Abstract
Tumour-derived microvesicles (TMVs) comprise a class of extracellular vesicles released from tumour cells that are now understood to facilitate communication between the tumour and the surrounding microenvironment. Despite their significance, the regulatory mechanisms governing the trafficking of bioactive cargos to TMVs at the cell surface remain poorly defined. Here we describe a molecular pathway for the delivery of microRNA (miRNA) cargo to nascent TMVs involving the dissociation of a pre-miRNA/Exportin-5 complex from Ran–GTP following nuclear export and its subsequent transfer to a cytoplasmic shuttle comprised of ARF6–GTP and GRP1. As such, ARF6 activation increases the pre-miRNA cargo contained within TMVs through a process that requires the casein kinase 2-mediated phosphorylation of RanGAP1. Furthermore, TMVs were found to contain pre-miRNA processing machinery including Dicer and Argonaute-2, which allow for cell-free pre-miRNA processing within shed vesicles. These findings offer cellular targets to block the loading and processing of pre-miRNAs within TMVs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Deep-sequencing (miRNA–seq) data that support the findings of this study have been deposited in the Gene Expression Omnibus under the accession code GSE130316. Previously published protein–protein interaction data that was analysed for this manuscript is available through the Intact database (www.ebi.ac.uk/intact) accession no. EBI-105937047, pubid:17353931 (ref. 45; Figs. 2b and 5e). Data used for the PRISM predictions was accessed through http://cosbi.ku.edu.tr/prism/ using structures stored in the RCSB Protein Data Bank (https://www.rcsb.org/; Figs. 2a, 5d and Supplementary Fig. 3a). The Cancer Genome Atlas data analysed in this manuscript was accessed via the Xena Functional Genomics Explorer52 (www.xenabrowser.net; Supplementary Figs. 4e,f and 5). All other data generated for and analysed in this manuscript is available by contacting the corresponding author. The source data for Figs. 1–6 and Supplementary Figs. 2–4,6 have been provided as Supplementary Table 1. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Abels, E. R. & Breakefield, X. O. Introduction to extracellular vesicles: biogenesis, RNA cargo selection, content, release, and uptake. Cell. Mol. Neurobiol. 36, 301–312 (2016).
Desrochers, L. M., Antonyak, M. A. & Cerione, R. A. Extracellular vesicles: satellites of information transfer in cancer and stem cell biology. Dev. Cell 37, 301–309 (2016).
Tricarico, C., Clancy, J. & D’Souza-Schorey, C. Biology and biogenesis of shed microvesicles. Small GTPases 8, 220–232 (2016).
van Niel, G., D’Angelo, G. & Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 19, 213–228 (2018).
Clancy, J. W. et al. Regulated delivery of molecular cargo to invasive tumour-derived microvesicles. Nat. Commun. 6, 6919 (2015).
Sedgwick, A. E., Clancy, J. W., Olivia Balmert, M. & D’Souza-Schorey, C. Extracellular microvesicles and invadopodia mediate non-overlapping modes of tumor cell invasion. Sci. Rep. 5, 14748 (2015).
Lazaro-Ibanez, E. et al. Metastatic state of parent cells influences the uptake and functionality of prostate cancer cell-derived extracellular vesicles. J. Extracell. Vesicles 6, 1354645 (2017).
Muralidharan-Chari, V. et al. ARF6-regulated shedding of tumor cell-derived plasma membrane microvesicles. Curr. Biol. 19, 1875–1885 (2009).
D’Souza-Schorey, C. & Clancy, J. W. Tumor-derived microvesicles: shedding light on novel microenvironment modulators and prospective cancer biomarkers. Genes Dev. 26, 1287–1299 (2012).
Wang, T. et al. Hypoxia-inducible factors and RAB22A mediate formation of microvesicles that stimulate breast cancer invasion and metastasis. Proc. Natl Acad. Sci. USA 111, E3234–E3242 (2014).
Nabhan, J. F., Hu, R., Oh, R. S., Cohen, S. N. & Lu, Q. Formation and release of arrestin domain-containing protein 1-mediated microvesicles (ARMMs) at plasma membrane by recruitment of TSG101 protein. Proc. Natl Acad. Sci. USA 109, 4146–4151 (2012).
Wang, Q. & Lu, Q. Plasma membrane-derived extracellular microvesicles mediate non-canonical intercellular NOTCH signaling. Nat. Commun. 8, 709 (2017).
Kuo, L. & Freed, E. O. ARRDC1 as a mediator of microvesicle budding. Proc. Natl Acad. Sci. USA 109, 4025–4026 (2012).
Melo, S. A. et al. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell 26, 707–721 (2014).
Menck, K. et al. Tumor-derived microvesicles mediate human breast cancer invasion through differentially glycosylated EMMPRIN. J. Mol. Cell Biol. 7, 143–153 (2015).
Skog, J. et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 10, 1470–1476 (2008).
Sansone, P. et al. Evolution of cancer stem-like cells in endocrine-resistant metastatic breast cancers is mediated by stromal microvesicles. Cancer Res. 77, 1927–1941 (2017).
Al-Nedawi, K., Meehan, B., Kerbel, R. S., Allison, A. C. & Rak, J. Endothelial expression of autocrine VEGF upon the uptake of tumor-derived microvesicles containing oncogenic EGFR. Proc. Natl Acad. Sci. USA 106, 3794–3799 (2009).
Al-Nedawi, K. et al. Intercellular transfer of the oncogenic receptor EGFRvIII by microvesicles derived from tumour cells. Nat. Cell Biol. 10, 619–624 (2008).
Calin, G. A. & Croce, C. M. MicroRNA signatures in human cancers. Nat. Rev. Cancer 6, 857–866 (2006).
Giovannetti, E. et al. MicroRNA-21 in pancreatic cancer: correlation with clinical outcome and pharmacologic aspects underlying its role in the modulation of gemcitabine activity. Cancer Res. 70, 4528–4538 (2010).
Melo, S. A. & Esteller, M. Disruption of microRNA nuclear transport in human cancer. Semin. Cancer Biol. 27, 46–51 (2014).
Jiang, Q. et al. MicroRNA-100 suppresses the migration and invasion of breast cancer cells by targeting FZD-8 and inhibiting Wnt/β-catenin signaling pathway. Tumour Biol. 37, 5001–5011 (2016).
Song, B. et al. MicroRNA-21 regulates breast cancer invasion partly by targeting tissue inhibitor of metalloproteinase 3 expression. J. Exp. Clin. Cancer Res. 29, 29 (2010).
Bohnsack, M. T., Czaplinski, K. & Gorlich, D. Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA 10, 185–191 (2004).
Yi, R., Qin, Y., Macara, I. G. & Cullen, B. R. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev. 17, 3011–3016 (2003).
Slezak-Prochazka, I., Durmus, S., Kroesen, B. J. & van den Berg, A. MicroRNAs, macrocontrol: regulation of miRNA processing. RNA 16, 1087–1095 (2010).
Wei, Z. et al. Coding and noncoding landscape of extracellular RNA released by human glioma stem cells. Nat. Commun. 8, 1145 (2017).
Shurtleff, M. J., Temoche-Diaz, M. M., Karfilis, K. V., Ri, S. & Schekman, R. Y-box protein 1 is required to sort microRNAs into exosomes in cells and in a cell-free reaction. eLife 5, e19276 (2016).
Villarroya-Beltri, C. et al. Sumoylated hnRNPA2B1 controls the sorting of miRNAs into exosomes through binding to specific motifs. Nat. Commun. 4, 2980 (2013).
Kosaka, N. et al. Neutral sphingomyelinase 2 (nSMase2)-dependent exosomal transfer of angiogenic microRNAs regulate cancer cell metastasis. J. Biol. Chem. 288, 10849–10859 (2013).
Kosaka, N. et al. Secretory mechanisms and intercellular transfer of microRNAs in living cells. J. Biol. Chem. 285, 17442–17452 (2010).
Guduric-Fuchs, J. et al. Selective extracellular vesicle-mediated export of an overlapping set of microRNAs from multiple cell types. BMC Genom. 13, 357 (2012).
Gibbings, D. J., Ciaudo, C., Erhardt, M. & Voinnet, O. Multivesicular bodies associate with components of miRNA effector complexes and modulate miRNA activity. Nat. Cell Biol. 11, 1143–1149 (2009).
McKenzie, A. J. et al. KRAS-MEK signaling controls Ago2 sorting into exosomes. Cell Rep. 15, 978–987 (2016).
Iorio, M. V. & Croce, C. M. MicroRNA dysregulation in cancer: diagnostics, monitoring and therapeutics. A comprehensive review. EMBO Mol. Med. 4, 143–159 (2012).
Croce, C. M. Causes and consequences of microRNA dysregulation in cancer. Nat. Rev. Genet. 10, 704–714 (2009).
Lin, S. & Gregory, R. I. MicroRNA biogenesis pathways in cancer. Nat. Rev. Cancer 15, 321–333 (2015).
Medina, P. P., Nolde, M. & Slack, F. J. OncomiR addiction in an in vivo model of microRNA-21-induced pre-B-cell lymphoma. Nature 467, 86–90 (2010).
Guttilla, I. K. & White, B. A. Coordinate regulation of FOXO1 by miR-27a, miR-96, and miR-182 in breast cancer cells. J. Biol. Chem. 284, 23204–23216 (2009).
Zheng, Y. S. et al. MiR-100 regulates cell differentiation and survival by targeting RBSP3, a phosphatase-like tumor suppressor in acute myeloid leukemia. Oncogene 31, 80–92 (2012).
Ding, J. et al. Gain of miR-151 on chromosome 8q24.3 facilitates tumour cell migration and spreading through downregulating RhoGDIA. Nat. Cell Biol. 12, 390–399 (2010).
Klein, S., Franco, M., Chardin, P. & Luton, F. Role of the Arf6 GDP/GTP cycle and Arf6 GTPase-activating proteins in actin remodeling and intracellular transport. J. Biol. Chem. 281, 12352–12361 (2006).
Brownawell, A. M. & Macara, I. G. Exportin-5, a novel karyopherin, mediates nuclear export of double-stranded RNA binding proteins. J. Cell Biol. 156, 53–64 (2002).
Ewing, R. M. et al. Large-scale mapping of human protein-protein interactions by mass spectrometry. Mol. Syst. Biol. 3, 89 (2007).
Keskin, O., Nussinov, R. & Gursoy, A. PRISM: protein-protein interaction prediction by structural matching. Methods Mol. Biol. 484, 505–521 (2008).
Stewart, M. Molecular mechanism of the nuclear protein import cycle. Nat. Rev. Mol. Cell Biol. 8, 195–208 (2007).
Pellon-Cardenas, O., Clancy, J., Uwimpuhwe, H. & D’Souza-Schorey, C. ARF6-regulated endocytosis of growth factor receptors links cadherin-based adhesion to canonical Wnt signaling in epithelia. Mol. Cell. Biol. 33, 2963–2975 (2013).
Takeda, E., Hieda, M., Katahira, J. & Yoneda, Y. Phosphorylation of RanGAP1 stabilizes its interaction with Ran and RanBP1. Cell Struct. Funct. 30, 69–80 (2005).
D’Souza-Schorey, C. & Chavrier, P. ARF proteins: roles in membrane traffic and beyond. Nat. Rev. Mol. Cell Biol. 7, 347–358 (2006).
Grossmann, A. H. et al. The small GTPase ARF6 stimulates β-catenin transcriptional activity during WNT5A-mediated melanoma invasion and metastasis. Sci. Signal. 6, ra14 (2013).
Goldman, M. et al. The UCSC Xena Platform for cancer genomics data visualization and interpretation. Preprint at bioRxiv https://doi.org/10.1101/326470 (2018).
Hongu, T. & Kanaho, Y. Activation machinery of the small GTPase Arf6. Adv. Biol. Regul. 54, 59–66 (2014).
Donaldson, J. G. & Jackson, C. L. ARF family G proteins and their regulators: roles in membrane transport, development and disease. Nat. Rev. Mol. Cell Biol. 12, 362–375 (2011).
Clancy, J. W., Tricarico, C. J., Marous, D. R. & D’Souza-Schorey, C. Coordinated regulation of intracellular fascin distribution governs tumor microvesicle release and invasive cell capacity. Mol. Cell. Biol. 39, e00264-18 (2019).
Martin del Campo, S. E. et al. MiR-21 enhances melanoma invasiveness via inhibition of tissue inhibitor of metalloproteinases 3 expression: in vivo effects of MiR-21 inhibitor. PLoS ONE 10, e0115919 (2015).
Ebert, M. S., Neilson, J. R. & Sharp, P. A. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nat. Methods 4, 721–726 (2007).
Li, Q. et al. MiR-21/Smad 7 signaling determines TGF-β1-induced CAF formation. Sci. Rep. 3, 2038 (2013).
Cohen, L. A. et al. Active Arf6 recruits ARNO/cytohesin GEFs to the PM by binding their PH domains. Mol. Biol. Cell. 18, 2244–2253 (2007).
Ritterhoff, T. et al. The RanBP2/RanGAP1*SUMO1/Ubc9 SUMO E3 ligase is a disassembly machine for Crm1-dependent nuclear export complexes. Nat. Commun. 7, 11482 (2016).
Reis-Sobreiro, M. et al. Emerin deregulation links nuclear shape instability to metastatic potential. Cancer Res. 78, 6086–6097 (2018).
Thind, A. & Wilson, C. Exosomal miRNAs as cancer biomarkers and therapeutic targets. J. Extracell. Vesicles 5, 31292 (2016).
Dror, S. et al. Melanoma miRNA trafficking controls tumour primary niche formation. Nat. Cell. Biol. 18, 1006–1017 (2016).
Sakha, S., Muramatsu, T., Ueda, K. & Inazawa, J. Exosomal microRNA miR-1246 induces cell motility and invasion through the regulation of DENND2D in oral squamous cell carcinoma. Sci. Rep. 6, 38750 (2016).
Zhang, L. et al. Microenvironment-induced PTEN loss by exosomal microRNA primes brain metastasis outgrowth. Nature 527, 100–104 (2015).
Schweitzer, J. K. & D’Souza-Schorey, C. Localization and activation of the ARF6 GTPase during cleavage furrow ingression and cytokinesis. J. Biol. Chem. 277, 27210–27216 (2002).
Hongu, T. et al. Arf6 regulates tumour angiogenesis and growth through HGF-induced endothelial beta1 integrin recycling. Nat. Commun. 6, 7925 (2015).
Jiang, J., Lee, E. J., Gusev, Y. & Schmittgen, T. D. Real-time expression profiling of microRNA precursors in human cancer cell lines. Nucleic Acids Res. 33, 5394–5403 (2005).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Acknowledgements
This work was supported in part by grants from the National Cancer Institute (grant no. R01CA115316) and the Catherine Peachey Foundation to C.D.-S.
Author information
Authors and Affiliations
Contributions
J.W.C. provided conceptual input, designed and performed experiments (shown in Figs. 1a-d; 2a,d,e,f,h; 3c,e; 4; 5; 6; and Supplementary Figs. S1a,e-g; S2a, c-g; S3b, d-i, k-n; S4; S5; S6a-c, e-i, k), analysed the data, proposed the model, assembled the figures and wrote the manuscript. Y.Z. provided conceptual input, designed and performed experiments (shown in Figs. 1e-l; 2a-e,i; 3a-e; 4a, 5; 6f; and Supplementary Figs. S1b,c; S2a-c; S3a,b, j; S6d, j), analysed the data and contributed to writing of the manuscript. C.S. designed and performed experiments (shown in Supplementary Figs. S1b; S6j) and assisted with experiments. C.D.-S. provided conceptual input, contributed to experimental design, analysed the data, wrote the manuscript and was responsible for the overall project administration.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 TMVs are distinct from other populations of extracellular vesicles.
a. 20 µg of total protein isolated from LOX cells, TMVs, or exosomes as outlined in methods, were separated by SDS-PAGE and probed, as indicated, by western blotting. Representative blots from N=3 biologically independent experiments shown. b. 2x106 LOX cells were plated and treated with vehicle control (water) or 50 nM okadaic acid for 24 hours to induce apoptosis. Apoptotic bodies and TMVs were fractionated according to methods, and the total secretome separated by SDS-PAGE and probed by western blotting. 5% of total cell lysate used as input control. Representative blots from N=4 biologically independent experiments shown. c. Bioanalyzer analysis for Small RNA TMV cargo from invasive breast cancer (MDA-MB-231) and prostate (PC-3) cells reveals the presence of miRNA cargo within shed TMVs. Representative images from N=3 biologically independent samples shown. Unprocessed blot images shown in Supplemental Image 7.
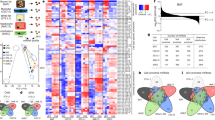
Supplementary Figure 2 ARF6 activation increases TMV shedding and miRNA content in shed TMVs.
a. Nanoparticle tracking analysis (NTA) and total particle concentration of TMVs released from equal numbers of LOX or LOXARF6-Q67L cells. Data represents mea±SEM for each diameter (NTA) or mean±SD (total particle concentration) for N=5 biologically independent samples. p-value determined by unpaired, two-tailed t-test. b. Heat-map analysis of the 50 most abundant TMV miRNAs from LOX or LOXARF6-Q67L cell lines reveals a consistent increase in TMV miRNA content with ARF6 activation. c. qRT–PCR analysis reveals an increase in miRNA cargo content in TMVs released by tumour cells of melanoma (LOX), ovarian (OvCar3), and breast (MDA-MB-231) origins. Data presented as mean±SD for N=3 biologically independent experiments. p-values determined by unpaired, two-tailed t-test between control and treatment conditions for each independent miRNA amplification reaction. p-values ≤0.05 were considered significant. d. qRT–PCR analysis of LOX cellular miRNA levels with expression of ARF6-Q67L. For each condition, data presented as mean±SD of N=3 biologically independent experiments. p-values determined by unpaired, two-tailed t-test between control and treatment conditions for each independent miRNA amplification reaction. p-values ≤0.05 were considered significant. e. ARF6 activity following expression of the fast-cycling ARF6-T157N mutant was measured using an MT2 ARF6–GTP specific pulldown as outlined in methods. Representative western blots and quantification for N=3 biologically independent samples shown. Data presented as mean±SD. p-value determined by unpaired, two-tailed t-test. f. NTA and total particle concentration of TMVs released from equal numbers of LOX or LOXARF6-T157N cells. Data represents mean±SEM for each diameter (NTA) or mean±SD (total particle concentration) for N=5 biologically independent samples. p-value determined by unpaired, two-tailed t-test. g. qRT–PCR analysis confirms an increase in pre-miRNA and miRNA cargo content in TMVs released by cells expressing ARF6-T157N. For each condition, data presented as mean±SD of N=3 biologically independent experiments. p-values determined by unpaired, two-tailed t-test for each independent miRNA amplification reaction. P-values ≤0.05 were considered significant. Unprocessed blot images shown in Supplemental Image 7. Statistical Source in Supplementary Table 1.
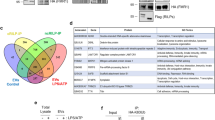
Supplementary Figure 3 Exportin-5 interacts with ARF6, affecting TMV cargo.
a. PRISM predicted interaction between Exportin-5 and ARF6–GDP. b. Myc immunoprecipitation from cells expressing ARF6-WT-HA and myc-Exportin-5. Representative blots (N=3 biologically independent experiments) shown. c. Myc-tag co-immunoprecipitation from cells expressing ARF6-T27N-HA or ARF6-WT-HA; and myc-Exportin-5. Representative blots (N=3 biologically independent experiments) shown. Endogenous Exportin-5 and ARF6 imaging in LOX cell TMVs (d) and perinuclear region (e). Panels d and e represent orthogonal view of Fig. 3h. Representative images (N=3 biologically independent experiments) shown. f. Immunofluorescence of endogenous Exportin-5 and ARF6 in isolated TMVs. Representative images shown (N=4 biologically independent samples). g. Western blot of Exportin-5 in TMVs from invasive tumour lines. Representative blots (N=4 biologically independent experiments) shown. h. Dicer and Argonaute-2 western blotting in control or LOXARF6-Q67L TMVs. Representative blots (N=3 biologically independent samples) shown. i. qRT–PCR of TMV miRNA cargo from control or TBB treated cells. Data presented as mean±SD of N=3 biologically independent experiments. p-values determined by unpaired, two-tailed t-test between control and treatment reactions for each miRNA. j. Western blot of Exportin-5 in 20 μg total protein from control or TBB treated LOX cells. Representative blots (N=3 biologically independent samples) shown. k. Western blot of equal amounts of nuclear and cytosolic fractions from control or TBB treated LOX cells. Representative blots (N=3 biologically independent samples) shown. Data presented as mean±SD. p-values determined by unpaired, two-tailed t-test between each control and treatment condition. l. Western blot of Exportin-5 in 20 μg total protein from control or TBB treated LOXARF6-Q67L cells. Representative blots (N=3 biologically independent samples) shown. m. Western blot of equal amounts of nuclear and cytosolic fractions from control or TBB treated LOXARF6-Q67L cells. Representative blots (N=3 biologically independent samples) shown. Data presented as mean±SD. p-values determined by unpaired, two-tailed t-test. n. Western blot of Ran and RanGAP1 in 20 µg total protein from LOX cells or TMVs. Representative blots (N=3 biologically independent experiments) shown. For all panels, P-values ≤0.05 were considered significant. Unprocessed blot images shown in Supplemental Image 7. Statistical Source in Supplementary Table 1.
Supplementary Figure 4 Cytohesin activity alters Exportin-5 localization and expression has varying effects on patient outcomes.
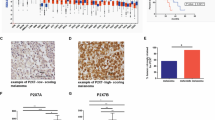
a. Control or SecinH3 treated LOX cells were fractionated as detailed in methods. Equal amounts of nuclear and cytosolic fractions were resolved by SDS-PAGE and probed as indicated by western blotting. Representative blots (N=3 biologically independent samples) shown. Data presented as mean±SD. p-values determined by unpaired, two-tailed t-test. b. Western blotting using 10 μg of total protein from cells treated with vehicle control or SecinH3 show no reduction in total levels of Exportin-5. Representative blots (N=3 biologically independent samples) shown. c. Similar to western blotting, qRT–PCR showed no change in Exportin-5 mRNA levels upon inhibition of the cytohesins by treatment with SecinH3. Data presented as mean±SD for N=3 biologically independent experiments. No significant relationship was found by one-way ANOVA with Tukey’s correction for multiple comparisons. d. LOX cells treated with SecinH3 alone or SecinH3 and Chloroquine were fractionated as outlined in methods. Cytosolic Exportin-5 levels were measured by western blotting of equal amounts of nuclear and cytosolic fractions. Representative blots (N=3 biologically independent samples) shown. Data presented as mean±SD. p-values determined by unpaired, two-tailed t-test. e. High levels of GRP1 expression correlate with poor overall survival. f. Cytohesin 1 and cytohesin 2 show an inverse correlation between expression levels and overall survival in pan-cancer analysis. Kaplan-Meier data reported with log-rank test statistic (χ2) and the p-value (χ2-distribution). In all panels, P-values ≤0.05 were considered significant. Unprocessed blot images shown in Supplemental Image 7. Statistical Source in Supplementary Table 1.
Supplementary Figure 5 GRP1 expression correlates with poor patient outcomes in multiple tumour types.
GRP1 expression data contained within the publicly available datasets comprising The Cancer Genome Atlas was analysed for correlation between GRP1 expression and patient survival. High levels of GRP1 (levels above the median values noted for each cancer type) expression correlate with poor outcomes in a. melanoma, b. ovarian, c. breast, d. lung squamous cell, and e. bladder cancers. Data reported with log-rank test statistic (χ2) and the p-value (χ2-distribution). For all panels, P-values ≤0.05 were considered significant.
Supplementary Figure 6 GRP1 is necessary for Exportin-5 and miRNA trafficking to TMVs.
a. Co-immunoprecipitation of GRP1. Representative blots (N=3 biologically independent experiments) shown. b. myc-Exportin-5 was precipitated from control or GRP1-shRNA transduced LOX cells. Blots are representative of N=3 biologically independent experiments. c. NTA and total particle concentration of TMVs released from equal numbers of LOX or LOXGRP1sh cells. Data represents mean±SEM for each diameter (NTA) or mean±SD (total particle concentration) for N=5 biologically independent samples. p-value determined by unpaired, two-tailed t-test. d. Western blotting of Exportin-5, Dicer, and Argonaute-2 in TMVs from LOXGRP1sh cells. Representative blots (N=3 biologically independent experiments). e. 15 μg of total protein from control or GRP1-shRNA cells; and control or GRP1-shRNA TMVs were separated by SDS-PAGE and protein cargo analysed by western blotting. Representative blots (N=3 biologically independent experiments) shown. f, g. Mature miRNA cargo is lost from shed TMVs upon the introduction of either of 2 independent shRNA sequences targeting GRP1. For each condition, data presented as mean±SD of N=3 biologically independent experiments. p-values determined by unpaired, two-tailed t-test between control and shRNA reactions for each independent miRNA amplification reaction. Cellular pre-miRNA (h) and mature miRNA (i) cargo is increased with depletion of GRP1. For each condition, data presented as mean±SD of N=3 biologically independent experiments. p-values determined by unpaired, two-tailed t-test between control and shRNA reactions for each independent miRNA amplification reaction. j. LOX cells were independently transduced with shRNA targeting the 3 cytohesin family members known to act on ARF6. Equal numbers of TMVs from control or shRNA cells were lysed and Exportin-5 cargo examined by western blotting. Representative blots (N=3 biologically independent experiments) shown. For all panels, P-values ≤0.05 were considered significant. k. Interaction between Exportin-5 and ARF6 is facilitated by the ARF GEF GRP1. Trafficking complex formation allows Exportin-5 and pre-miRNA cargo to be transferred from Ran–GTP to ARF6–GTP for outward trafficking and inclusion into nascent TMVs at peripheral sites of TMV biogenesis. Unprocessed blot images shown in Supplemental Image 7. Statistical Source in Supplementary Table 1.
Supplementary Figure 7 Unprocessed images of all gels and blots.
Unprocessed images of gels and blots presented in main and supplementary figures.
Supplementary information
Supplementary Information
Supplementary Figures 1–7 and Supplementary Table titles/legends.
Supplementary Table 1
Statistics source data.
Supplementary Table 2
Table of antibody information.
Rights and permissions
About this article
Cite this article
Clancy, J.W., Zhang, Y., Sheehan, C. et al. An ARF6–Exportin-5 axis delivers pre-miRNA cargo to tumour microvesicles. Nat Cell Biol 21, 856–866 (2019). https://doi.org/10.1038/s41556-019-0345-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0345-y
This article is cited by
-
Unravel the regulatory mechanism of Yrr1p phosphorylation in response to vanillin stress in Saccharomyces cerevisiae
Microbial Cell Factories (2023)
-
Nucleic acid drug vectors for diagnosis and treatment of brain diseases
Signal Transduction and Targeted Therapy (2023)
-
Application of the neuropeptide NPVF to enhance angiogenesis and osteogenesis in bone regeneration
Communications Biology (2023)
-
Context-specific regulation of extracellular vesicle biogenesis and cargo selection
Nature Reviews Molecular Cell Biology (2023)
-
Regulatory of miRNAs in tri-lineage differentiation of C3H10T1/2
Stem Cell Research & Therapy (2022)