Abstract
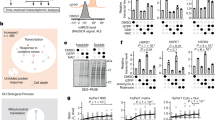
The cytosolic accumulation of mitochondrial precursors is hazardous to cellular fitness and is associated with a number of diseases. However, it is not observed under physiological conditions. Individual mechanisms that allow cells to avoid cytosolic accumulation of mitochondrial precursors have recently been discovered, but their interplay and regulation remain elusive. Here, we show that cells rapidly launch a global transcriptional programme to restore cellular proteostasis after induction of a ‘clogger’ protein that reduces the number of available mitochondrial import sites. Cells upregulate the protein folding and proteolytic systems in the cytosol and downregulate both the cytosolic translation machinery and many mitochondrial metabolic enzymes, presumably to relieve the workload of the overstrained mitochondrial import system. We show that this transcriptional remodelling is a combination of a ‘wideband’ core response regulated by the transcription factors Hsf1 and Rpn4 and a unique mitoprotein-induced downregulation of the oxidative phosphorylation components, mediated by an inactivation of the HAP complex.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Code availability
Custom code used for data analysis is available from the authors upon reasonable request.
Data availability
The data generated or analysed for the current study are included in this published article and its Supplementary Information. All numerical source data used for statistical analyses and graphical representations for Figs. 1–7 and Supplementary Figs. 1–7 are provided in Supplementary Table 9. The deep-sequencing (RNA-seq) raw data have been deposited into GEO with the accession number GSE116749. The MS proteomics data have been deposited to the ProteomeXchange Consortium via the PRIDE58 partner repository with the dataset identifier PXD011789. Visualizations of all RNA-seq and proteomics data are deposited into Figshare at https://doi.org/10.6084/m9.figshare.7611107. Previously published gene expression datasets59,60,61,62,63,64,65,66,67,68,69,70 (PMID numbers 11830665, 23643537, 18604275,18212068, 12740579, 26391581, 23222101, 16209719, 17327914, 11102521, 24952590, 12820961, 18753408) were downloaded from the SPELL database and are available at https://downloads.yeastgenome.org/expression/microarray/.
Change history
29 April 2019
In the version of this article originally published, parts of Figure 5 were misaligned because of a shift during production. In a, one data point was outside of the graph border. In b, axes lines were not connected, and graph lines did not reach the data points. In c and d, the axes lines were not connected. In e and g, the axes lines were not connected, and error bars and columns were not aligned. Shown below are the original and corrected versions of Figure 5. The errors have been corrected in the PDF and HTML versions of the paper.
References
Gold, V. A., Chroscicki, P., Bragoszewski, P. & Chacinska, A. Visualization of cytosolic ribosomes on the surface of mitochondria by electron cryo-tomography. EMBO Rep. 18, 1786–1800 (2017).
Costa, E. A., Subramanian, K., Nunnari, J. & Weissman, J. S. Defining the physiological role of SRP in protein-targeting efficiency and specificity. Science 359, 689–692 (2018).
Young, J. C., Hoogenraad, N. J. & Hartl, F. U. Molecular chaperones Hsp90 and Hsp70 deliver preproteins to the mitochondrial import receptor Tom70. Cell 112, 41–50 (2003).
Deshaies, R. J., Koch, B. D., Werner-Washburne, M., Craig, E. A. & Schekman, R. A subfamily of stress proteins facilitates translocation of secretory and mitochondrial precursor polypeptides. Nature 332, 800–805 (1988).
Hoseini, H. et al. The cytosolic cochaperone Sti1 is relevant for mitochondrial biogenesis and morphology. FEBS J. 283, 3338–3352 (2016).
Kim, H. E. et al. Lipid biosynthesis coordinates a mitochondrial-to-cytosolic stress response. Cell 166, 1539–1552 (2016).
Eisner, V., Picard, M. & Hajnoczky, G. Mitochondrial dynamics in adaptive and maladaptive cellular stress responses. Nat. Cell Biol. 20, 755–765 (2018).
Wrobel, L. et al. Mistargeted mitochondrial proteins activate a proteostatic response in the cytosol. Nature 524, 485–488 (2015).
Wang, X. & Chen, X. J. A cytosolic network suppressing mitochondria-mediated proteostatic stress and cell death. Nature 524, 481–484 (2015).
Weidberg, H., . & Amon, A. MitoCPR-A surveillance pathway that protects mitochondria in response to protein import stress. Science 360, eaan4146 (2018).
Eilers, M. & Schatz, G. Binding of a specific ligand inhibits import of a purified precursor protein into mitochondria. Nature 322, 228–232 (1986).
Xie, Y. & Varshavsky, A. RPN4 is a ligand, substrate, and transcriptional regulator of the 26S proteasome: a negative feedback circuit. Proc. Natl Acad. Sci. USA 98, 3056–3061 (2001).
Steffen, J., Seeger, M., Koch, A. & Kruger, E. Proteasomal degradation is transcriptionally controlled by TCF11 via an ERAD-dependent feedback loop. Mol. Cell 40, 147–158 (2010).
Owsianik, G., Balzi l, L. & Ghislain, M. Control of 26S proteasome expression by transcription factors regulating multidrug resistance in Saccharomyces cerevisiae. Mol. Microbiol. 43, 1295–1308 (2002).
Hahn, J. S., Neef, D. W. & Thiele, D. J. A stress regulatory network for co-ordinated activation of proteasome expression mediated by yeast heat shock transcription factor. Mol. Microbiol. 60, 240–251 (2006).
Zheng, X. et al. Dynamic control of Hsf1 during heat shock by a chaperone switch and phosphorylation. eLife 5, e18638 (2016).
Sontag, E. M., Samant, R. S. & Frydman, J. Mechanisms and functions of spatial protein quality control. Annu. Rev. Biochem. 86, 97–122 (2017).
Albanese, V., Yam, A. Y., Baughman, J., Parnot, C. & Frydman, J. Systems analyses reveal two chaperone networks with distinct functions in eukaryotic cells. Cell 124, 75–88 (2006).
Doring, K. et al. Profiling Ssb-nascent chain interactions reveals principles of Hsp70-assisted folding. Cell 170, 298–311 (2017).
Hibbs, M. A. et al. Exploring the functional landscape of gene expression: directed search of large microarray compendia. Bioinformatics 23, 2692–2699 (2007).
Morgenstern, M. et al. Definition of a high-confidence mitochondrial proteome at quantitative scale. Cell Rep. 19, 2836–2852 (2017).
Werner, T. et al. Ion coalescence of neutron encoded TMT 10-plex reporter ions. Anal. Chem. 86, 3594–3601 (2014).
Couvillion, M. T., Soto, I. C., Shipkovenska, G. & Churchman, L. S. Synchronized mitochondrial and cytosolic translation programs. Nature 533, 499–503 (2016).
Münch, C. & Harper, J. W. Mitochondrial unfolded protein response controls matrix pre-RNA processing and translation. Nature 534, 710–713 (2016).
Krakowiak, K. et al. Hsf1 and Hsp70 constitute a two-component feedback loop that regulates the yeast heat shock response. eLife 7, e31668 (2018).
Nargund, A. M., Pellegrino, M. W., Fiorese, C. J., Baker, B. M. & Haynes, C. M. Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337, 587–590 (2012).
Zambelli, F., Pesole, G. & Pavesi, G. Pscan: finding over-represented transcription factor binding site motifs in sequences from co-regulated or co-expressed genes. Nucleic Acids Res. 37, W247–W252 (2009).
Fleming, J. A., Lightcap, E. S., Sadis, S., Thoroddsen, V., Bulawa, C. E. & Blackman, R. K. Complementary whole-genome technologies reveal the cellular response to proteasome inhibition by PS-341. Proc. Natl Acad. Sci. USA 99, 1461–1466 (2002).
Buschlen, S., Amillet, J.-M., Guiard, B., Fournier, A., Marcireau, C. & Bolotin-Fukuhara, M. The S. cerevisiae HAP complex, a key regulator of mitochondrial function, coordinates nuclear and mitochondrial gene expression. Comp. Funct. Genomics 4, 37–46 (2003).
Woellhaf, M. W., Sommer, F., Schroda, M. & Herrmann, J. M. Proteomic profiling of the mitochondrial ribosome identifies Atp25 as a composite mitochondrial precursor protein. Mol. Biol. Cell 27, 3031–3039 (2016).
Brachmann, C. B. et al. Designer deletion strains derived from Saccharomyces cerevisiae S288C: a useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 14, 115–132 (1998).
Thomas, B. J. & Rothstein, R. Elevated recombination rates in transcriptionally active DNA. Cell 56, 619–630 (1989).
Chacinska, A. et al. Essential role of Mia40 in import and assembly of mitochondrial intermembrane space proteins. EMBO J. 23, 3735–3746 (2004).
Chacinska, A. et al. Mitochondrial presequence translocase: switching between TOM tethering and motor recruitment involves Tim21 and Tim17. Cell 120, 817–829 (2005).
Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004).
Ryan, M. T., Voos, W. & Pfanner, N. Assaying protein import into mitochondria. Methods Cell Biol. 65, 189–215 (2001).
Schmitt, M. E., Brown, T. A. & Trumpower, B. L. A rapid and simple method for preparation of RNA from Saccharomyces cerevisiae. Nucleic Acids Res. 18, 3091–3092 (1990).
Teste, M. A., Duquenne, M., Francois, J. M. & Parrou, J. L. Validation of reference genes for quantitative expression analysis by real-time RT-PCR in Saccharomyces cerevisiae. BMC Mol. Biol. 10, 99 (2009).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2–ΔΔCT method. Methods 25, 402–408 (2001).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Roy. Stat. Soc. B Met. 57, 289–300 (1995).
Yu, F. et al. Structural basis of intramitochondrial phosphatidic acid transport mediated by Ups1–Mdm35 complex. EMBO Rep. 16, 813–823 (2015).
Kwon, A. T., Arenillas, D. J., Worsley Hunt, R. & Wasserman, W. W. oPOSSUM-3: advanced analysis of regulatory motif over-representation across genes or ChIP-Seq datasets. G3 (Bethesda) 2, 987–1002 (2012).
Khan, A. et al. JASPAR 2018: update of the open-access database of transcription factor binding profiles and its web framework. Nucleic Acids Res. 46, D260–D266 (2018).
Hughes, C. S. et al. Ultrasensitive proteome analysis using paramagnetic bead technology. Mol. Syst. Biol. 10, 757 (2014).
Moggridge, S., Sorensen, P. H., Morin, G. B. & Hughes, C. S. Extending the compatibility of the sp3 paramagnetic bead processing approach for proteomics. J. Proteome Res. 17, 1730–1740 (2018).
Becher, I. et al. Pervasive protein thermal stability variation during the cell cycle. Cell 173, 1495–1507 (2018).
Franken, H. et al. Thermal proteome profiling for unbiased identification of direct and indirect drug targets using multiplexed quantitative mass spectrometry. Nat. Protoc. 10, 1567–1593 (2015).
Savitski, M. M., Wilhelm, M., Hahne, H., Kuster, B. & Bantscheff, M. A scalable approach for protein false discovery rate estimation in large proteomic data sets. Mol. Cell. Proteomics 14, 2394–2404 (2015).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Huber, W., von Heydebreck, A., Sultmann, H., Poustka, A. & Vingron, M. Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics 18, S96–S104 (2002).
Joe, H. & Ward, J. Hierarchical grouping to optimize an objective function. J. Am. Stat. Assoc. 58, 236–244 (1963).
Vizcaino, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
Worley, J., Luo, X. & Capaldi, A. P. Inositol pyrophosphates regulate cell growth and the environmental stress response by activating the HDAC Rpd3L. Cell Rep. 3, 1476–1482 (2013).
Willis, I. M. et al. Genetic interactions of MAF1 identify a role for Med20 in transcriptional repression of ribosomal protein genes. PLoS Genet. 4, e1000112 (2008).
Shivaswamy, S. & Iyer, V. R. Stress-dependent dynamics of global chromatin remodeling in yeast: dual role for SWI/SNF in the heat shock stress response. Mol. Cell. Biol. 28, 2221–2234 (2008).
Segal, E. et al. Module networks: identifying regulatory modules and their condition-specific regulators from gene expression data. Nat. Genet. 34, 166–176 (2003).
Oromendia, A. B., Dodgson, S. E. & Amon, A. Aneuploidy causes proteotoxic stress in yeast. Genes Dev. 26, 2696–2708 (2012).
Pastor-Flores, D. et al. Depletion of yeast PDK1 orthologs triggers a stress-like transcriptional response. BMC Genomics 16, 719 (2015).
Levy, S. et al. Strategy of transcription regulation in the budding yeast. PLoS One 2, e250 (2007).
Gasch, A. P. et al. Genomic expression programs in the response of yeast cells to environmental changes. Mol. Cell Biol. 11, 4241–4257 (2000).
O'Duibhir, E. et al. Cell cycle population effects in perturbation studies. Mol. Syst. Biol. 10, 732–732 (2014).
Düvel, K., Santhanam, A., Garrett, S., Schneper, L. & Broach, J. R. Multiple roles of Tap42 in mediating rapamycin-induced transcriptional changes in yeast. Mol. Cell 11, 1467–1478 (2003).
Berry, D. B., Gasch, A. P. & Weissman, J. S. Stress-activated genomic expression changes serve a preparative role for impending stress in yeast. Mol. Biol. Cell 19, 4580–4587 (2008).
Matsumoto, R., Akama, K., Rakwal, R. & Iwahashi, H. The stress response against denatured proteins in the deletion of cytosolic chaperones SSA1/2 is different from heat-shock response in Saccharomyces cerevisiae. BMC Genomics 6, 141 (2005).
Acknowledgements
The authors thank the following individuals: K. Hansen, S. Backes, J. Laborenz, K. Knoeringer, S. Knaus, A. Trinkaus, J. Pistolic and J. Provaznik for help with the experiments; M. Schuldiner, D. Pincus, S. Schuck, D. Wolf, G. Kramer, K. Morano, D. Thiele, M. Bolotin-Fukuhara and T. Becker for reagents and strains; and B. Morgan, S. Schuck and M. Schuldiner for comments on the manuscript. The authors acknowledge the Deutsche Forschungsgemeinschaft (DIP MitoBalance, IRTG1830 and HE2803/8-2 to J.M.H.; SFB 894 and IRTG 1830 to M.v.d.L.), the Joachim Herz Stiftung (to F.B.), the Forschungsinitiative Rheinland Pfalz BioComp (to J.M.H.) and the Boehringer Ingelheim Fonds (predoctoral fellowship to F.W.) for funding.
Author information
Authors and Affiliations
Contributions
F.B. and J.M.H. conceived the project. F.B., L.K., C.G. and A.G. designed, cloned and verified the constructs and strains, and F.B., C.G., F.J. and V.B. produced the cDNA and carried out the RNA-seq experiments. F.B., P.H. and M.M.S. carried out the MS experiments. F.B., E.Z., L.K. and C.G. measured the transcript and protein levels in cell extracts, and F.B., F.W. and M.v.d.L. performed the blue native–PAGE assays. F.B. performed the bioinformatics analyses of the RNA-seq data, and F.B. and F.S. performed the bioinformatics analyses of the proteomics data. F.B., L.K. and C.G. designed and set up the gene expression reporter assays. F.B., L.K., C.G., A.G. and J.M.H. analysed the data, and F.B. and J.M.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Clogger expression leads to characteristic changes of the transcriptome and prevents cell growth, but does not cause cell death or respiration deficiencies.
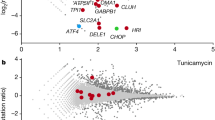
a, Clogger-expressing wild type cells and controls were grown to log phase on synthetic lactate medium. Tenfold serial dilutions were dropped onto lactate plates lacking (left) or containing 0.5% galactose (middle). For the plate shown on the right, samples were pre-grown for 4.5 h on galactose-containing medium to induce the expression of the DHFR fusion proteins. Cells were washed and spotted on galactose-free plates. Despite the strong growth arrest in the presence of galactose, clogger-expressing strains were fully viable as soon as clogger expression was repressed. b, Cell viability was measured by a CASY cell counter upon several hours of clogger induction. A wildtype strain was treated with CASYblue for 5 min according to the manufacturers instructions as reference for dying cells. c, Cells were grown in galactose-containing medium for 4.5 h, harvest, resuspended in 1 ml of lactate medium and their oxygen consumption rate was measured. Clogger induction for 4.5 h did not affect respiration rates. Experiments were repeated independently n = 4 (a) or n = 2 (b, c) times with similar results. d, Volcano Plots showing the relative log2 fold changes (mean of n = 4 independent biological samples) of cells expressing cytosolic DHFR, b2-DHFR or b2Δ-DHFR relative to strain containing just an empty vector. While cytosolic DHFR does not alter gene expression, the expression of four characteristic groups of genes is consistently changed upon induction of the cloggers: Several members of the chaperone system / protein folding (GO:0006457) are induced within 30 min. The proteasome complex (GO:0000502) is induced between 30 and 90 min after clogger induction. The cytosolic ribosome (GO:0022626) and components of the oxidative phosphorylation machinery (GO:0006119) are repressed between 30 and 90 min after clogger induction. P values were calculated with the likelihood ratio test and corrected for multiple hypothesis testing with the Benjamini-Hochberg procedure within the DESeq2 package.
Supplementary Figure 2 The transcription factor Rpn4 regulates the clogger-mediated expression of proteasome subunits via PACE promoter elements.
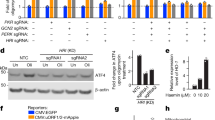
a Expression profiles of proteasome core subunits and other factors of the ubiquitin-proteasome system in clogger expressing and control cells. The read numbers were only corrected for the size differences of the four cDNA libraries and are directly shown. Please note the surprisingly homogeneous upregulation of all core subunits of the proteasome and other proteins of the ubiquitin-proteasome system between the 30 and 90 min time point after clogger induction. b, Logo plot presentation of PACE elements which is similar but different to the binding site of Met31/32. c, Overview of the synthetic Rpn4 activity reporter consisting of four PACE elements upstream of a minimal promoter and a YFP-coding reading frame. d, Schematic representation of the fluorescence reporter for the detection and quantification of Rpn4-mediated gene expression. e, An increase in temperature induces an Rpn4-driven YFP expression. f, In the absence of Rpn4, YFP expression is very low and does not increase upon clogger induction. This shows that the reporter gene specifically responds to Rpn4 activation. g, Transcript levels of RPN8 were measured upon clogger induction of 4.5 h in wild type or Δrpn4 cells. h, The levels of proteasome subunits in cell extracts of the indicated strains were analyzed by Western blotting. Bar graphs show mean values and standard deviations. Experiments were repeated independently n = 4 (a), n = 1 (e) or n = 3 (f, g, h) times; significance was assessed with a one-sided, paired Student’s t-test. Unprocessed blots are shown in Supplementary Fig. 8.
Supplementary Figure 3 On basis of their response to clogger induction, chaperones can be sorted into different categories.
a, Based on the profiles of transcript (left, n = 4 independent biological samples) or protein (right, n = 3 independent biological samples) levels relative to control strains, genes from the protein folding GO term (GO:0006457) were clustered according to correlation of their profiles over time. Both datasets gave rise to similar groups, which also matched with reported functions of the comprised genes: Components of the general posttranslational folding machinery (such as Ssa1-Ssa4, Hsp82 or Ydj1) are strongly and rapidly induced by clogger induction. Several cochaperones and, notably, all subunits of the cytosolic chaperonin show a moderate and late upregulation and drop again at late time points on the protein level. Cotranslationally acting chaperones (such as Ssb1/Ssb2, Ssz1, zuotin) are slightly downregulated upon clogger induction. Components of the general posttranslational folding machinery (such as Ssa1-Ssa4, Ssc1 or Ydj1) are strongly and rapidly induced by clogger induction. b, Overexpression of Hsp70 (Ssa2) and Hsp40 (Ydj1) from strong constitutive promoters partially restored growth upon clogger induction. The experiment was repeated independently n = 2 times with similar results.
Supplementary Figure 4 Mitoprotein-induced stress inhibits both cytosolic and mitochondrial protein synthesis.
a, Radiolabelling of newly synthesized proteins showed a strong reduction of translation rates 3 h after clogger induction. b, Protein components of the cytosolic ribosome slightly decreased in abundance upon clogger induction as measured by mass spectrometry. c, mRNA levels of the mitochondrially encoded gene ATP6 showed no decrease upon clogger induction. Means and standard deviations are shown. d, Protein levels of Cox1 and Cox2 moderately reduced over time, as measured by mass spectrometry. e, Synthesis of mitochondrial translation products upon clogger induction was analyzed by radioactive labeling in the presence of cycloheximide. Blocking mitochondrial protein import strongly decreased mitochondrial translation. f, g, Reduced levels of mitochondrial ribosomal proteins were observed in the proteomics analysis, including the mitochondrially encoded subunit Var1. This effect was not seen on the transcript level. Experiments were repeated independently n = 1 (a, e) or n = 3 (b, c, d, f, g) times. p values were calculated according to a two-sided Student’s t-test, adjusted for multiple comparisons using the Benjamini-Hochberg procedure (see Methods for details). Unprocessed blots are shown in Supplementary Fig. 8.
Supplementary Figure 5 Mitoprotein-induced stress induces downregulation of OXPHOS genes.
a, The repression of OXPHOS genes such as FUM1 is independent of Rpn4. b, Clogger induction affects genes that contain a HAP complex-binding CCAAT element in a similar manner than the deletion of Hap2. c, Hap4 levels are reduced upon induction of clogger proteins. d, Predominantly the highly abundant mitochondrial proteins are downregulated upon clogger induction. e, Overexpression of HAP4 from a GAL promoter did not affect clogger expression nor altered induction of chaperones. Means and standard deviations are shown. Experiments were repeated independently n = 3 (a), n = 1 (c) or n = 4 (e) times. Unprocessed blots are shown in Supplementary Fig. 8.
Supplementary Figure 6 Deep proteomic profiling of clogger-induced changes in protein abundance.
a, Samples were collected from clogger expressing and control cells after indicated times of induction. Protein extracts were digested with trypsin and labelled with TMT10 reagent. Samples were mixed and subjected to mass spectrometry. Reporter ions were quantified in the MS2 spectra. b, 3,511 proteins were quantified in at least two out of three replicates (independent biological samples). c, Mitoprotein-induced stress alters the cellular proteome in a consistent manner which is clearly distinct from the changes induced by the addition of galactose. d, Overall fold changes of clogger expressing versus control cells increased over time. Genes with the strongest up- or down-regulation at each time point are labelled. e, Changes in protein abundance of clogger-expressing cells relative to control cells over time were measured by mass spectrometry. Shown here are mean log2 fold changes from n = 3 independent biological samples. Consistent with the transcriptional responses, a consistent induction of the proteasome as well as of many chaperones is observed, as well as prominent decrease in OXPHOS components and reduced levels of cytosolic ribosomes and its associated chaperones. Some components of the mitochondrial protein import machinery decrease in abundance, while they were not affected on the transcript level. Experiments were repeated independently n = 3 (b-e) times. Significance was assessed with a two-sided Student’s t-test, adjusted for multiple comparisons using the Benjamini-Hochberg procedure (see Methods for details).
Supplementary Figure 7 The mitoprotein-induced stress response is similar to, but distinct from a heat stress response and comprises the Pdr3-mediated mitoCPR response as a late sub-routine.
a, The clogger-induced expression changes were compared to the patterns of the yeast transcription SPELL database. Shown are percentages of transcription datasets that are similar to that of clogger-induced expression changes. b, For all heat stress transcriptome analyses deposited in SPELL, the mean changes in gene expression for the four GO terms shown were calculated and compared to that of the clogger-mediated response (after 90 min). Note that in none of the studies except the present one on mitoprotein-induced stress, a combination of OXPHOS downregulation with the classical heat stress responses of chaperones, proteasome and 80S ribosome was observed59,60,61,62,63,64,65,66,67,68,69,70. c, Expression profile of PDR3 upon clogger induction. d, GO enrichment analysis of genes with expression profiles that are similar to that of PDR3. e, The promoter of PDR3 contains two PACE elements and two PDRE elements. f, Individual profiles of genes that are transcriptionally controlled by Pdr3. Experiments were repeated independently n = 4 times (a–f). P values for enrichment (a, d) were calculated assuming a hypergeometric distribution (one-sided Fisher’s exact test) and corrected for multiple hypothesis testing using the Benjamini-Hochberg procedure.
Supplementary Figure 8
Unprocessed images of all gels and blots.
Supplementary information
Supplementary Information
Supplementary Figures 1–8 and legends for Supplementary Tables 1–9.
Supplementary Table 1
Yeast strains used in this study.
Supplementary Table 2
Plasmids used in this study.
Supplementary Table 3
qPCR primers used in this study.
Supplementary Table 4
RNAseq data. Read counts, normalized with respect to library size.
Supplementary Table 5
RNAseq data. log2 fold changes normalized to the empty vector control.
Supplementary Table 6
Pairwise similarity scores for the transcript profiles of all genes measured in the RNA-seq experiment.
Supplementary Table 7
Experimentally verified target genes of the transcription factors Hsf1, Yap1 and Pdr1/Pdr3.
Supplementary Table 8
Proteomics data. log2 fold changes of b2-DHFR normalized to cyt DHFR.
Supplementary Table 9
Statistics source data.
Rights and permissions
About this article
Cite this article
Boos, F., Krämer, L., Groh, C. et al. Mitochondrial protein-induced stress triggers a global adaptive transcriptional programme. Nat Cell Biol 21, 442–451 (2019). https://doi.org/10.1038/s41556-019-0294-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0294-5
This article is cited by
-
Prefoldin 2 contributes to mitochondrial morphology and function
BMC Biology (2023)
-
Mitochondrial complexome reveals quality-control pathways of protein import
Nature (2023)
-
A cytosolic surveillance mechanism activates the mitochondrial UPR
Nature (2023)
-
Immunoproteasome-specific subunit PSMB9 induction is required to regulate cellular proteostasis upon mitochondrial dysfunction
Nature Communications (2023)
-
Selection of red fluorescent protein for genetic labeling of mitochondria and intercellular transfer of viable mitochondria
Scientific Reports (2022)