Abstract
Drug-based strategies to overcome tumour resistance to radiotherapy (R-RT) remain limited by the single-agent toxicity of traditional radiosensitizers (for example, platinums) and a lack of targeted alternatives. In a screen for compounds that restore radiosensitivity in p53 mutant zebrafish while tolerated in non-irradiated wild-type animals, we identified the benzimidazole anthelmintic oxfendazole. Surprisingly, oxfendazole acts via the inhibition of IRAK1, a kinase thus far implicated in interleukin-1 receptor (IL-1R) and Toll-like receptor (TLR) immune responses. IRAK1 drives R-RT in a pathway involving IRAK4 and TRAF6 but not the IL-1R/TLR–IRAK adaptor MyD88. Rather than stimulating nuclear factor-κB, radiation-activated IRAK1 prevented apoptosis mediated by the PIDDosome complex (comprising PIDD, RAIDD and caspase-2). Countering this pathway with IRAK1 inhibitors suppressed R-RT in tumour models derived from cancers in which TP53 mutations predict R-RT. Moreover, IRAK1 inhibitors synergized with inhibitors of PIN1, a prolyl isomerase essential for IRAK1 activation in response to pathogens and, as shown here, in response to ionizing radiation. These data identify an IRAK1 radiation-response pathway as a rational chemoradiation therapy target.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Code availability
All codes used are referenced in the Methods sections above.
Data availability
The whole-genome expression array data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE67614. The human HNSCC and breast cancer data were derived from TCGA Research Network: http://cancergenome.nih.gov/. The dataset derived from this resource that supports the findings of this study is available in Supplementary Table 4 (see sheet S3). Source data for Figs. 1e, 2b,d, 3c,e,f, 4a–e,g,h,m,n, 5b,d, 6c–j, 7a–d,f,g and Supplementary Figs. 1b, 2a,b, 3, 4a–f,k,l, 5, 6c–n and 7c,d are provided as Supplementary Table 4. The full image dataset that supports the findings in Fig. 3e,f has been deposited in Figshare at https://doi.org/10.6084/m9.figshare.7427942. All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
Change history
01 February 2019
In the sentence beginning “Moreover, IRAK1 inhibitors...” in the Abstract of the version of this Article originally published, it was unclear whether the present study shows synergy between IRAK1 and PIN1 inhibitors in response to pathogens or ionizing radiation. The sentence has been modified to clarify this.
References
Lawrence, Y. R. et al. NCI-RTOG translational program strategic guidelines for the early-stage development of radiosensitizers. J. Natl Cancer Inst. 105, 11–24 (2013).
Sharma, R. A. et al. Clinical development of new drug–radiotherapy combinations. Nat. Rev. Clin. Oncol. 13, 627–642 (2016).
Wilson, G. D., Bentzen, S. M. & Harari, P. M. Biologic basis for combining drugs with radiation. Semin. Radiat. Oncol. 16, 2–9 (2006).
Golden, E. B., Formenti, S. C. & Schiff, P. B. Taxanes as radiosensitizers. Anticancer Drugs 25, 502–511 (2014).
Adelstein, D. J. et al. An intergroup phase III comparison of standard radiation therapy and two schedules of concurrent chemoradiotherapy in patients with unresectable squamous cell head and neck cancer. J. Clin. Oncol. 21, 92–98 (2003).
Olivier, M. et al. The IARC TP53 database: new online mutation analysis and recommendations to users. Hum. Mutat. 19, 607–614 (2002).
Igney, F. H. & Krammer, P. H. Death and anti-death: tumour resistance to apoptosis. Nat. Rev. Cancer 2, 277–288 (2002).
Vogelstein, B., Lane, D. & Levine, A. J. Surfing the p53 network. Nature 408, 307–310 (2000).
Vousden, K. H. & Lu, X. Live or let die: the cell’s response to p53. Nat. Rev. Cancer 2, 594–604 (2002).
Poeta, M. L. et al. TP53 mutations and survival in squamous-cell carcinoma of the head and neck. N. Engl. J. Med. 357, 2552–2561 (2007).
Skinner, H. D. et al. TP53 disruptive mutations lead to head and neck cancer treatment failure through inhibition of radiation-induced senescence. Clin. Cancer Res. 18, 290–300 (2012).
Nam, T. K. et al. Molecular prognostic factors in rectal cancer treated by preoperative chemoradiotherapy. Oncol. Lett. 1, 23–29 (2010).
Sclafani, F.et al. TP53 mutational status and cetuximab benefit in rectal cancer: 5-year results of the EXPERT-C trial. J. Natl Cancer Inst. 106, dju121 (2014).
Eikesdal, H. P., Knappskog, S., Aas, T. & Lonning, P. E. TP53 status predicts long-term survival in locally advanced breast cancer after primary chemotherapy. Acta Oncol. 53, 1347–1355 (2014).
Li, S., Zhang, W., Chen, B., Jiang, T. & Wang, Z. Prognostic and predictive value of p53 in low MGMT expressing glioblastoma treated with surgery, radiation and adjuvant temozolomide chemotherapy. Neurol. Res. 32, 690–694 (2010).
Tabori, U. et al. Universal poor survival in children with medulloblastoma harboring somatic TP53 mutations. J. Clin. Oncol. 28, 1345–1350 (2010).
Berghmans, S. et al. tp53 mutant zebrafish develop malignant peripheral nerve sheath tumors. Proc. Natl Acad. Sci. USA 102, 407–412 (2005).
Sidi, S. et al. Chk1 suppresses a caspase-2 apoptotic response to DNA damage that bypasses p53, Bcl-2, and caspase-3. Cell 133, 864–877 (2008).
McAleer, M. F. et al. Novel use of zebrafish as a vertebrate model to screen radiation protectors and sensitizers. Int. J. Radiat. Oncol. Biol. Phys. 61, 10–13 (2005).
Lacey, E. Mode of action of benzimidazoles. Parasitol. Today 6, 112–115 (1990).
Prichard, R. K. & Ranjan, S. Anthelmintics. Vet. Parasitol. 46, 113–120 (1993).
Keiser, M. J. et al. Relating protein pharmacology by ligand chemistry. Nat. Biotechnol. 25, 197–206 (2007).
Jain, A., Kaczanowska, S. & Davila, E. IL-1 receptor-associated kinase signaling and its role in inflammation, cancer progression, and therapy resistance. Front. Immunol. 5, 553 (2014).
Janssens, S. & Beyaert, R. Functional diversity and regulation of different interleukin-1 receptor-associated kinase (IRAK) family members. Mol. Cell 11, 293–302 (2003).
Powers, J. P. et al. Discovery and initial SAR of inhibitors of interleukin-1 receptor-associated kinase-4. Bioorg. Med. Chem. Lett. 16, 2842–2845 (2006).
Wang, Z. et al. Crystal structures of IRAK-4 kinase in complex with inhibitors: a serine/threonine kinase with tyrosine as a gatekeeper. Structure 14, 1835–1844 (2006).
O’Neill, L. A., Golenbock, D. & Bowie, A. G. The history of Toll-like receptors—redefining innate immunity. Nat. Rev. Immunol. 13, 453–460 (2013).
Rolf, M. G. et al. In vitro pharmacological profiling of R406 identifies molecular targets underlying the clinical effects of fostamatinib. Pharmacol. Res. Perspect. 3, e00175 (2015).
Joh, E. H., Lee, I. A., Jung, I. H. & Kim, D. H. Ginsenoside Rb1 and its metabolite compound K inhibit IRAK-1 activation–the key step of inflammation. Biochem. Pharmacol. 82, 278–286 (2011).
Yao, J. et al. Interleukin-1 (IL-1)-induced TAK1-dependent versus MEKK3-dependent NFkappaB activation pathways bifurcate at IL-1 receptor-associated kinase modification. J. Biol. Chem. 282, 6075–6089 (2007).
Kollewe, C. et al. Sequential autophosphorylation steps in the interleukin-1 receptor-associated kinase-1 regulate its availability as an adapter in interleukin-1 signaling. J. Biol. Chem. 279, 5227–5236 (2004).
Wee, Z. N. et al. IRAK1 is a therapeutic target that drives breast cancer metastasis and resistance to paclitaxel. Nat. Commun. 6, 8746 (2015).
Muroi, M. & Tanamoto, K. IRAK-1-mediated negative regulation of Toll-like receptor signaling through proteasome-dependent downregulation of TRAF6. Biochim. Biophys. Acta 1823, 255–263 (2012).
Rhyasen, G. W. & Starczynowski, D. T. IRAK signalling in cancer. Br. J. Cancer 112, 232–237 (2015).
Conze, D. B., Wu, C. J., Thomas, J. A., Landstrom, A. & Ashwell, J. D. Lys63-linked polyubiquitination of IRAK-1 is required for interleukin-1 receptor- and toll-like receptor-mediated NF-kappaB activation. Mol. Cell. Biol. 28, 3538–3547 (2008).
Ando, K. et al. PIDD death-domain phosphorylation by ATM controls prodeath versus prosurvival PIDDosome signaling. Mol. Cell 47, 681–693 (2012).
Sladky, V., Schuler, F., Fava, L. L. & Villunger, A. The resurrection of the PIDDosome—emerging roles in the DNA-damage response and centrosome surveillance. J. Cell Sci. 130, 3779–3787 (2017).
Tinel, A. & Tschopp, J. The PIDDosome, a protein complex implicated in activation of caspase-2 in response to genotoxic stress. Science 304, 843–846 (2004).
Tinel, A. et al. Autoproteolysis of PIDD marks the bifurcation between pro-death caspase-2 and pro-survival NF-kappaB pathway. EMBO J. 26, 197–208 (2007).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Essmann, F., Engels, I. H., Totzke, G., Schulze-Osthoff, K. & Janicke, R. U. Apoptosis resistance of MCF-7 breast carcinoma cells to ionizing radiation is independent of p53 and cell cycle control but caused by the lack of caspase-3 and a caffeine-inhibitable event. Cancer Res. 64, 7065–7072 (2004).
Lu, Z. & Hunter, T. Prolyl isomerase Pin1 in cancer. Cell Res. 24, 1033–1049 (2014).
Wei, S. et al. Active Pin1 is a key target of all-trans retinoic acid in acute promyelocytic leukemia and breast cancer. Nat. Med. 21, 457–466 (2015).
Skinner, H. D. et al. Proteomic profiling identifies PTK2/FAK as a driver of radioresistance in HPV-negative head and neck cancer. Clin. Cancer Res. 22, 4643–4650 (2016).
Cancer Genome Atlas Network Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 517, 576–582 (2015).
Tun-Kyi, A. et al. Essential role for the prolyl isomerase Pin1 in Toll-like receptor signaling and type I interferon-mediated immunity. Nat. Immunol. 12, 733–741 (2011).
Moore, J. D. & Potter, A. Pin1 inhibitors: pitfalls, progress and cellular pharmacology. Bioorg. Med. Chem. Lett. 23, 4283–4291 (2013).
Urusova, D. V. et al. Epigallocatechin-gallate suppresses tumorigenesis by directly targeting Pin1. Cancer Prev. Res. (Phila.) 4, 1366–1377 (2011).
Marsolier, J. et al. Theileria parasites secrete a prolyl isomerase to maintain host leukocyte transformation. Nature 520, 378–382 (2015).
Candeias, S. M. & Testard, I. The many interactions between the innate immune system and the response to radiation. Cancer Lett. 368, 173–178 (2015).
Reisz, J. A., Bansal, N., Qian, J., Zhao, W. & Furdui, C. M. Effects of ionizing radiation on biological molecules–mechanisms of damage and emerging methods of detection. Antioxid. Redox. Signal. 21, 260–292 (2014).
Thomas, J. A. et al. Impaired cytokine signaling in mice lacking the IL-1 receptor-associated kinase. J. Immunol. 163, 978–984 (1999).
Liou, Y. C. et al. Role of the prolyl isomerase Pin1 in protecting against age-dependent neurodegeneration. Nature 424, 556–561 (2003).
Girardini, J. E. et al. A Pin1/mutant p53 axis promotes aggressiveness in breast cancer. Cancer Cell 20, 79–91 (2011).
Anderson, J. R., Cain, K. C. & Gelber, R. D. Analysis of survival by tumor response. J. Clin. Oncol. 1, 710–719 (1983).
Braselmann, S. et al. R406, an orally available spleen tyrosine kinase inhibitor blocks Fc receptor signaling and reduces immune complex-mediated inflammation. J. Pharmacol. Exp. Ther. 319, 998–1008 (2006).
Fujiki, H. et al. Anticarcinogenic effects of (–)-epigallocatechin gallate. Prev. Med. 21, 503–509 (1992).
Notredame, C., Higgins, D. G. & Heringa, J. T-Coffee: a novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 302, 205–217 (2000).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Eramian, D., Eswar, N., Shen, M. Y. & Sali, A. How well can the accuracy of comparative protein structure models be predicted? Protein Sci. 17, 1881–1893 (2008).
Ung, P. M. & Schlessinger, A. DFGmodel: predicting protein kinase structures in inactive states for structure-based discovery of type-II inhibitors. ACS Chem. Biol. 10, 269–278 (2015).
Sherman, W., Day, T., Jacobson, M. P., Friesner, R. A. & Farid, R. Novel procedure for modeling ligand/receptor induced fit effects. J. Med. Chem. 49, 534–553 (2006).
Ye, H. et al. Distinct molecular mechanism for initiating TRAF6 signalling. Nature 418, 443–447 (2002).
Langheinrich, U., Hennen, E., Stott, G. & Vacun, G. Zebrafish as a model organism for the identification and characterization of drugs and genes affecting p53 signaling. Curr. Biol. 12, 2023–2028 (2002).
Thompson, R. et al. An inhibitor of PIDDosome formation. Mol. Cell 58, 767–779 (2015).
Wei, S. et al. Active Pin1 is a key target of all-trans retinoic acid in acute promyelocytic leukemia and breast cancer. Nat. Med. 21, 457–466 (2015).
Yaffe, M. B. et al. Sequence-specific and phosphorylation-dependent proline isomerization: a potential mitotic regulatory mechanism. Science 278, 1957–1960 (1997).
Niesen, F. H., Berglund, H. & Vedadi, M. The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability. Nat. Protoc. 2, 2212–2221 (2007).
Chou, T. C. Theoretical basis, experimental design, and computerized simulation of synergism and antagonism in drug combination studies. Pharmacol. Rev. 58, 621–681 (2006).
Kawata, M. et al. Additive effect of radiosensitization by 2-deoxy-D-glucose delays DNA repair kinetics and suppresses cell proliferation in oral squamous cell carcinoma. J. Oral Pathol. Med. 46, 979–985 (2017).
Acknowledgements
The authors thank E. Farias, R. Cagan, A. Dar, R. Krauss, J. Ashwell, X. Li, M. Posner, P. Reddy, R. De Vita and R. Sanchez for helpful comments, technical advice and reagents, and C. Franco and D. Dominguez for zebrafish care. This work was supported in part by the following bodies: NIH/NCI (F30CA186448 to P.H.L.; P30 CA008748 to K.S.P.; RO1CA168485 to H.D.S. and J.V.H; and RO1CA178162 to S.S.); the National Cancer Center NCC CCP (postdoctoral fellowship (to Y.L.); and the JJR Foundation, the Pershing Square Sohn Cancer Research Alliance, the New York Community Trust and the Searle Scholars Program (to S.S.).
Author information
Authors and Affiliations
Contributions
P.H.L. performed the following experiments: zebrafish drug screen with the assistance of J.M.B and V.B.; all zebrafish experiments with the exception of the TUNEL and caspase-3 assays (V.B.), and the p53atgMO, myd88MO, and hIRAK1 WT, K/R and E3A rescue assays (Y.L); and SEA analyses under the supervision of A.S. and S.S. R.B.S. performed all the cell culture assays with the assistance of P.H.L. and R.T., and analysed the data with P.H.L., Y.L. and S.S. R.G.M. purchased the radioresistant cell lines and assisted with translational analyses with J.A.A.-G. E.d.S., R.M.W., J.A.A.-G. and S.S. conceived the xenograft study, which was performed by E.d.S. and analysed by R.R. A.A. performed TCGA analyses under the supervision of K.S.P. and analysed the data with R.M.W. and S.S. P.M.-U.U. performed docking studies under the supervision of A.S. A.G. generated the IRAK1 KO under the supervision of J.M.S. S.K. and X.Z.Z. performed the in vitro isomerase assays under the supervision of K.P.L. H.D.S. analysed PIN1 expression levels in the MD Anderson Cancer Center HNSCC cohort with the assistance of J.V.H. S.S. conceived the study, supervised the project and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Extended data for Main Fig. 1.
(a) Representative images of 25 hpf embryos stained with AO, with conditions indicated, 7 hpIR (15 Gy Cs γ-IR TBI delivered at 18 hpf). Scale bar, 0.5 mm. (b) Quantification of AO positivity in the neural tubes of embryos from panel (a). Number of quantified embryos were: p53+/+, DMSO-treated: n of non-irradiated = 7, n of irradiated = 7; p53+/+, Gö6976-treated: n of non-irradiated = 6, n of irradiated = 8. p53MK/MK, DMSO-treated: n of non-irradiated = 7, n of irradiated = 8; p53MK/MK, Gö6976-treated: n of non-irradiated = 7, n of irradiated = 8; altogether obtained from 2 independent experiments. Data expressed as means ± SD, ***P < 0.0001, n.s., not significant, two-tailed Student’s t-test. See Supplementary Table 4 for statistics source data including precise P values. (c) Schematic of primary screen. Embryos arrayed in multi-well plates and treated with drug libraries at 17 hpf, irradiated (15 Gy) at 18 hpf, drug-treated for 6 hr, washout at 24 hpf, and scored for DTCs at 120 hpf. (d) Primary screen results. 640 FDA-approved drugs (X-axis, 2 μg/mL each) scored for DTC penetrance (Y-axis) in >12 embryos/drug. Gö6976, used as positive control, labeled in red. Drugs producing >75% DTC were considered primary hits and carried over to secondary screen. (e) Secondary screen. For each primary hit in (d), embryos were arrayed in a similar schema but half of embryos were non-irradiated. Results plotted in (f-g) below. (f) Schematic of 136 drugs screened in secondary screen (e). Quadrants determined based on greater than or less than 50% potency (%DTC +IR) and greater or less than a selectivity ratio [(%DTC +IR)/(%DTC –IR)] of 4:1. Quadrant I: selective but not potent; II: both potent and selective; III: potent but not selective; IV: neither potent nor selective. Identity of each compound in the screen described in Supplementary Tables 1-2. (g) Each drug was investigated by SEA for possibly targeting IRAK1/4 or PIN1. Red, predicted to bind IRAK1/4; blue, predicted to bind PIN1/4; purple, predicted to bind both IRAK1/4 and PIN1/4. Additional, possibly lethal off-targets were not investigated.
Supplementary Figure 2 Extended data for Main Figs. 1 and 2.
(a) Representative images of anterior spinal cords (as depicted on top) of WT and p53MK/MK embryos injected with standard control MO (std MO) or p53 MO targeted to initiator ATG (p53 MO (atg)) and treated with 15 Gy IR with or without oxfendazole (20 µg/ml) at 18 hpf. Embryos stained with AO at 7.5 hpIR. (b) Quantification of spinal cord areas shown in (a). Data represented as means ± SD of n = 3 independent experiments with at least 8 spinal chord images analyzed per condition; ***, P < 0.0005, ns, non-significant, two-tailed student’s t test. See Supplementary Table 4 for statistics source data including precise P values. (c-d) Pooled embryonic lysates analyzed by western blot with indicated antibodies. (e) Confocal microscopy of p53MK/MK zebrafish tails stained with DAPI (blue) and anti-γH2AX (green) after 0 Gy or 15 Gy IR and DMSO vs oxfendazole (20 µg/ml). Arrowheads indicates positively staining cells. hpIR, hours post ionizing radiation. hpf, hours post fertilization. (f) HeLa cells treated with indicated IRAK1 inhibitors or PIN1 inhibitor (EGCG) were irradiated (10 Gy), harvested at indicated time points and analyzed by western blot with indicated antibodies. (g) Acridine orange (AO) staining of 48 hpf embryos after 0 or 15 Gy IR and increasing doses of oxfendazole, with magnified views of spinal cords shown below. (h) Magnified images of spinal cords stained with AO at 6 hpIR, 18 hpIR, and 30 hpIR. (i) Oxfendazole (20 µg/ml) suppression of R-RT requires a 4-hr post-IR window as shown through a temporal requirement experiment. Readouts of R-RT observed as penetrance (%, absolute numbers) after AO staining at 48 hpf and DTC at 120 hpf with drug incubation windows depicted in green. (j) Quantification of DTCs in >12 p53MK/MK embryos at 120 hpf treated with inhibitors to SEA-predicted oxfendazole targets and irradiated at 18 hpf. Highest doses tested here tested in Fig. 2b. Shown are means of n = 2 independent experiments. Scale bars, 0.2 (a,g-h) and 0.1 (e) mm. See Supplementary Fig. 8 for unprocessed immunoblots.
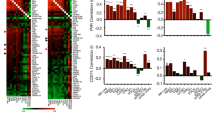
Supplementary Figure 3 IRAK1 and PIN1 mRNA expression levels in HNSCC and BC TCGA cohorts.
The Cancer Genome Atlas (TCGA) analysis shows the relative mRNA levels of IRAK1 (a-d, i-l) and PIN1 (e-h, m-p) in HNSCC (a-h) and BC (i-p) as a function of TP53 status (a, e, i, m), future RT treatment or lack thereof (b, f, j, n), or both TP53 status and future RT data sets (c-d, g-h, k-l, o-p). The n number of independent patient samples is indicated in each panel. *P < 0.05, **P < 0.005, ***P < 0.0001, NS, not significant; two sided Wilcoxon rank-sum test. See Supplementary Table 4 for statistics source data including precise P values.
Supplementary Figure 4 Extended data for main Figs. 4 and 5.
(a-b) AlamarBlue survival curves of SAS, MDA-MB-231 and CAL27 shTRIPZ (control) and shIRAK1 cells (a) and IRAK1/4i-treated HeLa cells (b) at indicated IR doses. Western blots showing shRNA knockdown efficacy also shown in (a). All data represented as means ± SD, n = 3 independent experiments performed in triplicate, *P < 0.05, **P < 0.005, ***P < 0.0005, two-tailed Student’s t-test. (c-f) Clonogenic assays of HeLa (c-d), CAL27 (e) and SAS (f) cells either stably expressing IRAK1 shRNA (c) or treated with R406 (1 μg/ml, d-f) and irradiated with indicated γIR doses (Gy). Representative images shown to the left, surviving fractions to the right. All data represented as means ± SD, n = 3 independent experiments performed in triplicate, *P < 0.05, **P < 0.005, ***P < 0.0005, two-tailed Student’s t-test. (g) HeLa cells transfected with control LACZ or MyD88 siRNA and treated with IL-1β (100 ng/ml) analyzed by immunofluorescence with indicated antibodies 5 minutes after stimulation. siMyD88 abrogates recruitment of IRAK1 to cell surface, and is thus sufficient to eliminate MyD88 function. Scale bar, 20 μm. (h) Control and IRAK1 shRNA HeLa cells treated with IL-1β (100 ng/ml) or IR (10 Gy), harvested at indicated time points after stimulus and analyzed by western blot. (i) Extended data for western blot shown in Fig. 4k to show RAIDD, Chk2 phosphorylation (marker of ATM activity), total Chk2 and GAPDH levels. (j) Extended data for western blot shown in Fig. 4d to show caspase-2 cleavage levels. (k-l) Quantification and representative images of p53MK/MK embryos injected with indicated MOs, irradiated (15 Gy) at 18 hpf and scored for DTCs at 5 dpf. Data in (k) are means ± SD of n = 3 independent experiments (bars 3,4,7,8) or n = 4 independent experiments (bars 1,2,5,6). *P < 0.05; ***P < 0.0005; ns, not significant, two-tailed Student’s t-test. Scale bar, 0.5 mm. See Supplementary Table 4 and Supplementary Figure 8 for statistics source data including precise P values and unprocessed immunoblots, respectively.
Supplementary Figure 5 Extended data for main Fig. 5.
Line graph representations of alamarBlue-based cell viability assays shown in Fig. 5d-f. Indicated cancer-cell lines after treatment with cisplatin, ginsenoside Rb1, and R406 after 0, 2.5, 5, and 7.5 Gy IR. TP53 status indicated next to cell line name. Data represented as means of n = 3 independent experiments, *P < 0.05; **P < 0.005; ***P < 0.0005; ns, not significant, two-tailed Student’s t-test. See Supplementary Table 4 for statistics source data including precise P values.
Supplementary Figure 6 Extended data for main Fig. 6.
(a) PPIase, cyclophilin and FKBP12 enzymatic assays of oxfendazole (top row) and corresponding positive controls (bottom row). Fluorescein absorbance measured every 3 seconds. (b) Thermal shift assay. PIN1 alone (blue) or with indicated drugs in two replicates. Tm, melting temperature. (c) Quantification of DTCs represented in Fig. 6a; n = 4 independent experiments (bars 2 and 4) and n = 3 independent experiments (bars 1,2,5–8). (d) Quantification of individual embryos as shown in Fig. 6b. Number of quantified embryo images over 2 experiments: DMSO-treated: n of non-irradiated = 2, n of irradiated = 5; oxfendazole-treated: n of non-irradiated = 4, n of irradiated = 5; stdMO-treated: n of non-irradiated = 5, n of irradiated = 7; pin1MO-treated: n of non-irradiated = 7, n of irradiated = 7. (e) PIN1 MO dose-response curve as scored in DTC assay; n = 4 independent experiments (0.0 mM), n = 3 independent experiments (0.2 mM) and n = 2 independent experiments (0.1, 0.3 and 0.4 mM). (f-g) Dose-response DTC curves for indicated PIN1 inhibitors, >12 embryos scored per condition; n = 3 independent experiments. (h) Oxfendazole synergizes with EGCG at doses which independently do not produce DTCs. Data are means of 2 independent experiments. (i) TUNEL assay of HeLa cells after 10 Gy IR with indicated siRNA or oxfendazole; n = 3 independent experiments in triplicates. (j) Clonogenic assay of HeLa cells treated with buparvaqone (5 μg/ml) and up to 5 Gy IR; n = 5 independent experiments in triplicates. (k) shTRIPZ (control) and shPIN1 CAL27 cells treated with indicated IR doses assayed by alamarBlue at 5 dpIR; n = 3 independent experiments in triplicates. Western blot showing PIN1 knockdown also shown. (l - m) Clonogenic assays of CAL27 (l) and SAS cells (m) treated with buparvaquone (5 μg/ml); n = 5 independent experiments in triplicates. (n) Survival curves for Fig. 6f; n = 3 independent experiments in triplicates. All data represented as means ± SD, *P < 0.05, **P < 0.005, ***P < 0.0005, two-tailed Student’s t-test. See Supplementary Table 4 for statistics source data including precise P values.
Supplementary Figure 7 Extended data for Main Fig. 7.
(a) HeLa cells treated as in Fig. 7a analyzed by alamarBlue at 5 dpIR. Data expressed as means of 2 independent experiments in triplicates. CI, combination index, see (e). (b) Indicated cell lines treated and analyzed as in (a) with buparvaquone (0.5 µg/mL in BHY, 1 µg/mL in CAL27, SAS, and SW480) and R406 (0.1 µg/mL in BHY, 0.5 µg/mL in CAL27, SAS, and SW480). Data expressed as means of 2 independent experiments in triplicates. CI, see (g). (c-d) Clonogenic assays of CAL27 (c) and SAS (d) cells treated with subtherapeutic concentrations of the indicated drugs alone or in combination with IR (Gy); buparvaquone: 0.5 µg/mL (BHY), 1 µg/mL otherwise; R406: 0.1 µg/mL (BHY), 0.5 µg/mL otherwise. Data represented as means ± SD of n = 3 independent experiments in triplicates. (e-g) CI values for panel a (e), Fig. 7c (f) and panel b (g). (e) Each drug tested at two doses, thereby representing four data points after treatment with 0, 2.5, or 5 Gy IR. After 2.5 or 5 Gy IR, every combination of IRAK1+PIN1 inhibitor produced CI’s <0.8–1, indicating synergy. (f) CI values for p53MK/MK embryos treated with four possible combinations of two doses of EGCG and R406. At greater than 50% fractions affected, EGCG + R406 combination showed <0.8 CI, indicating synergy. Bottom-right data point is dose combination in Fig. 7c. (g) CI values for BHY, CAL27, SAS, and SW480 cells after indicated drug/IR combinations. Each data point represents mean of n = 3 independent experiments. (h) Extended data for Fig. 7e-g showing tumor volume (top) and animal weight (bottom) assays. Data expressed as means ± SEM, n = 5 mice per group. (i) Cells treated and analyzed as in (a), but substituting TP53 WT and null HCT116 cells for HeLa. Subtherapeutic concentrations as established for HCT116 cells were: R406, 0.1 μg/ml; Gin-Rb1, 5 μg/ml; buparvaquone 0.5 μg/ml; EGCG, 5 μg/ml. Data expressed as means of 2 independent experiments in triplicates. *P <0.05, **P <0.005, ***P <0.0005, two-tailed Student’s t-test. See Supplementary Table 4 for statistics source data including precise P values.
Supplementary Figure 8 Unprocessed immunoblots.
Unprocessed immunoblots for indicated Figures panels.
Supplementary Figure 9 Unprocessed immunoblots.
Unprocessed immunoblots for indicated Figures panels.
Supplementary Figure 10 Unprocessed immunoblots.
Unprocessed immunoblots for indicated Figures panels.
Supplementary information
Supplementary Information
Supplementary Figures 1–8 and Supplementary Table legends.
Supplementary Table 1
Contents of FDA-approved library with secondary screen quadrant status.
Supplementary Table 2
Small molecule screen information.
Supplementary Table 3
Patient tumour characteristics.
Supplementary Table 4
Statistics source data.
Rights and permissions
About this article
Cite this article
Liu, P.H., Shah, R.B., Li, Y. et al. An IRAK1–PIN1 signalling axis drives intrinsic tumour resistance to radiation therapy. Nat Cell Biol 21, 203–213 (2019). https://doi.org/10.1038/s41556-018-0260-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0260-7
This article is cited by
-
Reciprocal antagonism of PIN1-APC/CCDH1 governs mitotic protein stability and cell cycle entry
Nature Communications (2024)
-
Radiation induces IRAK1 expression to promote radioresistance by suppressing autophagic cell death via decreasing the ubiquitination of PRDX1 in glioma cells
Cell Death & Disease (2023)
-
Promotion of microRNA-146a by histone deacetylase 4 silencing contributes to radiosensitization of esophageal carcinoma
Journal of Translational Medicine (2022)
-
Metabolic disorders sensitise endometrial carcinoma through endoplasmic reticulum stress
Cellular & Molecular Biology Letters (2022)
-
A comprehensive interaction study provides a potential domain interaction network of human death domain superfamily proteins
Cell Death & Differentiation (2021)