Abstract
Balanced chromosomal rearrangements such as inversions and translocations can cause congenital disease or cancer by inappropriately rewiring promoter–enhancer contacts1,2. To study the potentially pathogenic consequences of balanced chromosomal rearrangements, we generated a series of genomic inversions by placing an active limb enhancer cluster from the Epha4 regulatory domain at different positions within a neighbouring gene-dense region and investigated their effects on gene regulation in vivo in mice. Expression studies and high-throughput chromosome conformation capture from embryonic limb buds showed that the enhancer cluster activated several genes downstream that are located within asymmetric regions of contact, the so-called architectural stripes3. The ectopic activation of genes led to a limb phenotype that could be rescued by deleting the CCCTC-binding factor (CTCF) anchor of the stripe. Architectural stripes appear to be driven by enhancer activity, because they do not form in mouse embryonic stem cells. Furthermore, we show that architectural stripes are a frequent feature of developmental three-dimensional genome architecture often associated with active enhancers. Therefore, balanced chromosomal rearrangements can induce ectopic gene expression and the formation of asymmetric chromatin contact patterns that are dependent on CTCF anchors and enhancer activity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Code availability
Custom code to predict domains with asymmetric stripes is available at https://github.com/VerenaHeinrich/HiC2Imbalance.
Data availability
Hi-C, cHi-C, ChIP-seq and RNA-seq data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE116794. Previously published ChIP-seq data that were reanalysed here are available under accession code GSE847959 and GSE3103936. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Hnisz, D. et al. Activation of proto-oncogenes by disruption of chromosome neighborhoods. Science 351, 1454–1458 (2016).
Lupiáñez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Vian, L. et al. The energetics and physiological impact of cohesin extrusion. Cell 173, 1165–1178 (2018).
Dixon, J. R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012).
Rao, S. S. P. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Vietri Rudan, M. et al. Comparative Hi-C reveals that CTCF underlies evolution of chromosomal domain architecture. Cell Rep. 10, 1297–1309 (2015).
Andrey, G. et al. Characterization of hundreds of regulatory landscapes in developing limbs reveals two regimes of chromatin folding. Genome Res. 27, 223–233 (2017).
Mumbach, M. R. et al. Enhancer connectome in primary human cells identifies target genes of disease-associated DNA elements. Nat. Genet. 49, 1602–1612 (2017).
Franke, M. et al. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature 538, 265–269 (2016).
Spielmann, M., Lupiáñez, D. G. & Mundlos, S. Structural variation in the 3D genome. Nat. Rev. Genet. 19, 453–467 (2018).
Dali, R. & Blanchette, M. A critical assessment of topologically associating domain prediction tools. Nucleic Acids Res. 45, 2994–3005 (2017).
Will, A. J. et al. Composition and dosage of a multipartite enhancer cluster control developmental expression of Ihh (Indian hedgehog). Nat. Genet. 49, 1539–1545 (2017).
Jäger, R. et al. Capture Hi-C identifies the chromatin interactome of colorectal cancer risk loci. Nat. Commun. 6, 6178 (2015).
Schuettengruber, B., Chourrout, D., Vervoort, M., Leblanc, B. & Cavalli, G. Genome regulation by polycomb and trithorax proteins. Cell 128, 735–745 (2007).
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125, 315–326 (2006).
Mammana, A. & Chung, H.-R. Chromatin segmentation based on a probabilistic model for read counts explains a large portion of the epigenome. Genome Biol. 16, 151 (2015).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Zabidi, M. A. et al. Enhancer–core-promoter specificity separates developmental and housekeeping gene regulation. Nature 518, 556–559 (2015).
Kraft, K. et al. Deletions, inversions, duplications: engineering ofstructural variants using CRISPR/Cas in mice. Cell Rep. 10, 833–839 (2015).
George, S. H. L. et al. Developmental and adult phenotyping directly from mutant embryonic stem cells. Proc. Natl Acad. Sci. USA 104, 4455–4460 (2007).
Artus, J. & Hadjantonakis, A.-K. Generation of chimeras by aggregation of embryonic stem cells with diploid or tetraploid mouse embryos. Methods Mol. Biol. 693, 37–56 (2011).
Kragesteen, B. K. et al. Dynamic 3D chromatin architecture contributes to enhancer specificity and limb morphogenesis. Nat. Genet. 50, 1463–1473 (2018).
Knight, P. A. & Ruiz, D. A fast algorithm for matrix balancing. IMA J. Numer. Anal. 33, 1029–1047 (2013).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Shin, H. et al. TopDom: an efficient and deterministic method for identifying topological domains in genomes. Nucleic Acids Res. 44, e70 (2016).
Wingett, S. et al. HiCUP: pipeline for mapping and processing Hi-C data. F1000 Res. 4, 1310 (2015).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Love, M. I., Huber, W. & Anders, M. I. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Lee, T. I., Johnstone, S. E. & Young, R. A. Chromatin immunoprecipitation and microarray-based analysis of protein location. Nat. Protoc. 1, 729–748 (2006).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
ENCODE Project Consortium An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Acknowledgements
This study was supported by grants from Deutsche Forschungsgemeinschaft to S.M. and K.K. We thank J. Fiedler and K. Macura from the transgenic facility; M. Hochradel from the sequencing facility MPIMG; I. Jerkovic for the Rad21 protocol; N. Brieske for WISH; the Mundlos group and B. Flensburger in Berlin; members of the Chang lab and B. Tekila in Stanford; and E. Nora and G. Fudenberg for scientific discussions and support.
Author information
Authors and Affiliations
Contributions
K.K. and S.M. conceived the project. K.K. and A.M. performed cHi-C. V.H., J.M and R.S. performed the computational analysis with input from M.V. K.K., C.R. and A.M. produced transgenics, carried out transgenic validation and expressional/phenotypical analysis. D.M.I., R.A.H. and A.D. performed Hi-C. G.A. and A.M. performed ChIP-seq. B.T. sequenced the cHi-C samples. L.W. performed morula aggregation. K.K., S.M. and A.M. wrote the manuscript with input from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Chromatin states, phenotypical and expressional analysis study in serial genomic inversions.
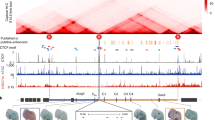
A) Zoom-in of the promoter region of the first gene at the breakpoint in the different inversions. B-1) Genome-wide segmentation was performed based on ChIP-seq histone modification data (H3K27ac, H3K27me3, H3K4me1, H3K4me3, H3K9me3) with EpiCSeg using seven states. For all HMs a minimum mapping quality threshold of 10 was used, except for H3K9me3. The Average mark intensities (log) per state are shown in the heatmap. Apart from H3K9me3, ChIP-seq data were obtained from a previously published data set (accession code: GSE84795). H3K9me3 ChIP-seq was performed once. B-2) Genome-wide segmentation as in B-1) but without H3K9me3 and with six defined chromatin states. C) Relation of contact frequency and log2-fold change dependent on H3K27me3 presence at promoter prior to activation. Contact frequency between enhancer and new target promoter is calculated from cHi-C for each Inversion and logFC is calculated from RNA-seq. D) Whole mount in situ hybridization with Ihh probe stained and stopped simultaneously and skeletal staining for each inversion. Guides used to generate mutants are indicated by different colors corresponding to the different rearrangements. Wild type serves as the negative control, the Dbf mouse mutant is shown as a positive control. Mice develop polydactyly when Ihh is ectopically expressed at stage E11.5 in the anterior/posterior portion of the limb bud (Dbf, Inv2, and Inv1). In Inv3 and Inv4 Ihh misexpression is also present, but here it is distributed across the limb bud and no polydactyly is observed. WISH experiments were performed three times independently with similar results.
Supplementary Figure 2 The occurrence of the architectural stripe is connected to a significant misexpression of several genes downstream of the enhancer cluster.
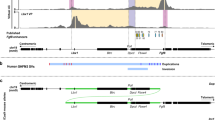
Zoom-in at Inv1 featuring the architectural stripe. The anchor point of the structure is indicated by a blue hexagon. The enhancer underneath the structure (indicated by purple oval) is able to activate several genes (see RNA-seq track with fold changes; WT: n=5 biologically independent samples; Inv1: n=3 biologically independent samples; each biologically independent sample refers to one pair of limb buds collected from a single embryo; significance is shown for adjusted p-values which was derived from DESEQ2) and skips a convergent CTCF site which binds cohesin (CTCF/Rad21 ChIP-seq was performed once with limb buds pooled from 10 embryos per condition).
Supplementary Figure 3 Contact frequencies for each genomic inversion between the promoter and individual enhancers or the boundary, respectively.
The contact frequency (cf; calculated from cHi-C data) illustrates that overall the anchor point contacts promoter at higher rates than the individual enhancers. As expected, the cf decays with increasing distance to centromeric side (left) and is directly insulated at the telomeric side (right) because of the stripe anchor. The green line indicates the inverted region. cHiC was performed once with limb buds collected from 10 embryos per condition.
Supplementary Figure 4 Relation between chromatin states (output of EpiCSeg) and their relative occurrence in all binned asymmetric chromatin domains.
The only correlation between stripe anchors and chromatin state abundance can be found for the state ‘Active 2’ (active enhancer state). Additionally, for the states ‘active 1’ (found at promoters) and ‘repressed 2’ the frequencies are very low, indicating that these chromatin states are almost absent at asymmetric chromatin domains. Smoothed curves (solid lines) are fitted using loess regression (‘stat_smooth’ function, ggplot2 R package, v3.1.0). Additionally, 95% confidence intervals are displayed in shaded tones.
Supplementary information
Supplementary Information
Supplementary Figures 1–4 and legend for Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Kraft, K., Magg, A., Heinrich, V. et al. Serial genomic inversions induce tissue-specific architectural stripes, gene misexpression and congenital malformations. Nat Cell Biol 21, 305–310 (2019). https://doi.org/10.1038/s41556-019-0273-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-019-0273-x
This article is cited by
-
BRWD1 orchestrates small pre-B cell chromatin topology by converting static to dynamic cohesin
Nature Immunology (2024)
-
Boundary stacking interactions enable cross-TAD enhancer–promoter communication during limb development
Nature Genetics (2024)
-
Pan-3D genome analysis reveals structural and functional differentiation of soybean genomes
Genome Biology (2023)
-
Three-dimensional genome organization in immune cell fate and function
Nature Reviews Immunology (2023)
-
The spatial organization of transcriptional control
Nature Reviews Genetics (2023)