Abstract
Wnt signalling stimulated by binding of R-spondin (Rspo) to Lgr-family members is crucial for gastrointestinal stem cell renewal. Infection of the stomach with Helicobacter pylori stimulates increased secretion of Rspo by myofibroblasts, leading to an increase in proliferation of Wnt-responsive Axin2+Lgr5− stem cells in the isthmus of the gastric gland and finally gastric gland hyperplasia. Basal Lgr5+ cells are also exposed to Rspo3, but their response remains unclear. Here, we demonstrate that—in contrast to its known mitogenic activity—Rspo3 induces differentiation of basal Lgr5+ cells into secretory cells that express and secrete antimicrobial factors, such as intelectin-1, into the lumen. The depletion of Lgr5+ cells or the knockout of Rspo3 in myofibroblasts leads to hypercolonization of the gastric glands with H. pylori, including the stem cell compartment. By contrast, systemic administration or overexpression of Rspo3 in the stroma clears H. pylori from the gastric glands. Thus, the Rspo3–Lgr5 axis simultaneously regulates both antimicrobial defence and mucosal regeneration.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The microarray data from this manuscript have been deposited in the GEO under accession code GSE108729. Single-cell RNA-seq data from this manuscript have been deposited in the GEO under accession code GSE124930. Bulk RNA-seq data are available from GEO under accession code GSE129107. Quantitative data that support the findings of this study are available within the paper and the Supplementary Information. Source data for graphical representations and statistical descriptions have been provided as Supplementary Table 6. All other data that support the findings of this study are available from the corresponding author on reasonable request.
Code availability
The computational code for GSEA analysis of microarray data of this manuscript can be accessed at https://github.com/MPIIB-Department-TFMeyer/Sigal_et_al_Rspo3_secretory_role. The code used for single cell RNA-seq analysis can be accessed at https://gist.github.com/cofischer/ab41169b5212046a59ba831e280835b3.
References
Parsonnet, J. et al. Helicobacter pylori infection and the risk of gastric carcinoma. N. Engl. J. Med. 325, 1127–1131 (1991).
Amieva, M. R. et al. Disruption of the epithelial apical-junctional complex by Helicobacter pylori CagA. Science 300, 1430–1434 (2003).
Toller, I. M. et al. Carcinogenic bacterial pathogen Helicobacter pylori triggers DNA double-strand breaks and a DNA damage response in its host cells. Proc. Natl Acad. Sci. USA 108, 14944–14949 (2011).
Sigal, M. et al. Helicobacter pylori activates and expands Lgr5(+) stem cells through direct colonization of the gastric glands. Gastroenterology 148, 1392–404.e21 (2015).
Sigal, M. et al. Stromal R-spondin orchestrates gastric epithelial stem cells and gland homeostasis. Nature 548, 451–455 (2017).
Munoz, J. et al. The Lgr5 intestinal stem cell signature: robust expression of proposed quiescent ‘+4’ cell markers. EMBO J. 31, 3079–3091 (2012).
Barker, N. et al. Lgr5+ve stem cells drive self-renewal in the stomach and build long-lived gastric units in vitro. Cell Stem Cell 6, 25–36 (2010).
Arnold, I. et al. Tolerance rather than immunity protects from Helicobacter pylori-induced gastric preneoplasia. Gastroenterology 140, 199–209 (2010).
Wunder, C. et al. Cholesterol glucosylation promotes immune evasion by Helicobacter pylori. Nat. Med. 12, 1030–1038 (2006).
Komiya, T., Tanigawa, Y. & Hirohashi, S. Cloning of the novel gene intelectin, which is expressed in intestinal Paneth cells in mice. Biochem. Biophys. Res. Commun. 251, 759–762 (1998).
Wesener, D. A. et al. Recognition of microbial glycans by human intelectin-1. Nat. Struct. Mol. Biol. 22, 603–610 (2015).
Powell, J. J., Greenfield, S. M. & Thompson, R. P. Concentrations of metals in gastric juice in health and peptic ulcer disease. Gut 33, 1617–1620 (1992).
Stange, D. E. et al. Differentiated Troy + chief cells act as reserve stem cells to generate all lineages of the stomach epithelium. Cell 155, 357–368 (2013).
Leushacke, M. et al. Lgr5-expressing chief cells drive epithelial regeneration and cancer in the oxyntic stomach. Nat. Cell Biol. 19, 774–786 (2017).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Farin, H. F. et al. Visualization of a short-range Wnt gradient in the intestinal stem-cell niche. Nature 530, 340–343 (2016).
Wehkamp, J. et al. Paneth cell antimicrobial peptides: topographical distribution and quantification in human gastrointestinal tissues. FEBS Lett. 580, 5344–5350 (2006).
Koslowski, M. J. et al. Association of a functional variant in the Wnt co-receptor LRP6 with early onset ileal Crohn’s disease. PLoS Genet. 8, e1002523 (2012).
Saenz, J. B., Vargas, N. & Mills, J. C. Tropism for spasmolytic polypeptide-expressing metaplasia allows Helicobacter pylori to expand its intragastric niche. Gastroenterology 156, 160–174 (2019).
Naik, S., Larsen, S. B., Cowley, C. J. & Fuchs, E. Two to tango: dialog between immunity and stem cells in health and disease. Cell 175, 908–920 (2018).
Bartfeld, S. et al. In vitro expansion of human gastric epithelial stem cells and their responses to bacterial infection.Gastroenterology 148, 126–136.e6 (2015).
Boccellato, F. et al. Polarised epithelial monolayers of the gastric mucosa reveal insights into mucosal homeostasis and defence against infection. Gut 68, 400–413 (2019).
Zimmermann, S. et al. ALPK1- and TIFA-dependent innate immune response triggered by the Helicobacter pylori type IV secretion system. Cell Rep. 20, 2384–2395 (2017).
Pfannkuch, L. et al. ADP heptose, a novel pathogen-associated molecular pattern identified in Helicobacter pylori. FASEB J. https://doi.org/10.1096/fj.201802555R (2019).
Arnold, I. C. et al. Tolerance rather than immunity protects from Helicobacter pylori-induced gastric preneoplasia. Gastroenterology 140, 199–209 (2011).
Mejias-Luque, R. et al. Lymphotoxin β receptor signalling executes Helicobacter pylori-driven gastric inflammation in a T4SS-dependent manner. Gut 66, 1369–1381 (2017).
Amieva, M. R. & El-Omar, E. M. Host-bacterial interactions in Helicobacter pylori infection. Gastroenterology 134, 306–323 (2008).
Tian, H. et al. A reserve stem cell population in small intestine renders Lgr5-positive cells dispensable. Nature 478, 255–259 (2011).
Neufeld, S. et al. A conditional allele of Rspo3 reveals redundant function of R-spondins during mouse limb development. Genesis 50, 741–749 (2012).
Herring, B. P., Hoggatt, A. M., Burlak, C. & Offermanns, S. Previously differentiated medial vascular smooth muscle cells contribute to neointima formation following vascular injury. Vasc. Cell 6, 21 (2014).
Hilkens, J. et al. RSPO3 expands intestinal stem cell and niche compartments and drives tumorigenesis. Gut 66, 1095–1105 (2017).
Dyer, V. et al. Genomic features of the Helicobacter pylori strain PMSS1 and its virulence attributes as deduced from its in vivo colonisation patterns. Mol. Microbiol. 110, 761–776 (2018).
Churchill, G. A. Fundamentals of experimental design for cDNA microarrays. Nat. Genet. 32, 490–495 (2002).
Yu, D., Huber, W. & Vitek, O. Shrinkage estimation of dispersion in negative binomial models for RNA-seq experiments with small sample size. Bioinformatics 29, 1275–1282 (2013).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Kamburov, A., Stelzl, U., Lehrach, H. & Herwig, R. The ConsensusPathDB interaction database: 2013 update. Nucleic Acids Res. 41, D793–D800 (2013).
Sergushichev, A. An algorithm for fast preranked gene set enrichment analysis using cumulative statistic calculation. Preprint at bioRxiv https://doi.org/10.1101/060012 (2016).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Backert, S. et al. Translocation of the Helicobacter pylori CagA protein in gastric epithelial cells by a type IV secretion apparatus. Cell. Microbiol. 2, 155–164 (2000).
Koch, M., Mollenkopf, H. J. & Meyer, T. F. Macrophages recognize the Helicobacter pylori type IV secretion system in the absence of toll-like receptor signalling. Cell. Microbiol. 18, 137–147 (2016).
Hatzios, S. K. et al. Chemoproteomic profiling of host and pathogen enzymes active in cholera. Nat. Chem. Biol. 12, 268–274 (2016).
Acknowledgements
We thank K. Hoffmann, S. Lehmann, Y. Giesecke, I. Wagner and C. Braeuning for technical support, V. Brinkmann for help with live imaging, A. Kühl and members of the Charité Core Facility for Histopathology for sample processing and histopathological scoring, and R. Zietlow for editing the manuscript. This work was performed in the Department of Molecular Biology of the MPIIB with extramural support from the DFG to M.S. (SI1983/2-1 and SI1983/3-1) and T.F.M. (ME705/18-1). M.S. received financial support from the George-Will-Foundation and is a participant of the BIH Charité Clinician Scientist Program funded by the Charité University Medicine Berlin and the Berlin Institute of Health.
Author information
Authors and Affiliations
Contributions
The study was designed by M.S. and supervised by T.F.M. M.S. and S.M. performed most of the in vivo experiments. M.E.R. and M.S. performed Lgr5 cell sorting and RNA bulk sequencing experiments. M.d.M.R. performed the functional experiments with Itln1. M.K., C.F. and S.S. performed single-cell RNA-seq experiments and data analysis. H.-J.M. performed microarray studies and H.B. and H.-J.M. analysed the data. E.R.M.B. and B.W. contributed materials required for the study. The manuscript was written by M.S. with guidance and discussion from T.F.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Fig. 1 R-spondin 3 affects Lgr5 signature and colonization with H. pylori.
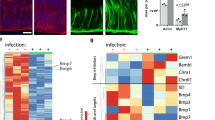
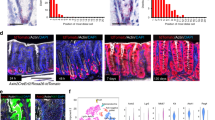
a) Single molecule RNA-in situ hybridization (red dots) for Rspo3 in corpus tissue. b) GSEA from microarray data comparing the expression profile of corpus tissue of Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice treated with tamoxifen 14 days before sacrifice to published intestinal Lgr5 signatures and TROY+ cell signature in the corpus. A permutation test was used. p-values from all gene sets were jointly adjusted for multiple testing using the method of Benjamini-Hochberg (FDR). c) qPCR analysis of Lgr5 expression normalized to Gapdh from Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice. d) H&E images from the corpus of Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice 2 months before sacrifice. e) Immunofluorescence images of stomach antrum tissue from Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice infected with H. pylori (green) 2 months before sacrifice. f) Expression levels of genes encoding for Reg proteins in the corpus of 2-month-infected Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice. Data from n = 2 independent biological samples per group. g) qPCR for Reg3g expression normalized to Gapdh in the corpus of uninfected and 2-month-infected Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice. Data are displayed as mean±SD from 3 biological replicates, analyzed by two-sided Student’s t-test. Images are representative of at least 3 independent biological samples. Scale bars: 100 μm. Microarray data are from 2 independent biological samples per group.
Supplementary Fig. 2 Bulk RNAseq from Lgr5+ cells.
a) Schematic visualization of a gland with Lgr5eGFP+ cells in the base b) Table of genes with highest FPKM values in uninfected Lgr5+ cells (data from n = 3 independent biological samples) c) FPKM values for EPCAM and Lgr5 in uninfected and infected samples. d) FPKM values of TFF2 as a marker of basal cells and Muc5ac as a marker of surface pit cells. e) Gating and sorting strategy of Lgr5+ EPCAM+ cells (note that all Lgr5+ cells are EPCAM+). Data are displayed as mean ± SD from 3 independent biological samples
Supplementary Fig. 3 Acidity is not affected by Rspo3.
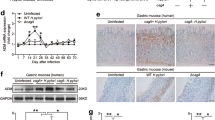
a) Immunofluorescence for gastrin in mouse antrum from uninfected and 2-month-infected Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice. b) Expression of gastrin from microarray data comparing the transcriptome of Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mouse antrum (mice were sacrificed 14 d post tamoxifen). c) Relative expression of gastrin mRNA normalized to Gapdh in the antrum of WT and Rspo3 KO mice either uninfected (n = 3 and n = 4) or infected with H. pylori for 2 months (n = 3 and n = 4). d) Relative expression of Atp4b mRNA normalized to Gapdh in the corpus of WT and Rspo3 KO mice either uninfected (n = 3 and n = 4) or infected with H. pylori for 2 months (n = 3 and n = 4). e) pH measured in the stomachs of WT and Rspo3 KO mice either uninfected (n = 4 and n = 5) or infected with H. pylori for 2 months (n = 5 and n = 4). Data are displayed as mean ± SD, analyzed by one-way ANOVA. Images are representative of at least 3 independent biological samples. Scale bars: 100 μm.
Supplementary Fig. 4 Expression of Lgr5 and intelectin is Rspo3-dependent, but overexpression is not observed early (2 weeks) after infection.
a) qPCR data from uninfected (n = 3) and 2 week-infected (n = 4) WT mice relative to Gapdh, normalized to uninfected control. b) qPCR data from 2 week-infected Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice (n = 4 per group) relative to Gapdh, normalized to Myh11CreErt2/Rspo3+/+. Data are displayed as mean ± SD, analyzed by two-sided Student’s t-test.
Supplementary Fig. 5 Functional T4SS and Rspo3 are required for antimicrobial effects but Rspo3 does not affect the inflammatory response.
a) Quantification of colony forming units from Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice infected with WT or ΔCagE H. pylori strain PMSS1 and treated with tamoxifen 2 months before sacrifice (for WT PMSS1 infection, data from Fig. 3a is reproduced here), each dot represents one mouse (n = 6,7,3 and 6 per group as indicated in the figure). b) qPCR for Lgr5 expression normalized to Gapdh from mouse antrum of Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl infected with WT and ΔCagE PMSS1 strain of H. pylori 2 months before sacrifice (n = 5,3,3,5 per group). c) qPCR for Itln1 expression normalized to Gapdh from antrum of Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice infected with WT and ΔCagE H. pylori strain PMSS1 2 months before sacrifice (n = 5,3,3,5 per respective group as indicated in the figure). d) Inflammation score for uninfected mice and Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice infected with WT and ΔCagE H. pylori strain PMSS1 2 months before sacrifice (n = 7,6,3,4 per respective group as indicated in the figure). e) Quantification of F4/80 immunostaining from Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl infected with WT and ΔCagE H. pylori strain PMSS1 2 months before sacrifice (data from 3 mice per group, comparison between Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl for each pair (uninfected, WT infected, ΔCagE infected) was done by two-sided t test). f) Quantification of CD3 immunostaining from Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice infected with WT and ΔCagE H. pylori strain PMSS1 2 months before sacrifice (data from 3 mice per group, comparison between Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl for each pair (uninfected, WT infected, ΔCagE infected) was done using two-sided t test). g) Representative images for F4/80 immunostaining from Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl infected with WT and ΔCagE H. pylori strain PMSS1 2 months before sacrifice. Images representative of 3 independent biological samples per group. h) Representative images for CD3 immunostaining from Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice infected with WT and ΔCagE H. pylori strain PMSS1 2 months before sacrifice. Images representative of 3 independent biological samples per group. i) Quantification of CD4+ T-bet+ cells measured in 10 high power fields per mouse from Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl infected with WT and ΔCagE H. pylori strain PMSS1 2 months before sacrifice (n = 4,3,4,5 per respective group), one way ANOVA. j) qPCR for interferon-γ and CXCL9 normalized to Gapdh from Myh11CreErt2/Rspo3+/+ and Myh11CreErt2/Rspo3fl/fl mice infected with WT H. pylori for 2 months, n = 4 per group, two-sided t-test. Scale bars: 100 µm.
Supplementary Fig. 6 Intelectin expression is Rspo-dependent in organoids and restricted to the gland base.
a-c) qPCR for Lgr5, Axin2 and Itln1 in primary 3D organoids derived from the mouse stomach antrum and grown in medium supplemented with different concentrations of Rspo1, normalized to Gapdh (0 and 50%: n = 2 independent biological replicates, 100%: n = 3 independent biological samples). d) smFISH for Itln1 in the antrum of an uninfected and a 2-month-infected mouse. Images are representative of 3 independent biological samples. Scale bars: 100 μm. e) Histogram displaying the position of Itln1+ cells in the antrum glands in 2-month-infected mice, data derived from n = 3 mice. f) Staining for Ki67, as well as high magnification image of co-staining for GSII and Ki67 to demonstrate that the gland base contains mainly GSII+ cells, whereas proliferation is mainly present in the cells above the base. Immunofluorescence for CD44 demonstrates expression in the gland base. Data are displayed as mean ± SD. Images are representative of at least 3 independent biological samples. Scale bars: 100 μm.
Supplementary Fig. 7 Intelectin 1 agglutinates H. pylori.
a) Survival curves: CFU data upon exposure of H. pylori cultures to Itln1, HBD3 and polymyxin B at different concentrations. Data derived from n = 3 independent biological samples. b) Agglutination assay: Crystal violet images of H. pylori cultures exposed to polymyxin B right: Quantification from n = 3 independent biological samples per group. c) Combined confocal microscopy/phase contrast images of H. pylori cultures either exposed to recombinant Itln1 (right) or the same volume of PBS. The cultures were recorded for a total of 52 minutes and images are from minute 26. The experiment was repeated twice in the lab. The area of GFP+ clusters that exceeded the size of single bacteria was measured every 60 seconds and plotted as GFP+ area over time. Red dots represent Itln1-treated cultures; black dots represent PBS-treated controls. d) Confocal microscopy images of primary bone marrow-derived macrophages that were infected with H. pylori in the presence or absence of recombinant Itln1. Middle: Inserts from the images at higher magnification. The GFP+ area per macrophage (n = 51 and n = 52 per group) was measured and two sided t-test was performed. Data are displayed as mean±SD. Scale bars: 10 µm.
Supplementary Fig. 8
Unprocessed blot of Fig. 7c
Supplementary Information
Supplementary Information
Supplementary Figs. 1–8
Supplementary Table 1
FPKM values from bulk sequenced Lgr5eGFP+ cells
Supplementary Table 2
Top 50 cluster specific genes for each cluster from scRNAseq data from LgreGFP+ cells of uninfected and H. pylori infected mice.
Supplementary Table 3
Gene sets of differentially expressed genes between cluster 1 and non-cluster 1 genes from data 18 of infected cells were subjected to over-representation analysis using the web-based tool 19 ConsensusPathDB-mouse.
Supplementary Table 4
Differential gene expression analysis from cluster 1 and cluster 2 from uninfected and infected cells using the merged dataset from single cells RNAsec (negative binomial exact test).
Supplementary Table 5
Summary of technical and statistical information from single cell RNAseq experiment.
Supplementary Table 6
Statistics source data
Supplementary Video 1
Time lapse movie (total of 50 minutes) displaying bacterial clustering upon exposure to intelectin 1.
Supplementary Video 2
Real-time visualization of bacterial motility from movie 1 to visualize altered bacterial motility upon intelectin 1 treatment.
Rights and permissions
About this article
Cite this article
Sigal, M., Reinés, M.d.M., Müllerke, S. et al. R-spondin-3 induces secretory, antimicrobial Lgr5+ cells in the stomach. Nat Cell Biol 21, 812–823 (2019). https://doi.org/10.1038/s41556-019-0339-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0339-9
This article is cited by
-
Role of PDGFRA+ cells and a CD55+ PDGFRALo fraction in the gastric mesenchymal niche
Nature Communications (2023)
-
Endothelial cell-derived RSPO3 activates Gαi1/3-Erk signaling and protects neurons from ischemia/reperfusion injury
Cell Death & Disease (2023)
-
Helicobacter pylori–activated fibroblasts as a silent partner in gastric cancer development
Cancer and Metastasis Reviews (2023)
-
The composition and roles of gastric stem cells in epithelial homeostasis, regeneration, and tumorigenesis
Cellular Oncology (2023)
-
Helicobacter pylori shows tropism to gastric differentiated pit cells dependent on urea chemotaxis
Nature Communications (2022)