Abstract
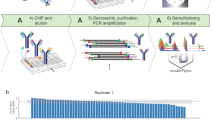
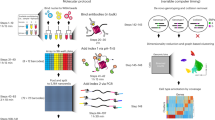
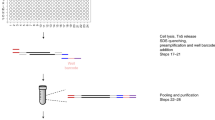
Single-cell measurement of chromatin states, including histone modifications and non-histone protein binding, remains challenging. Here, we present a low-cost, efficient, simultaneous indexing and tagmentation-based ChIP-seq (itChIP-seq) method, compatible with both low cellular input and single cells for profiling chromatin states. itChIP combines chromatin opening, simultaneous cellular indexing and chromatin tagmentation within a single tube, enabling the processing of samples from tens of single cells to, more commonly, thousands of single cells per assay. We demonstrate that single-cell itChIP-seq (sc-itChIP-seq) yields ~9,000 unique reads per cell. Using sc-itChIP-seq to profile H3K27ac, we sufficiently capture the earliest epigenetic priming event during the cell fate transition from naive to primed pluripotency, and reveal the basis for cell-type specific enhancer usage during the differentiation of bipotent cardiac progenitor cells into endothelial cells and cardiomyocytes. Our results demonstrate that itChIP is a widely applicable technology for single-cell chromatin profiling of epigenetically heterogeneous cell populations in many biological processes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Code availability
Codes for analysis of the figures are available at https://github.com/Helab-bioinformatics/itChIP.
Data availability
All sequencing data have been deposited in the GEO database at accession code GSE109762. Previously published data reanalysed in this work are available under accession codes GSE22075 (Nano-ChIP), GSE63523 (ULI-NChIP), GSE65360 (scATAC-seq H1ESC, Fig. 3b,c), GSE31039 (bulk H3K27ac mESC), GSE56098 (bulk H3K27ac EpiLCs), GSM2474111 (bulk Pol II ChIP), GSM590132 (bulk EZH2 ChIP), GSE70253 (scDrop-ChIP, H3K4me2 profile for mESC), GSE100033 (scATAC-seq, profile at postnatal day (P) 56, Supplementary Table 4), GSE105012 (scChIC-seq, H3K4me3 profile for human white blood cells), GSE111121 (scCUT&RUN, SOX2 ChIP for mESC), GSE124557 (CUT&Tag, H3K4me2 profile for K562 cells) and GSE108963 (scRNA-seq, E7.75 and E8.25 heart). Source data for Figs. 1b,d, 2, 3b,c, 4b and 6c,e and Supplementary Figs. 1b,d,f,h, 2b,c and 4 have been provided in Supplementary Table 7. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Rivera, C. M. & Ren, B. Mapping human epigenomes. Cell 155, 39–55 (2013).
Roh, T.-Y., Ngau, W. C., Cui, K., Landsman, D. & Zhao, K. High-resolution genome-wide mapping of histone modifications. Nat. Biotechnol. 22, 1013–1016 (2004).
Kharchenko, P. V., Tolstorukov, M. Y. & Park, P. J. Design and analysis of ChIP-seq experiments for DNA-binding proteins. Nat. Biotechnol. 26, 1351–1359 (2008).
Stegle, O., Teichmann, S. A. & Marioni, J. C. Computational and analytical challenges in single-cell transcriptomics. Nat. Rev. Genet. 16, 133–145 (2015).
Adli, M. & Bernstein, B. E. Whole-genome chromatin profiling from limited numbers of cells using nano-ChIP-seq. Nat. Protoc. 6, 1656–1668 (2011).
Brind’Amour, J. et al. An ultra-low-input native ChIP-seq protocol for genome-wide profiling of rare cell populations. Nat. Commun. 6, 6033 (2015).
Zheng, H. et al. Resetting epigenetic memory by reprogramming of histone modifications in mammals. Mol. Cell 63, 1066–1079 (2016).
Cao, Z., Chen, C., He, B., Tan, K. & Lu, C. A microfluidic device for epigenomic profiling using 100 cells. Nat. Methods 12, 959–962 (2015).
Skene, P. J., Henikoff, J. G. & Henikoff, S. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat. Protoc. 13, 1006–1019 (2018).
Harada, A. et al. A chromatin integration labelling method enables epigenomic profiling with lower input. Nat. Cell. Biol. 21, 287–296 (2019).
Rotem, A. et al. Single-cell ChIP-seq reveals cell subpopulations defined by chromatin state. Nat. Biotechnol. 33, 1165–1172 (2015).
Schmidl, C., Rendeiro, A. F., Sheffield, N. C. & Bock, C. ChIPmentation: fast, robust, low-input ChIP-seq for histones and transcription factors. Nat. Methods 12, 963–965 (2015).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Daley, T. & Smith, A. D. Predicting the molecular complexity of sequencing libraries. Nat. Methods 10, 325–327 (2013).
Lake, B. B. et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol. 36, 70–80 (2018).
Nord, A. S. et al. Rapid and pervasive changes in genome-wide enhancer usage during mammalian development. Cell 155, 1521–1531 (2013).
Zhou, P. et al. Mapping cell type-specific transcriptional enhancers using high affinity, lineage-specific Ep300 bioChIP-seq. Elife 6, e22039 (2017).
Cui, K. R. et al. Chromatin signatures in multipotent human hematopoietic stem cells indicate the fate of bivalent genes during differentiation. Cell Stem Cell 4, 80–93 (2009).
Amini, S. et al. Haplotype-resolved whole-genome sequencing by contiguity-preserving transposition and combinatorial indexing. Nat. Genet. 46, 1343–1349 (2014).
Vitak, S. A. et al. Sequencing thousands of single-cell genomes with combinatorial indexing. Nat. Methods 14, 302–308 (2017).
Kurimoto, K. et al. Quantitative dynamics of chromatin remodeling during germ cell specification from mouse embryonic stem cells. Cell Stem Cell 16, 517–532 (2015).
Buecker, C. et al. Reorganization of enhancer patterns in transition from naive to primed pluripotency. Cell Stem Cell 14, 838–853 (2014).
Preissl, S. et al. Single-nucleus analysis of accessible chromatin in developing mouse forebrain reveals cell-type-specific transcriptional regulation. Nat. Neurosci. 21, 432–439 (2018).
Ibarra-Soria, X. et al. Defining murine organogenesis at single-cell resolution reveals a role for the leukotriene pathway in regulating blood progenitor formation. Nat. Cell Biol. 20, 127–134 (2018).
Loebel, D. A., Watson, C. M., De Young, R. A. & Tam, P. P. Lineage choice and differentiation in mouse embryos and embryonic stem cells. Dev. Biol. 264, 1–14 (2003).
Wu, S. M., Chien, K. R. & Mummery, C. Origins and fates of cardiovascular progenitor cells. Cell 132, 537–543 (2008).
Stanley, E. G. et al. Efficient Cre-mediated deletion in cardiac progenitor cells conferred by a 3’UTR-ires-Cre allele of the homeobox gene Nkx2-5. Int. J. Dev. Biol. 46, 431–439 (2002).
Qiu, X. J. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
de Jonge, H. J. M. et al. Evidence based selection of housekeeping genes. PLoS One 2, e898 (2007).
Dix, A. et al. Biomarker-based classification of bacterial and fungal whole-blood infections in a genome-wide expression study. Mycoses 58, 57–58 (2015).
Hainer, S. J., Boskovic, A., McCannell, K. N., Rando, O. J. & Fazzio, T. G. Profiling of pluripotency factors in single cells and early embryos. Cell 177, 1319–1329 (2019).
Ku, W. L. et al. Single-cell chromatin immunocleavage sequencing (scChIC-seq) to profile histone modification. Nat. Methods 16, 323–325 (2019).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Buenrostro, J. D. et al. Single-cell chromatin accessibility reveals principles of regulatory variation. Nature 523, 486–490 (2015).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Ying, Q. L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008).
Islam, S. et al. Highly multiplexed and strand-specific single-cell RNA 5′ end sequencing. Nat. Protoc. 7, 813–828 (2012).
Islam, S. et al. Quantitative single-cell RNA-seq with unique molecular identifiers. Nat. Methods 11, 163–166 (2014).
Xiao, Y. et al. Hippo signaling plays an essential role in cell state transitions during cardiac fibroblast development. Dev. Cell 45, 153–169 (2018).
Ai, S. et al. ItChIP-simultaneous indexing and tagmentation-based ChIP-seq. Protoc. Exch. https://doi.org/10.21203/rs.2.11366/v1 (2019).
Ai, S. et al. EED orchestration of heart maturation through interaction with HDACs is H3K27me3-independent. Elife 6, e24570 (2017).
Picelli, S. et al. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative genomics viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. 14, 178–192 (2013).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Cusanovich, D. A. et al. The cis-regulatory dynamics of embryonic development at single-cell resolution. Nature 555, 538–542 (2018).
Acknowledgements
We thank W.T. Pu at Boston Children’s Hospital for the critical and constructive improvement of the manuscript, all members of He laboratory for discussion and the High Performance Computing Platform of the Center for Life Sciences, Peking University. A.H. was supported by grants from the National Basic Research Program of China (2017YFA0103402), the National Natural Science Foundation of China (31571487 and 31771607), Peking-Tsinghua Center for Life Sciences, and the 1000 Youth Talents Program of China.
Author information
Authors and Affiliations
Contributions
A.H. conceived the project and designed experiments. S.A., C.C.L., Y. Luo and X.Y. performed experiments. H.X., C.C.L., Q.S. and Y. Liu performed bioinformatic analyses with technical assistance from C.L. A.H. supervised the study. A.H., S.A., H.X., C.C.L., and Y. Luo wrote the paper with inputs from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors have submitted patent applications (application nos 2018101211756 and 2018101428753) in China based on the results reported in this paper.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Optimization of key steps in the itChIP protocol.
a, b, Optimization of the SDS concentration for chromatin opening with 10,000 RT 7 min fixed ESCs. We picked the condition yielding desired DNA fragments of < 1 kb (a), as those > 1 kb were undesired for sequencing, and higher ChIP-qPCR enrichment (b) for itChIP. 0.3% SDS was selected accordingly. P values were calculated by one-way ANOVA followed by Bonferroni’s post hoc test for multiple comparisons. c, d, Optimization of the Tn5 complex amount for efficient tagmentation. The same as in (a), except for the indicated Tn5 amount. We picked the condition yielding DNA fragments of < 1 kb (c) and higher ChIP-qPCR enrichment (d) for itChIP. 3 μl of 12.5 μM Tn5 transposase was selected accordingly. P values were calculated by one-way ANOVA followed by Bonferroni’s post hoc test for multiple comparisons. e, f, Optimization of the temperature for chromatin opening. The same as in (a), except for chromatin opening conditions, 37 °C for 1 h and 62 °C for 10 min. We picked the condition yielding DNA fragments of < 1 kb (e) and higher ChIP-qPCR enrichment (f) for itChIP. In this regard, both conditions exhibited no difference. For simplicity, we used 62 °C for 10 min to open chromatin in this study. P values were calculated by two-tailed paired t-test. g, h, Optimization of tagmentation conditions. The same as in (a), except for the indicated temperature and time for tagmentation. We picked the condition yielding DNA fragments of < 1 kb (g) and higher ChIP-qPCR enrichment (h) for itChIP. In this regard, both conditions can be used for chromatin tagmentation. For simplicity, we used 37 °C for 1 h for chromatin tagmentation in this study. P values were calculated by two-tailed paired t-test. Optimal conditions were boxed. Dish lines in a, c, e, g indicate the position of 1 kb DNA marker. In b, d, f and h, data represent means ± s.d. from n=3 biological replicates. Arrowhead indicates free adaptors. Source data for b, d, f, h is shown in Supplementary Table 7. Unprocessed blots are shown in Supplementary Fig. 7.
Supplementary Figure 2 Characterization of H3K4me3 itChIP-seq across different sonication conditions for soluble chromatin release and varying cell numbers.
a, Track view displaying H3K4me3 itChIP signals at representative loci using indicated chromatin releasing conditions in 500 ESCs. Notably, although the group under no shearing to aid in chromatin releasing gave rise to the indistinguishable ChIP signals from 500 cells, it required a few more PCR cycles to achieve the similar yield in the library preparation. We reasoned that for single-cell itChIP experiments brief sonication may produce the better yield and complexity of the resulting libraries. Sonication-ChIP from 1 M ESCs. Two biological replicates were shown above. b, Scatter plots showing pairwise comparison between three groups and sonication-ChIP as in (a). Pearson correlation coefficients were calculated using the signals at ± 5 kb of gene TSS. Two biological replicates were shown above. c, Scatter plots showing pairwise comparison of H3K4me3 in between itChIP using varying cell numbers and bulk sonication-ChIP. Data were measured in fragments per million reads at regions of ± 5 kb of gene TSS and Pearson correlation coefficients are shown in each box. n = 2 biologically independent experiments, and the aggregation of two replicates for each condition was used here. d, Characteristic fraction of itChIP-seq reads. H3K4me3 from two independent biological replicates using indicated cell numbers (upper) and an example of the average yield and PCR cycles in two library preparations (bottom). e, The library complexity is shown for each itChIP group. The number of distinct reads over all mapped reads in each group was plotted.
Supplementary Figure 3 Examples of a repressive histone marker and non-histone proteins itChIP-seq in low-input cells.
a, e, Track view displaying RNA polymerase II (Pol II) (a), EZH2 (e) and itChIP signals on representative regions in ESCs. b, f, Heatmap showing Pol II (b), EZH2 (f) and itChIP signals at peak regions. All ChIP signals represent the average ChIP read intensity of the biological replicates. The rows were sorted by the descending signals in 6623 peaks as in (b) and 9192 peaks in (f). c, g, Average Pol II (c), EZH2 (g) and H3K27me3 itChIP signals from different groups plotted centered the peaks. Each curve represents the average signals of the biological replicates. Cell numbers are indicated in each box. Bulk data of Pol II and EZH2 were downloaded from GSM2474111 and GSM590132. Data shown represent 2 biologically independent experiments, and the aggregation of two replicates for each condition was used here. d, h, Genomic feature distributions of itChIP peaks for Pol II (d) and EZH2 (h). Merged peaks of two biological replicates of Pol II and EZH2 itChIP data were used to calculate the distribution fraction.
Supplementary Figure 4 Custom Nextera sequencing strategy and data quality of sc-itChIP libraries.
a, T5 and T7 barcodes are introduced during Tn5 complex assembly with transposon oligonucleotides, and i5 and i7 barcodes are introduced during PCR enrichment for sequencing library preparation on Illumina Hiseq 2500 before size selection with AMPure XP beads. Modified from the study by Amini et al. 19. b, Size distribution of single-cell itChIP-seq data from two independent experiments. c, Scatter plots showing genome-wide correlation of two independent experiments as in (b). ChIP signals were measured in non-overlapping 5 kb windows genome wide. Pearson correlation coefficient (r) is shown in the box. d, Distribution of unique, non-duplicated reads per cell in two independent experiments of 1:10 mixed ESCs and differentiating ESCs (EpiLCs) after filtering single cells of potential barcodes with <3,000 reads.
Supplementary Figure 5 Clustering of sc-itChIP data of Nkx2-5 lineages from E7.75 and E8.25 hearts.
a, Figure exemplifying the gating strategy. FACS scheme for the isolation of E7.75 and E8.25 Tomato positive NKX2.5 lineage cells using fluorescent-minus-cell as controls. The gate shown on each plot is indicated above the plot. b, c, MDS plot of single cells colored by clusters (b) and embryonic stages (c). CP, cardiac progenitor cells; EC, endothelial cells; CM, cardiomyocytes. Cells were clustered using the method in Preissl at al23. d, Scatter plot showing the enhancer activity of feature genes. The representative marker genes were selected for the cell type validation of CP, EC and CM respectively. Red and grey indicate on/off of enhancer activity in each single cell. Single cells were from two biologically independent experiments.
Supplementary Figure 6 Validation of scRNA-seq and sc-itChIP-seq data of Nkx2-5 cardiac cell lineages from E7.75 and E8.25 hearts.
a, Pseudotime analysis showing two trajectories of cardiomyocyte (CM) and endothelial cell (EC) lineage starting from cardiac progenitor cell population (CP) at E7.75 and E8.25. Single cells were from two biologically independent experiments. b, Scatter plot showing the marker gene expression of single cells over the developmental pseudotime. Three representative marker genes in each row were selected for the cell type validation of EC, CM and CP respectively. c, Heatmap showing H3K27ac signals around the housekeeping genes. The columns represent single cells as ordered in Fig. 6f. Single cells were from two biologically independent experiments. The housekeeping gene lists are from the studies by Dix et al.30. and De Jonge et al.29. d, Scatter plot showing the H3K27ac signals at two representative housekeeping gene regions on tSNE as in Fig. 6e. Single cells were from two biologically independent experiments. e, Pseudotime analysis showing two trajectories of cardiomyocyte (CM) and endothelial cell (EC) lineage starting from cardiac progenitor cell population (CP) at E7.75 and E8.25. Single cells were from two biologically independent experiments. f, Scatter plot showing H3K27ac signals at peak regions nearby feature genes in single cells of CM and EC lineages corresponding to Fig. 6e. H3K27ac signals were calculated by counting the reads at peak regions and then normalized by read depth. g, A full heatmap of all clusters showing the normalized H3K27ac signals of three “pseudocells” (CP, EC and CM) as in Fig. 6g at peak regions, related to Fig. 6h. Peaks were clustered into 4 groups by K-means. C1, C2 and C3 were positively correlated with CP, CM and EC respectively, and C4 was the negatively correlated peaks. Of note, no signals were detected in a large number of regions (rows), probably due to the fact that those are enriched in other single cells rather than from the ending cell populations analyzed here.
Supplementary Figure 7 Unprocessed gel images used in this study.
Unprocessed gel images used in Supplementary Fig. 1. Boxes indicated cropped region shown in representative figures as indicated.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7 and titles and legends for Supplementary Tables 1–7
Rights and permissions
About this article
Cite this article
Ai, S., Xiong, H., Li, C.C. et al. Profiling chromatin states using single-cell itChIP-seq. Nat Cell Biol 21, 1164–1172 (2019). https://doi.org/10.1038/s41556-019-0383-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0383-5
This article is cited by
-
Single-cell sequencing technology applied to epigenetics for the study of tumor heterogeneity
Clinical Epigenetics (2023)
-
SingleScan: a comprehensive resource for single-cell sequencing data processing and mining
BMC Bioinformatics (2023)
-
BIND&MODIFY: a long-range method for single-molecule mapping of chromatin modifications in eukaryotes
Genome Biology (2023)
-
Single-cell sortChIC identifies hierarchical chromatin dynamics during hematopoiesis
Nature Genetics (2023)
-
Characterizing cis-regulatory elements using single-cell epigenomics
Nature Reviews Genetics (2023)