Abstract
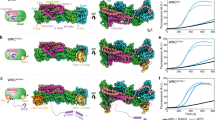
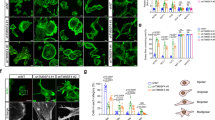
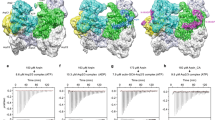
RPEL proteins, which contain the G-actin-binding RPEL motif, coordinate cytoskeletal processes with actin dynamics. We show that the ArhGAP12- and ArhGAP32-family GTPase-activating proteins (GAPs) are RPEL proteins. We determine the structure of the ArhGAP12/G-actin complex, and show that G-actin contacts the RPEL motif and GAP domain sequences. G-actin inhibits ArhGAP12 GAP activity, and this requires the G-actin contacts identified in the structure. In B16 melanoma cells, ArhGAP12 suppresses basal Rac and Cdc42 activity, F-actin assembly, invadopodia formation and experimental metastasis. In this setting, ArhGAP12 mutants defective for G-actin binding exhibit more effective downregulation of Rac GTP loading following HGF stimulation and enhanced inhibition of Rac-dependent processes, including invadopodia formation. Potentiation or disruption of the G-actin/ArhGAP12 interaction, by treatment with the actin-binding drugs latrunculin B or cytochalasin D, has corresponding effects on Rac GTP loading. The interaction of G-actin with RPEL-family rhoGAPs thus provides a negative feedback loop that couples Rac activity to actin dynamics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The ArhGAP12/G-actin structure has been deposited in the Protein Data Bank (PDB; https://www.rcsb.org) with the primary accession code 6GVC. Structures of MRTF-A RPEL2/G-actin and ArhGAP15 that were re-analysed in this study were obtained from PDB under the accession codes 2V52 and 3BYI, respectively.
Previously published RNA sequencing data that were re-analysed here are available under the accession code GSE45888.
The human melanoma survival data were derived from the TCGA Research Network (http://cancergenome.nih.gov). The dataset derived from this resource that supports the findings of this study is available in OncoLnc (http://www.oncolnc.org).
The source data for Figs. 1c, 2a,d, 3a,f, 4a–c, 5a–d, 6a–d, 7a–c,e,f,h and Supplementary Figs. 1e,f,i, 2f, 3c,d, 4b–d,f,–h, 5a–d,g–i, 6a–e have been provided as Supplementary Table 1. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Krause, M. & Gautreau, A. Steering cell migration: lamellipodium dynamics and the regulation of directional persistence. Nat. Rev. Mol. Cell Biol. 15, 577–590 (2014).
Przybyla, L., Muncie, J. M. & Weaver, V. M. Mechanical control of epithelial-to-mesenchymal transitions in development and cancer. Annu. Rev. Cell Dev. Biol. 32, 527–554 (2016).
Skau, C. T. & Waterman, C. M. Specification of architecture and function of actin structures by actin nucleation factors. Annu. Rev. Biophys. 44, 285–310 (2015).
Dominguez, R. & Holmes, K. C. Actin structure and function. Annu. Rev. Biophys. 40, 169–186 (2011).
Lawson, C. D. & Ridley, A. J. Rho GTPase signaling complexes in cell migration and invasion. J. Cell Biol. 217, 447–457 (2018).
Hodge, R. G. & Ridley, A. J. Regulating Rho GTPases and their regulators. Nat. Rev. Mol. Cell Biol. 17, 496–510 (2016).
Cook, D. R., Rossman, K. L. & Der, C. J. Rho guanine nucleotide exchange factors: regulators of Rho GTPase activity in development and disease. Oncogene 33, 4021–4035 (2014).
Laurin, M. & Cote, J. F. Insights into the biological functions of dock family guanine nucleotide exchange factors. Genes Dev. 28, 533–547 (2014).
Tcherkezian, J. & Lamarche-Vane, N. Current knowledge of the large RhoGAP family of proteins. Biol. Cell 99, 67–86 (2007).
Amin, E. et al. Deciphering the molecular and functional basis of RHOGAP family proteins: a systematic approach toward selective inactivation of Rho family proteins. J. Biol. Chem. 291, 20353–20371 (2016).
Miralles, F., Posern, G., Zaromytidou, A. I. & Treisman, R. Actin dynamics control SRF activity by regulation of its coactivator MAL. Cell 113, 329–342 (2003).
Vartiainen, M. K., Guettler, S., Larijani, B. & Treisman, R. Nuclear actin regulates dynamic subcellular localization and activity of the SRF cofactor MAL. Science 316, 1749–1752 (2007).
Mouilleron, S., Guettler, S., Langer, C. A., Treisman, R. & McDonald, N. Q. Molecular basis for G-actin binding to RPEL motifs from the serum response factor coactivator MAL. EMBO J. 27, 3198–3208 (2008).
Wiezlak, M. et al. G-actin regulates the shuttling and PP1 binding of the RPEL protein Phactr1 to control actomyosin assembly. J. Cell Sci. 125, 5860–5872 (2012).
Huet, G. et al. Actin-regulated feedback loop based on Phactr4, PP1 and cofilin maintains the actin monomer pool. J. Cell Sci. 126, 497–507 (2013).
Esnault, C. et al. Rho-actin signaling to the MRTF coactivators dominates the immediate transcriptional response to serum in fibroblasts. Genes Dev. 28, 943–958 (2014).
Cen, B. et al. Megakaryoblastic leukemia 1, a potent transcriptional coactivator for serum response factor (SRF), is required for serum induction of SRF target genes. Mol. Cell. Biol. 23, 6597–6608 (2003).
Allen, P. B., Greenfield, A. T., Svenningsson, P., Haspeslagh, D. C. & Greengard, P. Phactrs 1-4: a family of protein phosphatase 1 and actin regulatory proteins. Proc. Natl Acad. Sci. USA 101, 7187–7192 (2004).
Sagara, J. et al. Scapinin, a putative protein phosphatase-1 regulatory subunit associated with the nuclear nonchromatin structure. J. Biol. Chem. 278, 45611–45619 (2003).
Pawłowski, R., Eeva Kaisa Rajakylä, E. K., Vartiainen, M. K. & Treisman, R. An actin-regulated importin α/β-dependent extended bipartite NLS directs nuclear import of MRTF-A. EMBO J. 29, 3448–3458 (2010).
Hirano, H. & Matsuura, Y. Sensing actin dynamics: structural basis for G-actin-sensitive nuclear import of MAL. Biochem. Biophys. Res. Commun. 414, 373–378 (2011).
Furukawa, Y. et al. Isolation of a novel human gene, ARHGAP9, encoding a Rho-GTPase activating protein. Biochem. Biophys. Res. Commun. 284, 643–649 (2001).
Gentile, A. et al. Met-driven invasive growth involves transcriptional regulation of Arhgap12. Oncogene 27, 5590–5598 (2008).
Seoh, M. L., Ng, C. H., Yong, J., Lim, L. & Leung, T. ArhGAP15, a novel human RacGAP protein with GTPase binding property. FEBS Lett. 539, 131–137 (2003).
Zhao, C. et al. GC-GAP, a Rho family GTPase-activating protein that interacts with signaling adapters Gab1 and Gab2. J. Biol. Chem. 278, 34641–34653 (2003).
Matsuda, M. et al. Identification of adherens junction-associated GTPase activating proteins by the fluorescence localization-based expression cloning. Exp. Cell Res. 314, 939–949 (2008).
Monastyrskaya, K. et al. miR-199a-5p regulates urothelial permeability and may play a role in bladder pain syndrome. Am. J. Pathol. 182, 431–448 (2013).
Rudnicki, A. et al. Next-generation sequencing of small RNAs from inner ear sensory epithelium identifies microRNAs and defines regulatory pathways. BMC Genom. 15, 484 (2014).
Lecat, S., Matthes, H. W., Pepperkok, R., Simpson, J. C. & Galzi, J. L. A fluorescent live imaging screening assay based on translocation criteria identifies novel cytoplasmic proteins implicated in G protein-coupled receptor signaling pathways. Mol. Cell. Proteomics 14, 1385–1399 (2015).
Schlam, D. et al. Phosphoinositide 3-kinase enables phagocytosis of large particles by terminating actin assembly through Rac/Cdc42 GTPase-activating proteins. Nat. Commun. 6, 8623 (2015).
Ba, W. et al. ARHGAP12 functions as a developmental brake on excitatory synapse function. Cell Rep. 14, 1355–1368 (2016).
Nakamura, T. et al. PX-RICS mediates ER-to-Golgi transport of the N-cadherin/β-catenin complex. Genes Dev. 22, 1244–1256 (2008).
Mouilleron, S., Wiezlak, M., O’Reilly, N., Treisman, R. & McDonald, N. Q. Structures of the Phactr1 RPEL domain and RPEL motif complexes with G-actin reveal the molecular basis for actin binding cooperativity. Structure 20, 1960–1970 (2012).
Guettler, S., Vartiainen, M. K., Miralles, F., Larijani, B. & Treisman, R. RPEL motifs link the serum response factor cofactor MAL but not myocardin to Rho signaling via actin binding. Mol. Cell. Biol. 28, 732–742 (2008).
Posern, G., Sotiropoulos, A. & Treisman, R. Mutant actins demonstrate a role for unpolymerized actin in control of transcription by serum response factor. Mol. Biol. Cell 13, 4167–4178 (2002).
Mouilleron, S., Langer, C. A., Guettler, S., McDonald, N. Q. & Treisman, R. Structure of a pentavalent G-actin*MRTF-A complex reveals how G-actin controls nucleocytoplasmic shuttling of a transcriptional coactivator. Sci. Signal. 4, ra40 (2011).
Radu, M. et al. ArhGAP15, a Rac-specific GTPase-activating protein, plays a dual role in inhibiting small GTPase signaling. J. Biol. Chem. 288, 21117–21125 (2013).
Zamboni, V. et al. Disruption of ArhGAP15 results in hyperactive Rac1, affects the architecture and function of hippocampal inhibitory neurons and causes cognitive deficits. Sci. Rep. 6, 34877 (2016).
Graziano, B. R. et al. A module for Rac temporal signal integration revealed with optogenetics. J. Cell Biol. 216, 2515–2531 (2017).
Kurisu, S., Suetsugu, S., Yamazaki, D., Yamaguchi, H. & Takenawa, T. Rac-WAVE2 signaling is involved in the invasive and metastatic phenotypes of murine melanoma cells. Oncogene 24, 1309–1319 (2005).
Nakahara, H. et al. Involvement of Cdc42 and rac small G proteins in invadopodia formation of RPMI7951 cells. Genes Cells 8, 1019–1027 (2003).
Yamaguchi, H. et al. Sphingosine-1-phosphate receptor subtype-specific positive and negative regulation of Rac and haematogenous metastasis of melanoma cells. Biochem. J. 374, 715–722 (2003).
Stengel, K. & Zheng, Y. Cdc42 in oncogenic transformation, invasion, and tumorigenesis. Cell. Signal. 23, 1415–1423 (2011).
Revach, O. Y., Winograd-Katz, S. E., Samuels, Y. & Geiger, B. The involvement of mutant Rac1 in the formation of invadopodia in cultured melanoma cells. Exp. Cell Res. 343, 82–88 (2016).
Komatsu, N. et al. Development of an optimized backbone of FRET biosensors for kinases and GTPases. Mol. Biol. Cell 22, 4647–4656 (2011).
Eddy, R. J., Weidmann, M. D., Sharma, V. P. & Condeelis, J. S. Tumor cell invadopodia: invasive protrusions that orchestrate metastasis. Trends Cell Biol. 27, 595–607 (2017).
Medjkane, S., Perez-Sanchez, C., Gaggioli, C., Sahai, E. & Treisman, R. Myocardin-related transcription factors and SRF are required for cytoskeletal dynamics and experimental metastasis. Nat. Cell Biol. 11, 257–268 (2009).
Jaiswal, M. et al. Functional cross-talk between ras and Rho pathways: a Ras-specific GTPase-activating protein (p120RasGAP) competitively inhibits the RhoGAP activity of deleted in liver cancer (DLC) tumor suppressor by masking the catalytic arginine finger. J. Biol. Chem. 289, 6839–6849 (2014).
Fujii, T., Iwane, A. H., Yanagida, T. & Namba, K. Direct visualization of secondary structures of F-actin by electron cryomicroscopy. Nature 467, 724–728 (2010).
Murakami, K. et al. Structural basis for actin assembly, activation of ATP hydrolysis, and delayed phosphate release. Cell 143, 275–287 (2010).
Aktories, K., Lang, A. E., Schwan, C. & Mannherz, H. G. Actin as target for modification by bacterial protein toxins. FEBS J. 278, 4526–4543 (2011).
Ceccarelli, D. F. et al. Non-canonical interaction of phosphoinositides with pleckstrin homology domains of Tiam1 and ArhGAP9. J. Biol. Chem. 282, 13864–13874 (2007).
Kong, L. & Ge, B. X. MyD88-independent activation of a novel actin-Cdc42/Rac pathway is required for Toll-like receptor-stimulated phagocytosis. Cell Res. 18, 745–755 (2008).
Winter, G., Lobley, C. M. & Prince, S. M. Decision making in xia2. Acta Crystallogr. D 69, 1260–1273 (2013).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of coot. Acta Crystallogr. D 66, 486–501 (2010).
Vaguine, A. A., Richelle, J. & Wodak, S. J. SFCHECK: a unified set of procedures for evaluating the quality of macromolecular structure-factor data and their agreement with the atomic model. Acta Crystallogr. D 55, 191–205 (1999).
The PyMOL Molecular Graphics System, Version 2.1.1 (Schrödinger, LLC, 2010).
Edelstein, A. D. et al. Advanced methods of microscope control using muManager software. J. Biol. Methods 1, e10 (2014).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
We thank the Crick Science Technology Platforms for their support and advice during this work, especially M. Renshaw and K. Anderson (Advanced Light Microscopy), P. Chakravarty and A. Stewart (Bioinformatics and Biostatistics); C. Watkins and J. Bee (Biological Research); N. Patel and A. Alidoust (Fermentation Facility); D. Davis (Flow Cytometry); G. Clark (Genomics Equipment Park); M. Howell (High-throughput screening); N. O’Reilly (Peptide Chemistry) and P. Walker (Structural Biology). X-ray data were collected at the Diamond Light Source (ID24 beamline, mx8015). We thank M. Matsuda (Kyoto University) for the RaichuEV-Rac plasmid and M. Way and members of the R.T. and N.Q.M. groups for their helpful discussions. This work was supported by Cancer Research UK core funding until 31 March 2015. Since then, support to R.T. and N.Q.M. has been provided by the Francis Crick Institute, which receives its core funding from Cancer Research UK (FC001-190 and FC001-115), the UK Medical Research Council (FC001-190 and FC001-115) and the Wellcome Trust (FC001-190 and FC001-115), and by an ERC Advanced Grant (grant no. 268690) to R.T.
Author information
Authors and Affiliations
Contributions
All authors designed and interpreted the experiments. J.D. conducted the biochemical and cell biology studies. S.M. determined the structure of the actin/ArhGAP12 complex and conducted the comparative structural analysis. J.D. and R.T. wrote the manuscript with input from S.M. and N.Q.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–8 and legend for Supplementary Table 1
Supplementary Table 1
Statistics source data.
Rights and permissions
About this article
Cite this article
Diring, J., Mouilleron, S., McDonald, N.Q. et al. RPEL-family rhoGAPs link Rac/Cdc42 GTP loading to G-actin availability. Nat Cell Biol 21, 845–855 (2019). https://doi.org/10.1038/s41556-019-0337-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0337-y
This article is cited by
-
Patterning of the cell cortex by Rho GTPases
Nature Reviews Molecular Cell Biology (2024)
-
TRIM56 acts through the IQGAP1-CDC42 signaling axis to promote glioma cell migration and invasion
Cell Death & Disease (2023)
-
Subtype-specific kinase dependency regulates growth and metastasis of poor-prognosis mesenchymal colorectal cancer
Journal of Experimental & Clinical Cancer Research (2023)
-
ARHGEF37 overexpression promotes extravasation and metastasis of hepatocellular carcinoma via directly activating Cdc42
Journal of Experimental & Clinical Cancer Research (2022)
-
Multiplexed live-cell profiling with Raman probes
Nature Communications (2021)