Green, R. High-resolution ribosome profiling defines discrete ribosome elongation states and translational regulation during cellular stress. Mol. Cell 73, 959–970.e5 (2019).
Murray, L. M. et al. Selective vulnerability of motor neurons and dissociation of pre- and post-synaptic pathology at the neuromuscular junction in mouse models of spinal muscular atrophy. Hum. Mol. Genet. https://doi.org/10.1093/hmg/ddm367 (2008).
Kariya, S. et al. Reduced SMN protein impairs maturation of the neuromuscular junctions in mouse models of spinal muscular atrophy. Hum. Mol. Genet. https://doi.org/10.1093/hmg/ddn156 (2008).
Martínez-Hernández, R. et al. Synaptic defects in type i spinal muscular atrophy in human development. J. Pathol. https://doi.org/10.1002/path.4080 (2013).
Niebroj‐Dobosz, I. & Hausmanowa‐Petrusewicz, I. Serum cholinesterase activity in infantile and juvenile spinal muscular atrophy. Acta Neurol. Scand. https://doi.org/10.1111/j.1600-0404.1989.tb03864.x (1989).
Hutchinson, D. O. et al. Congential endplate acetylocholinesterase deficiency. Brain https://doi.org/10.1093/brain/116.3.633 (1993).
Hsieh-Li, H. M. et al. A mouse model for spinal muscular atrophy. Nat. Genet. 24, 66–70 (2000).
Riessland, M. et al. SAHA ameliorates the SMA phenotype in two mouse models for spinal muscular atrophy. Hum. Mol. Genet. https://doi.org/10.1093/hmg/ddq023 (2010).
Huang, Y. T. et al. Robust comparison of protein levels across tissues and throughout development using standardized quantitative western blotting. J. Vis. Exp. https://doi.org/10.3791/59438 (2019).
Viero, G. et al. Three distinct ribosome assemblies modulated by translation are the building blocks of polysomes. J. Cell Biol. 208, 581–596 (2015).
Chen, E., Sharma, M. R., Shi, X., Agrawal, R. K. & Joseph, S. Fragile X mental retardation protein regulates translation by binding directly to the ribosome. Mol. Cell https://doi.org/10.1016/j.molcel.2014.03.023 (2014).
Wessel, D. & Flügge, U. I. A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids. Anal. Biochem. https://doi.org/10.1016/0003-2697(84)90782-6 (1984).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Sleigh, J. N., Gillingwater, T. H. & Talbot, K. The contribution of mouse models to understanding the pathogenesis of spinal muscular atrophy. Dis. Model. Mech. https://doi.org/10.1242/dmm.007245 (2011).
Jones, R. A. et al. Cellular and molecular anatomy of the human neuromuscular junction. Cell Rep. https://doi.org/10.1016/j.celrep.2017.11.008 (2017).
Lauria, F. et al. riboWaltz: optimization of ribosome P-site positioning in ribosome profiling data. PLoS Comput. Biol. https://doi.org/10.1371/journal.pcbi.1006169 (2018).
Acknowledgements
We thank N. Polacek, University of Bern, for reading the manuscript and suggestions; D. Beeson, University of Oxford, for providing us with the FCC–Alexa Fluor 488 conjugate; staff at the IMPACT imaging facility at the University of Edinburgh for assistance with imaging; H. Zhou and F. Muntoni, University College London, for providing us with the ASO tissues; and staff at the Core Facilities Next Generation Sequencing Facility (NGS) and High Throughput Screening (HTS) at Department CIBIO University of Trento for technical support. This work was supported by Provincia Autonoma di Trento, Italy (AxonomiX research project), the UK SMA Research Consortium, SMA Europe, the Wellcome Trust (106098/Z/14/Z), Stichting Spieren voor Spieren (the Netherlands), the Slovenian Research Agency (program grant no. P1-0391), COST Action (CA15126), AFM-Telethon (reference no. 22129), Caritro Foundation (young post-doc funding grant) and Telethon (reference no. GGP19115). We also acknowledge financial support from IMMAGINA Biotechnology (Italy).
Author information
Authors and Affiliations
Contributions
F.L. and T.T. performed all of the high-throughput computational analyses. P.B. performed the SMN-specific Ribo-seq and sub-cellular fractionation experiments. P.B., E.P. and M.C. performed all other Ribo-seq library preparation. E.J.N.G. and T.H.G. generated and maintained all experimental animals, performed mouse tissue collection and the related western blotting and fluorescence microscopy of NMJs. F.M., A.I., J.O. and A.R. performed the cloning for dual luciferase experiments and the dual luciferase analysis. F.M. and A.R. performed all RT–qPCR analysis. D.D., N.O. and G.A. performed the SPR analysis. M.M, M.D.S. and G.V. performed the in vitro transcription translation experiments and data analysis. G.V. performed all polysomal purifications, RNA and protein extractions, western blotting and data analysis. F.L., T.T., E.J.N.G. and G.V. prepared the figures. G.V. conceived experiments and directed the research. A.Q., T.H.G. and G.V. obtained the funding. F.L., T.T., E.J.N.G., T.H.G. and G.V. wrote the manuscript. All of the authors contributed to the preparation, revision and writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.C. is the CEO of IMMAGINA Biotechnology; G.V. is a scientific advisor to IMMAGINA Biotechnology; T.H.G. has served on SMA advisory boards for Roche. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 SMN interacts with the translation machinery in vitro and in vivo in an RNA-independent manner.
a, Samples were ultracentrifuged to evaluate the aspecific sedimentation of SMN. The experiment was performed as in Fig. 1c, in the absence of ribosomes. The presence of SMN in the pellet was determined by western blot and densitometric analysis. b, Saturation curve obtained using experiments as in Fig. 1. The unspecific binding was obtained as in panel (a). The data represent the average ± SEM among n=3 independent experiments. c, Western blot analysis on P7 cytoplasmic lysates from control P7 spinal cord. The experiment was performed as in Fig. 1e and was repeated independently 2 times with similar results. d-e, Polysomal profiling and co-sedimentation profiles of lysates obtained from control brain before (d) and after (e) RNAse I treatment. RPL26 and RPS6 were used as sedimentation controls for the large and small ribosomal subunits, respectively. The sedimentation distribution of proteins along the sucrose gradient is expressed as percentage in each fraction considering as 100% the sum of all fractions. Data in (d) report the average ± SEM among n=3 biologically independent samples. f, Polysomal profile and co-sedimentation analysis of SMN and ribosome markers RPL26 and RPS6 in Hek-293 using a lysis buffer with high 10 mM Mg2+. The distribution of SMN along the profile was fitted with three Gaussian curves to identify different populations. g, Polysomal profile and co-sedimentation analysis of SMN and ribosome markers in Hek-293 lysed in the absence of Mg2+ to release the large and small subunits. RPL26 and RPS6 were used as sedimentation controls for the large and small ribosomal subunits, respectively. The sedimentation distribution of proteins along the sucrose gradient is expressed as in (d). The distributions of RPS6, RPL26 and SMN along the profile were fitted with one or two Gaussian curves. Statistical source data and unprocessed blots are provided in Source data Extended data Fig. 1.
Extended Data Fig. 2 SMN interacts with the translation machinery in a concentration dependent manner across different tissues and is associated to actively translating ribosomes positively regulating translation.
a, Co-sedimentation of SMN proteins with RiboNucleoParticles (snRNP/RNPs) and ribosomal subunits in Hek-293 lysates. Each lane of the western blot corresponds to a fraction along the profile. Gemin 5 was used as marker of the Gemin granules and HuR of RNA-granules. b-d, Polysomal profiling and co-sedimentation profiles of lysates obtained from P3 control mouse spinal cord (b), heart (c) and kidney (d). The ribosomal proteins RPL26 and RPS6 were used as sedimentation controls for the large and small ribosomal subunits, respectively. The sedimentation distribution of proteins along the sucrose gradient is expressed as percentage in each fraction considering as 100% the sum of all fractions. The percentage shown are average ± SEM from n=3 biologically independent experiments. Statistical source data and unprocessed blots are provided in Source data Extended data Fig. 2.
Extended Data Fig. 3 Ribosome profiling of SMN-primed ribosomes reveals enriched mRNAs organized in functionally well-defined communities.
a, Examples of immunoprecipitation of SMN-ribosomes from ribo-pellet in control mouse brain lysates after RNaseI treatment. RPL26 was used as control. The experiment was repeated independently three times. b, Transcript types enriched in fragments protected by SMN-primed ribosomes that protect predominantly RNAs associated with protein coding genes. c, Venn diagram showing the intersection of SMN-specific protein coding transcripts covered by short (24–26 nucleotides) and long (32–34 nucleotides) reads. d, Percentage of P-sites according to the three reading frames for the first five codons of the CDSs and for the remaining region of the sequences. A clear trinucleotide periodicity in the correct frame is detectable only at the beginning of the coding sequence. Results are shown as the average ± SEM among n=3 biologically independent samples. The statistical significance from two-sided t-test comparing frames 1 and 2 with respect to frame 0 are reported. e, Representative RPF coverage tracks for SMN-specific transcripts in SMN-specific RiboSeq. The average normalized coverage and the standard error among replicates are represented in each profile. The structure of the transcript, showing the boundaries of CDS and UTR regions, is outlined below each profile. f, Over-representation of tissue-specific markers among genes enriched in SMN-primed ribosomes. Tissues where SMN levels were previously measured are displayed. The enrichment score was calculated with enrichR. g, Gene ontology and pathway enrichment analysis of genes enriched in SMN-primed ribosomes. The number of genes is displayed on the right of the bars. (GO_BP:biological process, GO_CC:cellular component, GO_MF: molecular function). h, Comparative annotation enrichment analysis of transcripts belonging to the seven SMN-specific communities. The heatmaps are colored according to the significance of the enrichments. The analysis was performed on Gene Ontology terms: Biological Process (left) and cellular component (right). Statistical source data and unprocessed blots are provided in Source data Extended data Fig. 3.
Extended Data Fig. 4 Transcripts bound by SMN-primed ribosomes display defects in positioning of active ribosomes at early stages of SMA.
a, mTORC1 is activated only at late stages of SMA. Upper panel, total proteins from brains at early and late stages of disease (SMA) and from littermate controls (Ctrl) were probed for phosphorylation responses by immunoblotting of known mTORC1 downstream targets (4E-BP) and PERK (eIF2a). In the right panels the quantification for 4E-BP is shown as the average signal ± SEM among n=3 biologically independent samples (normalized to total protein stain, not shown). The statistical significance from one-way ANOVA with Tukey’s multiple comparisons test is reported. b, Fraction of ribosomes in polysomes obtained from polysomal profiling of control and SMA mouse brain at early stage of disease (P5). The data represent the average ± SEM (n = 3 mice). Significant decreases were tested by two-sided t-test. c, Pairwise correlations of RPKM per protein coding transcripts (n=49825 mRNAs) between the two replicas of control and SMA Active-RiboSeq and control RiboSeq (upper panels) and between the three replicas of control SMN-specific RiboSeq (lower panels). Pearson’s correlation coefficients and statistical significance from two-sided Williams’ test are shown (*** pvalue < 1e-16). d, Over-representation of tissue-specific markers among genes with significantly increased (yellow) or decreased (red) active ribosome occupancies in SMA. Tissues where SMN levels were previously measured are displayed in bold. The enrichment score was calculated with enrichR. Statistical source data and unprocessed blots are provided in Source data Extended data Fig. 4.
Extended Data Fig. 5 Translationally defective transcripts in SMA display specific features.
a, Localization of proteins synthesized by mRNAs enriched in SMN-primed ribosomes. Manually annotated and reviewed proteins from the UniProt database were considered. b, Lengths comparison of regions of SMN-specific transcripts (n=874) compared with unspecific transcripts (n=4010). Statistical significance was determined with one-sided Wilcoxon-Mann-Whitney tests. c, Logo representation of top enriched motifs in the 5`UTR (left panel) and 3’ UTR (right panel) of SMN-specific transcripts, by discriminative analysis of k-mer composition. d, Over-representation of translational enhancer sequences among SMN-specific transcripts (619, in blue). Annotation of translational enhancer was retrieved from50. two-sided Wilcoxon rank-sum test to calculate the p-value shown. e, Logo-like representation of the most frequent amino-acids in SMN-specific (left) and unspecific RNAs (right) at the beginning of the CDS. f, SMN levels in NSC-34 expressing high and low SMN levels. Comparison between the fraction of ribosomes in polysomes (FRP) in NSC-34 expressing high and low levels of SMN. The results are the average ± SEM. Significant differences were determined using a two-sided t-test. g, Luciferase assays were performed in NSC-34 expressing high or low levels of SMN. Alanine repeats values were set to 1. h, The 5’UTR of Tuba4a was used as control. Results for luciferase assays with or without treatment with rapamycin are reported. In panels (g) and (h) the number of biologically independent experiments is reported. Results are shown as the average ± SEM. Significant changes were assessed using one-sided t-test. i, Fold change of Renilla and Firefly luciferase in cells expressing high vs low levels of SMN. The level of the two transcripts in cells expressing high levels of SMN was set to 1. The number of biologically independent samples is reported and the data represent the average ± SEM. Statistical source data and unprocessed blots are provided in Source data Extended data Fig. 5.
Extended Data Fig. 6 Communities of mRNAs bound by SMN-primed ribosomes show reduced ribosome occupancy.
a, Comparison between total RNA fold changes (SMA vs CTRL) in SMN-specific genes, binned in the 7 SMN-communities. Data were retrieved from Bernabò et al., 201711. Significant shifts in each community were identified with the two-sided one-sample Wilcoxon rank-sum test. The number of genes for each community is reported in Fig. 3f. b, Representative polysomal profiles obtained from CTRL and SMA mouse brains (early-symptomatic). c, Representative active RiboSeq coverage tracks for SMN-specific transcripts selected for validation by polysomal profiling. The average normalized coverage and the standard error among replicates are represented in each profile. The structure of the transcript, showing the boundaries of CDS and UTR regions, is outlined below each profile. Statistical source data are provided in Source data Extended data Fig. 6.
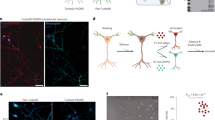
Extended Data Fig. 7 The acetylcholinesterase transcript shows ribosome drop-off and defective production of protein at the NMJ in SMA.
a, Translation efficiency of Acetylcholinesterase from brain at early and late stage and in spinal cord at late stage, and after SMN-restoring Antisense Oligonucleotides treatment11. The average ± SEM from n=3 biologically replicates in n=2 technical replicates is shown. For control brain early stage the average ± SEM from n=3 biologically replicates in n=3 technical replicate is shown. Translation efficiencies were obtained as the ratio of changes in polysomal and total RNA normalized to Tuba4a. Significant differences were determined using a two-sided t- test. b, Luciferase assays were performed in NSC34 expressing high or low levels of SMN in n=9 biologically independent replicates. c, Luciferase assays for testing the contribution of the first five codons of AChE with respect to four alanines. In (b) and (c) the number of biologically independent experiments is reported. Results are the average ± SEM. Significant changes were assessed using one-sided t-test. d, Brain, spinal cord and muscle lysates of early-symptomatic SMA mice were compared to controls using western blot. Experiments were repeated independently 3 times with similar results. e, Representative images for control (left) and early-symptomatic SMA mouse (right) neuromuscular endplates. Acetylcholine receptors (AChR) were labelled using alpha-bungarotoxin (BTX) and acetylcholinesterase (AChE) using fasciculin-2 (FCC). Top panels show FCC / BTX overlap, middle and bottom panels individual channels in greyscale. Endplates n=65 from control and n=73 endplates from SMA mice from 6 muscles and 3 mice/genotype were imaged and analysed. Scale bar: 10 μm. f, FCC and BTX average intensity were determined for 2 FDB muscles in 3 control and 3 SMA mice. The number of endplates per mouse is reported. Results are the average fluorescence intensity ± SEM. Significant differences were determined using a two-sided t-test. Statistical source data and unprocessed blots are provided in Source data Extended data Fig. 7.
Extended Data Fig. 8 Schematic representation of the role of SMN-primed ribosomes in translation based on SMN-specific ribosome profiling1.
After recruitment of ribosomes at the AUG, SMN-primed ribosomes are translating the first codons and stop at the fifth codon in a rotated state, according to the results shown in Fig. 3 and to Laureau et al., 201437. The ribosome pause at the fifth codon ensures productive translation as proposed by Han et al., 201454 and in accordance with our results (Figs. 3 and 4). The blue star represents SMN2. When SMN expression is lost in SMA, SMN-specific transcripts are bound by SMN-free ribosomes. These ribosomes cannot properly pause within the first 5 codons, causing ribosomes to drop-off (as observed by Han et al., 201454, leading to disruption of normal protein synthesis.
Supplementary information
Supplementary Tables
Supplementary Table 1: binding parameters from SPR experiments. Supplementary Table 2a: list of transcripts significantly enriched for SMN-primed ribosomes. Supplementary Table 2b: list of transcripts displaying significant alterations in active ribosome occupancy in SMA. Supplementary Table 2c: data and mapping statistics from all RNA-seq analyses of brains control and at early-symptomatic SMA stage. Supplementary Table 3: list of genes significantly enriched for SMN-primed ribosomes with significant alterations in active ribosome occupancy in SMA. Supplementary Table 4: list of primers for 5′ UTR cloning and SMN-specific motifs. Supplementary Table 5: list of primers for ddPCR and RT–qPCR. Supplementary Table 6: list of primers for ribosome profiling.
Source data
Source Data Fig. 1
Statistical source data for Fig. 1.
Source Data Fig. 1
Unprocessed western blots for Fig. 1.
Source Data Fig. 2
Statistical source data for Fig. 2.
Source Data Fig. 2
Unprocessed western blots for Fig. 2.
Source Data Fig. 3
Statistical source data for Fig. 3.
Source Data Fig. 4
Statistical source data for Fig. 4.
Source Data Fig. 5
Statistical source data for Fig. 5.
Source Data Fig. 6
Statistical source data for Fig. 6.
Source Data Fig. 6
Unprocessed western blots for Fig. 6.
Source Data Fig. 7
Statistical source data for Fig. 7.
Source Data Fig. 7
Unprocessed western blots for Fig. 7.
Source Data Extended Data Fig. 1
Statistical source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 1
Unprocessed western blots for Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Statistical source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 2
Unprocessed western blots for Extended Data Fig. 2.
Source Data Extended Data Fig. 3
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 3
Unprocessed western blots for Extended Data Fig. 3.
Source Data Extended Data Fig. 4
Statistical source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 4
Unprocessed western blots for Extended Data Fig. 4.
Source Data Extended Data Fig. 5
Statistical source data for Extended Data Fig. 5.
Source Data Extended Data Fig. 5
Unprocessed western blots for Extended Data Fig. 5.
Source Data Extended Data Fig. 6
Statistical source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7
Statistical source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 7
Unprocessed western blots for Extended Data Fig. 7.
Rights and permissions
About this article
Cite this article
Lauria, F., Bernabò, P., Tebaldi, T. et al. SMN-primed ribosomes modulate the translation of transcripts related to spinal muscular atrophy. Nat Cell Biol 22, 1239–1251 (2020). https://doi.org/10.1038/s41556-020-00577-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-020-00577-7
This article is cited by
-
SMN deficiency perturbs monoamine neurotransmitter metabolism in spinal muscular atrophy
Communications Biology (2023)
-
Principles, challenges, and advances in ribosome profiling: from bulk to low-input and single-cell analysis
Advanced Biotechnology (2023)
-
The Akt/mTOR and MNK/eIF4E pathways rewire the prostate cancer translatome to secrete HGF, SPP1 and BGN and recruit suppressive myeloid cells
Nature Cancer (2023)
-
Impaired dynamic interaction of axonal endoplasmic reticulum and ribosomes contributes to defective stimulus–response in spinal muscular atrophy
Translational Neurodegeneration (2022)
-
Mouse models of SMA show divergent patterns of neuronal vulnerability and resilience
Skeletal Muscle (2022)