Abstract
Cells of multicellular organisms need to adopt specific morphologies. However, the molecular mechanisms bringing about membrane topology changes are far from understood—mainly because knowledge of membrane-shaping proteins that can promote local membrane curvatures is still limited. Our analyses unveiled that several members of a large, previously unrecognised protein family, which we termed N-Ank proteins, use a combination of their ankyrin repeat array and an amino (N)-terminal amphipathic helix to bind and shape membranes. Consistently, functional analyses revealed that the N-Ank protein ankycorbin (NORPEG/RAI14), which was exemplarily characterised further, plays an important, ankyrin repeat-based and N-terminal amphipathic helix-dependent role in early morphogenesis of neurons. This function furthermore required coiled coil-mediated self-assembly and manifested as ankycorbin nanodomains marked by protrusive membrane topologies. In summary, here, we unveil a class of powerful membrane shapers and thereby assign mechanistic and cell biological functions to the N-Ank protein superfamily.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Peter, B. J. et al. BAR domains as sensors of membrane curvature: the amphiphysin BAR structure. Science 303, 495–499 (2004).
McMahon, H. T. & Gallop, J. L. Membrane curvature and mechanisms of dynamic cell membrane remodelling. Nature 438, 590–596 (2005).
Frost, A., Unger, V. M. & De Camilli, P. The BAR domain superfamily: membrane-molding macromolecules. Cell 137, 191–196 (2009).
Qualmann, B., Koch, D. & Kessels, M. M. Let’s go bananas: revisiting the endocytic BAR code. EMBO J. 30, 3501–3515 (2011).
Voeltz, G. K., Prinz, W. A., Shibata, Y., Rist, J. M. & Rapoport, T. A. A class of membrane proteins shaping the tubular endoplasmic reticulum. Cell 124, 573–586 (2006).
Breeden, L. & Nasmyth, K. Similarity between cell-cycle genes of budding yeast and fission yeast and the notch gene of Drosophila. Nature 329, 651–654 (1987).
Pluckthun, A. Designed ankyrin repeat proteins (DARPins): binding proteins for research, diagnostics, and therapy. Annu. Rev. Pharmacol. Toxicol. 55, 489–511 (2015).
Mosavi, L. K., Cammett, T. J., Desrosiers, D. C. & Peng, Z. Y. The ankyrin repeat as molecular architecture for protein recognition. Protein Sci. 13, 1435–1448 (2004).
Li, J., Mahajan, A. & Tsai, M. D. Ankyrin repeat: a unique motif mediating protein–protein interactions. Biochemistry 45, 15168–15178 (2006).
Sedgwick, S. G. & Smerdon, S. J. The ankyrin repeat: a diversity of interactions on a common structural framework. Trends Biochem. Sci. 24, 311–316 (1999).
Takahashi, N. et al. TRPV4 channel activity is modulated by direct interaction of the ankyrin domain to PI(4,5)P2. Nat. Commun. 5, 4994 (2014).
Kim, D. H. et al. An ankyrin repeat domain of AKR2 drives chloroplast targeting through coincident binding of two chloroplast lipids. Dev. Cell 30, 598–609 (2014).
Peng, Y. F. et al. Ankycorbin: a novel actin cytoskeleton-associated protein. Genes Cells 5, 1001–1008 (2000).
Kutty, R. K. et al. Molecular characterization and developmental expression of NORPEG, a novel gene induced by retinoic acid. J. Biol. Chem. 276, 2831–2840 (2001).
Sakai, T., Liu, L., Shishido, Y. & Fukui, K. Identification of a novel, embryonal carcinoma cell-associated molecule, Nucling, that is up-regulated during cardiac muscle differentiation. J. Biochem. 133, 429–436 (2003).
Gallop, J. L. et al. Mechanism of endophilin N-BAR domain-mediated membrane curvature. EMBO J. 25, 2898–2910 (2006).
Schneider, K. et al. ProSAP1 and membrane nanodomain-associated syndapin I promote postsynapse formation and function. J. Cell Biol. 205, 197–215 (2014).
Seemann, E. et al. Deciphering caveolar functions by syndapin III KO-mediated impairment of caveolar invagination. eLife 6, e29854 (2017).
Dang, H. V. et al. Nucling, a novel apoptosis-associated protein, controls mammary gland involution by regulating NF-κB and STAT3. J. Biol. Chem. 290, 24626–24635 (2015).
Burikhanov, R. et al. Paracrine apoptotic effect of p53 mediated by tumor suppressor Par-4. Cell Rep. 6, 271–277 (2014).
Yamada, K. et al. Identification of a novel autoantigen UACA in patients with panuveitis. Biochem. Biophys. Res. Commun. 280, 1169–1176 (2001).
Lavrov, A. V. et al. Frequent variations in cancer-related genes may play prognostic role in treatment of patients with chronic myeloid leukemia. BMC Genet. 17, 14 (2016).
Yin, Y., Arkhipov, A. & Schulten, K. Simulations of membrane tubulation by lattices of amphiphysin N-BAR domains. Structure 17, 882–892 (2009).
Sorre, B. et al. Nature of curvature coupling of amphiphysin with membranes depends on its bound density. Proc. Natl Acad. Sci. USA 109, 173–178 (2012).
Vural, B. et al. Seroreactivity against PTEN-induced putative kinase 1 (PINK1) in Turkish patients with Behçet’s disease. Clin. Exp. Rheumatol. 27, S67–S72 (2009).
De Camilli, P. et al. The synaptic vesicle-associated protein amphiphysin is the 128-kD autoantigen of stiff-man syndrome with breast cancer. J. Exp. Med. 178, 2219–2223 (1993).
Folli, F. et al. Autoantibodies to a 128-kd synaptic protein in three women with the stiff-man syndrome and breast cancer. N. Engl. J. Med. 328, 546–551 (1993).
Ahuja, R. et al. Cordon-bleu is an actin nucleation factor and controls neuronal morphology. Cell 131, 337–350 (2007).
Park, W. S. et al. Comprehensive identification of PIP3-regulated PH domains from C. elegans to H. sapiens by model prediction and live imaging. Mol Cell 30, 381–392 (2008).
Hou, W. et al. The actin nucleator Cobl is controlled by calcium and calmodulin. PLoS Biol. 13, e1002233 (2015).
Qualmann, B. & Kelly, R. B. Syndapin isoforms participate in receptor-mediated endocytosis and actin organization. J. Cell Biol. 148, 1047–1062 (2000).
Schwintzer, L. et al. The functions of the actin nucleator cobl in cellular morphogenesis critically depend on syndapin I. EMBO J. 30, 3147–3159 (2011).
Koch, D. et al. Proper synaptic vesicle formation and neuronal network activity critically rely on syndapin I. EMBO J. 30, 4955–4969 (2011).
Zobel, T. et al. Cooperative functions of the two F-BAR proteins Cip4 and Nostrin in the regulation of E-cadherin in epithelial morphogenesis. J. Cell Sci. 128, 499–515 (2015).
Izadi, M. et al. Cobl-like promotes actin filament formation and dendritic branching using only a single WH2 domain. J. Cell Biol. 217, 211–230 (2018).
Beetz, C. et al. A spastic paraplegia mouse model reveals REEP1-dependent ER shaping. J. Clin. Invest. 123, 4273–4282 (2013).
Qualmann, B., Boeckers, T. M., Jeromin, M., Gundelfinger, E. D. & Kessels, M. M. Linkage of the actin cytoskeleton to the postsynaptic density via direct interactions of Abp1 with the ProSAP/Shank family. J. Neurosci. 24, 2481–2495 (2004).
Pinyol, R., Haeckel, A., Ritter, A., Qualmann, B. & Kessels, M. M. Regulation of N-WASP and the Arp2/3 complex by Abp1 controls neuronal morphology. PLoS One 2, e400 (2007).
Sholl, D. A. Dendritic organization in the neurons of the visual and motor cortices of the cat. J. Anat. 87, 387–406 (1953).
Drozdetskiy, A., Cole, C., Procter, J. & Barton, G. J. JPred4: a protein secondary structure prediction server. Nucleic Acids Res. 43, W389–W394 (2015).
Chen, C. C., Hwang, J. K. & Yang, J. M. (PS)2-v2: template-based protein structure prediction server. BMC Bioinformatics 10, 366 (2009).
Lupas, A., Van Dyke, M. & Stock, J. Predicting coiled coils from protein sequences. Science 252, 1162–1164 (1991).
McDonnell, A. V., Jiang, T., Keating, A. E. & Berger, B. Paircoil2: improved prediction of coiled coils from sequence. Bioinformatics 22, 356–358 (2006).
Wolf, E., Kim, P. S. & Berger, B. MultiCoil: a program for predicting two- and three-stranded coiled coils. Protein Sci. 6, 1179–1189 (1997).
Dereeper, A. et al. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 36, W465–W469 (2008).
Acknowledgements
We thank B. Schade and A. Kreusch for technical assistance. This work was supported by DFG grants KE685/3-2 and KE685/4-2 to M.M.K. as, well as QU116/5-2 and QU116/6-2 to B.Q.
Author information
Authors and Affiliations
Contributions
D.W., S.A.H.-M., M.I. and E.S. designed and performed the experiments and interpreted the data. F.S. helped with the cryoEM image recordings. D.K. provided endophilinA2N-BAR and advice. L.S. made initial observations and generated plasmids important for the study. M.M.K. and B.Q. conceived the project, designed the experiments, interpreted the data and co-wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Ankycorbin is a seven ankyrin repeat-containing protein predominantly expressed in early neuromorphogenesis.
(a) Ankyrin repeat consensus according to10,8,S1,S2. (b) Overlap of seven ankyrin repeats of mouse ankycorbin with the Mosavi consensus8. Residues of high and of medium conservation in the Mosavi consensus are highlighted in red and green, respectively, in the seven ankyrin repeats of ankycorbin as well. (c,d) Immunoblotting analyses of 150 µg mouse brain homogenate per lane revealing that ankycorbin expression decreases from E16 to 8W. The representative data shown in c represents examples of the quantitative analysis shown in d. Data, mean±SEM of n=3 biologically independent samples as indicated in the figure (d, n represents number of biologically independent brain lysates). Bar plots overlaid with dot plots (d). Statistical analyses, one-way-ANOVA/Tukey’s. (e) Immunoblot of cortical mouse neurons cultured from DIV0 to DIV14. Anti-GAPDH and anti-laminB1 signals are shown for comparison of equal loadings (representative data reflecting two independent experiments yielding similar results). For numerical source data see Supplementary Table 3; for raw image data see Supplementary Fig. 9.
Supplementary Figure 2 Repetition of the ankycorbin phenotypical study as fully blinded study done by a naïve, previously not involved, independent experimenter confirms the identified role of ankycorbin in dendritic arborisation of developing hippocampal neurons.
(a-c) Quantitative analyses of a set of experiments independent of those shown in Fig. 1m–q conducted as fully blinded study by an experimenter not previously involved in the project. DIV4 primary hippocampal neurons were transfected as indicated, fixed at DIV6, anti-MAP2 immunostained, systematically imaged and morphometrically analysed based on the MAP2 immunostaining using Imaris software. Ankycorbin-GFP expression resulted in an increase of intersections with the Sholl circles in the dendritic arbor (a) as well as in an increase of branch points (b) and terminal points compared to GFP-expressing cells (c). A statistically highly significant increase of dendritic branching in ankycorbin-GFP overexpressing neurons when compared to control neurons was observed. Data, mean±SEM of biologically independent samples with n numbers as indicated in the figure (a-c, ankycorbin-GFP, n=31; GFP, n=29); a-c, n represents number of biologically independent neurons). Statistical analyses, two-way-ANOVA/Bonferroni’s (a), two-tailed t-test (b,c). *P<0.05; **P<0.01; ***P<0.001. For numerical source data see Supplementary Table 3.
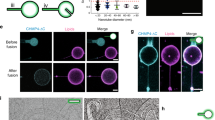
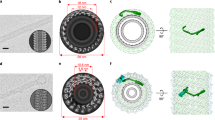
Supplementary Figure 3 The N terminal part of ankycorbin shapes membranes in in vitro-reconstitutions whereas other membrane-binding modules, such as the cytohesin 3/ARNO3 PH domain, do not.
(a-c) CryoEM analyses of in vitro-reconstitutions of liposomes without (a) and with ankycorbin1-252 (b) and quantitative determinations of liposome diameters of the same liposome preparation with and without incubation with ankycorbin1-252 (c). Bars, 100 nm. The representative data shown (a,b) represents examples of the quantitative analysis shown in c, n=674 (no protein control); n=531 (ankycorbin1-252) biologically independent liposomes (from two independent experiments). (d) Dose-dependence of ankycorbin1-252-mediated membrane shaping unveiled by quantitative determinations of liposome diameters of liposomes without ankycorbin1-252 added and upon incubations with ankycorbin1-252 at different protein/lipid ratios, n=600 (protein:lipids 0:1); n=1207 (protiein:lipids 0.05:1); n=601 (protein:lipids 0.15:1) (from two independent experiments visualized by freeze-fracturing and TEM). (e-j) Comparison of visual examination and average diameter determinations of liposomes sized by different extrusion cut-offs by TEM of freeze-fractured samples (e-i) and by photometric analyses of turbidity (j). Bars, 500 nm. (k) SDS-PAGE analyses of the binding of the cytohesin3/ARNO3 PH domain to both LUVs and SUVs. Note that the PH domain binds liposomes well irrespective of their size (that is membrane curvature) (representative data reflecting two independent experiments yielding similar results). (l) Photometric analyses of a putative membrane shaping of the cytohesin3/ARNO3 PH domain in comparison to GST serving as unrelated, not membrane-binding protein control. (m) Quantitative assessment of membrane rearrangement effects detectable by changes in light scattering caused by cytohesin3/ARNO3 PH domain in comparison to the control protein GST via integration over both averaged curves shown in l. l,m, n=4 independent biochemical assays each. Note that no changes occur when liposomes are incubated with GST or cytohesin3/ARNO3 PH domain (two-tailed t-test, n.s.). Data, mean±SEM of biologically independent samples with n numbers as indicated in the figure (c,d,i, n represents number of biologically independent liposomes; l,m, n represents number of independent biochemical assays). Bar plots overlaid with dot plots (m). Statistical analyses, two-tailed t-test (c) and one-way-ANOVA/Tukey’s (d), respectively. **P<0.1; ****P<0.0001. For P<0.0001, no P values are available. For other individual P values see figure. For numerical source data see Supplementary Table 3; for raw image data see Supplementary Fig. 9.
Supplementary Figure 4 Validation of ankycorbin RNAi.
(a) Immunoblots of Hepa1-6 cell lysates obtained 48 h post transfection with either scrambled RNAi/GFP (control) or ankycorbin RNAi/GFP showing that ankycorbin endogenously expressed in Hepa1-6 cells is efficiently knocked down. The representative data shown (a) represents examples of the quantitative analysis shown in b. (b) Quantitative RNAi validation depicting that ankycorbin RNAi significantly reduces ankycorbin protein levels. (b) n=5 independent experiments each. (c,d) Representative MIPs of anti-ankycorbin signals in Hepa1-6 cells 48 h post transfection (two independent experiments yielding similar results). Transfected cells are marked by expression of the marker GFP (asterisks). Please note the reduced anti-ankycorbin immunosignals in RNAi cells (d) in comparison to untransfected control cells, whereas scrambled RNAi does not show such an effect. (e) Quantitative analysis of ankycorbin RNAi efficiency comparing the anti-ankycorbin immunofluorescence signal of transfected cells to adjacent untransfected cells. Images were acquired using a constant exposure time. n=72 (scrambled RNAi/GFP); n=71 (ankycorbin RNAi/GFP) ratios of cell pairs. Bars, 10 µm. Data, mean±SEM of biologically independent samples with n numbers as indicated in the figure (b, n represents number of biologically independent assays; e, n represents number of biologically independent, transfected cell/neighbouring untransfected cell anti-ankycorbin immunosignal intensity ratios); bar plots with overlaid dot plots of all individual points of measurement. Two-sided Mann-Whitney-U. **P<0.01;****P<0.0001. For P<0.0001, no P values are available. For other individual P values see figure. For numerical source data see Supplementary Table 3; for raw image data see Supplementary Fig. 9.
Supplementary Figure 5 Ankycorbin’s N-terminal amphipathic helix is critically involved in membrane targeting in cells.
(a-e) MIPs of HeLa cells (24 h post transfection) coexpressing GFP (a), ankycorbin1-252-GFP (b), ankycorbin1-252 K2A,K5A,K7A,R9A,K10A-GFP (c), ankycorbin1-252 L4A,F8A-GFP (d) and ankycorbin17-252-GFP (e), respectively, with the plasma membrane marker cherryF. Note that only wild-type ankycorbin1-252 colocalises with cherryF, whereas the negative control (GFP) and all three ankycorbin mutants do not. The representative data shown reflects seven independent experiments (a,b) and three independent experiments (c,d,e), respectively, yielding similar results. Bars, 10 µm.
Supplementary Figure 6 Ankycorbin-mediated dendrite formation relies on its N-terminal amphipathic helix.
(a-e) MIPs of anti-MAP2-stained DIV4+2 hippocampal neurons co-expressing the membrane marker cherryF and GFP (a), ankycorbin-GFP (b) as well as an ankycorbin mutant lacking the amphipathic helix (ankycorbinΔAH-GFP) (c) and two different amphipathic helix mutants, respectively (d.e). The representative data shown (a-e) represents examples of the quantitative analysis shown in f-h (images of cells were obtained from two independent experiments with several independent coverslips). Transfected neurons are marked by red asterisks. Bars, 10 µm. (f-h) Quantitative analyses of the dendritic arbor of DIV4+2-transfected neurons. Data, mean±SEM of biologically independent samples with n numbers as indicated in the figure (f-h, n represents number of biologically independent, transfected neurons), n=30 (GFP), (ankycorbin-GFP) and (ankycorbinK2A,K5A,K7A,R9A,K10A); n=26 (ankycorbin∆AH-GFP); n=28 (ankycorbinL4A,F8A-GFP). Two-way-ANOVA/Bonferroni’s (f). One-way-ANOVA/Tukey’s (g,h). *P<0.05; **P<0.001. ***P<0.001. ****P<0.0001. For P<0.0001, no P values are available. For other individual P values see figure. For numerical source data see Supplementary Table 3.
Supplementary Figure 7 TEM characterisation of liposomes converted into small sizes by sonification.
(a-c) Visual examination of sonicated (SUV) versus untreated liposomes (LUV) by TEM of freeze-fractured samples (a,b) and determinations of their average diameters (n=208 SUVs and n=213 LUVs) (c). Some examples of SUVs are marked by arrows. Boxed area in (a) is shown as enlargement beneath. Bars, 500 nm. Data, mean±SEM of biologically independent samples with n numbers as also indicated in the figure (c, n represents number of biologically independent liposomes). Statistical analyses, two-tailed Student’s t-test. ****P<0.0001. For P<0.0001, no P values are available. For numerical source data see Supplementary Table 3.
Supplementary Figure 8 The ankycorbin N-Ank module alone is incapable to induce dendritic branching and proteins of the N-Ank family contain an amphipathic helix, five to seven ankyrin repeats and coiled coil domains, which mediate self-assembly.
(a-e) Fully blinded quantitative analyses of the potential of the ankycorbin N-Ank module in comparison to full-length ankycorbin to induce dendritic branching. DIV4 primary hippocampal neurons were transfected as indicated, fixed at DIV6, anti-MAP2 immunostained, systematically imaged and morphometrically analysed based on the MAP2 immunostaining using Imaris software (a-c). The representative data shown (a-c) represents examples of the quantitative analysis shown in d,e (the images of cells were obtained from two independent experiments with several independent coverslips). (d,e) Quantitative determination of branch point (d) and terminal point numbers (e) using Imaris software. Note that ankycorbin1-252 fails to phenocopy the full-length ankycorbin effects on dendritic branching. Data, mean±SEM of biologically independent samples with n numbers as indicated in the figure (d,e, n represents number of biologically independent, transfected neurons), n=29 (GFP); n=31 (ankycorbin-GFP); n=30 (ankycorbin1-252-GFP) neurons. Data for GFP and ankycorbin-GFP represent data shown in Supplementary Fig. 2. (GFP, n=29; ankycorbin-GFP, n=31; ankycorbin1-252-GFP, n=30 cells). Statistical analyses, One-way-ANOVA/Tukey’s (d,e). *P<0.05; ***P<0.001. (f) Domain organisation of 11 different N-Ank proteins (drawn to scale; ankycorbin (Q9EP71); UACA (Q8CGB3), Ankrd24 (Q80VM7), Ankrd35 (E9Q9D8), Ankrd30A (Q9BXX3), Ankrd26 (Q811D2), Ankrd20A1 (Q5TYW2), Ankrd18A (Q8IVF6), Ankrd18B (A2A2Z9), Ankrd62 (A6NC57), Ankrd36 (Q5QNR3)). Note that all of these proteins seem to occur in a variety of isoforms that are not depicted. Amphipathic helix in blue, ankyrin repeats (5–7 according to Uniprot; note that for some of these proteins more repeats may be present according to consensus similarities) in different shades of yellow and coiled coil domains in green (prediction according to Uniprot). Note that Ankrd7 (see Fig. 6a; not shown here) may lack a coiled coil domain. The four members of the N-Ank family analysed for their membrane binding functions in this study (that is ankycorbin, UACA, Ankrd35 and Ankrd20A1) are highlighted by coloured background. (g-i) Immunoblotting analyses showing EDC-cross-linked complexes of GFP-ankycorbin and of GFP-ankycorbin400-979 but not of GFP-ankycorbinΔCC (GFP-ankycorbin1-400) expressed in HEK293 cells (g and h represent the full blot images of the details shown in Fig. 8c,d). The representative data shown reflects two independent experiments yielding similar results. For numerical source data see Supplementary Table 3. For raw image data see Supplementary Fig. 9.
Supplementary Figure 9 Multi-page supplementary compilation of unprocessed gel and Western blot images (Continued).
Gels and blots appear in the order as in the manuscript and are marked for the corresponding figure panel. Coomassie-stained gels were converted into black and white images. Image parts used for the respective figure in the main manuscript or supplementary figures are boxed. The compilation contains uncropped gel images and Western blots images recorded using LiCOR (infrared fluorescence-based immunoblot detection). As in the figures, all blot images are shown as inverted images. Size standards recorded in the second infrared channel accompanying the immunoblotting analyses shown here are usually not shown. The shown raw image data are focused on the detected grey values in the grey value distributions to visualise signals but have not been processed further. Note that therefore the raw data signals shown here usually appear with low intensity and need to be processed further for full visualisation of all fluorescence signals.
Supplementary information
Supplementary Information
Supplementary Figures 1–9 and Supplementary table titles and legends
Supplementary Table 1
Overview and sequence information on all primers used in this study.
Supplementary Table 2
Overview and information on primary and secondary antibodies used in this study.
Supplementary Table 3
Numerical source data used for the quantitative analyses in this study.
Rights and permissions
About this article
Cite this article
Wolf, D., Hofbrucker-MacKenzie, S.A., Izadi, M. et al. Ankyrin repeat-containing N-Ank proteins shape cellular membranes. Nat Cell Biol 21, 1191–1205 (2019). https://doi.org/10.1038/s41556-019-0381-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0381-7
This article is cited by
-
Long-term depression in neurons involves temporal and ultra-structural dynamics of phosphatidylinositol-4,5-bisphosphate relying on PIP5K, PTEN and PLC
Communications Biology (2023)
-
Ubiquitination regulates ER-phagy and remodelling of endoplasmic reticulum
Nature (2023)
-
Heteromeric clusters of ubiquitinated ER-shaping proteins drive ER-phagy
Nature (2023)
-
Reduced Mrp2 surface availability as PI3Kγ-mediated hepatocytic dysfunction reflecting a hallmark of cholestasis in sepsis
Scientific Reports (2020)