Abstract
Mutations in, and the altered expression of, epigenetic modifiers are pervasive in human tumours, making epigenetic factors attractive antitumour targets. The open-versus-closed chromatin state within the cells-of-origin of cancer correlates with the uneven distribution of mutations. However, the long-term effect of targeting epigenetic modifiers on mutability in patients with cancer is unclear. Here, we increased chromatin accessibility by deleting the histone H3 lysine 9 (H3K9) methyltransferase G9a in murine epidermis and show that this does not alter the single nucleotide variant burden or global genomic distribution in chemical mutagen-induced squamous tumours. G9a-depleted tumours develop after a prolonged latency compared with their wild-type counterparts, but are more aggressive and have an expanded cancer progenitor pool, pronounced genomic instability and frequent loss-of-function p53 mutations. Thus, we call for caution when assessing long-term therapeutic benefits of chromatin modifier inhibitors, which may promote more aggressive disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Whole-genome expression, ATAC-seq and WES data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE99956. WGS data that support the findings of this study are available from the Sequence Read Archive (SRA) under accession code SRP133918. The epigenomic data for seven normal tissues (epidermis, heart, intestine, kidney, liver, lung and stomach) that describe local levels of various histone marks and DNase hypersensitivity were downloaded from https://www.encodeproject.org. Secondary accessions are as follows: DHS (ENCFF513QAB, ENCFF650OFZ, ENCFF299KAN, ENCFF890IPV, ENCFF540VTK, ENCFF417SAZ, ENCFF197TQR); H3K36me3 (ENCFF256PWZ, ENCFF085HHF, ENCFF103TPF, ENCFF706SAT, ENCFF555EBK, ENCFF165DVJ, ENCFF307IYH); H3K9me3 (ENCFF485MEK, ENCFF363IUI, ENCFF664YAD, ENCFF033IOU, ENCFF040LFN, ENCFF399KQJ, ENCFF591NXE). Replication time data were from ReplicationDomain.com (Int52769503, Ext42528275, Int20705995, Int61896107) and https://www.encodeproject.org (ENCFF001JTQ, ENCFF001JTL, ENCFF001JVQ). Source data for Figs. 1–6 and Supplementary Figs. 1–4 and 6 have been provided as Supplementary Table 8. All other data supporting the findings of this study are available from the corresponding authors upon reasonable request.
References
Laugesen, A. & Helin, K. Chromatin repressive complexes in stem cells, development, and cancer. Cell Stem Cell 14, 735–751 (2014).
Vavouri, T. & Lehner, B. Human genes with CpG island promoters have a distinct transcription-associated chromatin organization. Genome Biol. 13, R110 (2012).
Avgustinova, A. & Benitah, S. A. Epigenetic control of adult stem cell function. Nat. Rev. Mol. Cell Biol. 17, 643–658 (2016).
Ezhkova, E. et al. EZH1 and EZH2 cogovern histone H3K27 trimethylation and are essential for hair follicle homeostasis and wound repair. Gene Dev. 25, 485–498 (2011).
Adam, R. C. et al. Pioneer factors govern super-enhancer dynamics in stem cell plasticity and lineage choice. Nature 521, 366–370 (2015).
Ge, Y. et al. Stem cell lineage infidelity drives wound repair and cancer. Cell 169, 636–650 (2017).
Tough, D. F., Lewis, H. D., Rioja, I., Lindon, M. J. & Prinjha, R. K. Epigenetic pathway targets for the treatment of disease: accelerating progress in the development of pharmacological tools: IUPHAR Review 11. Br. J. Pharmacol. 171, 4981–5010 (2014).
Ahuja, N., Sharma, A. R. & Baylin, S. B. Epigenetic therapeutics: a new weapon in the war against cancer. Annu. Rev. Med. 67, 73–89 (2016).
Avgustinova, A. & Benitah, S. A. The epigenetics of tumour initiation: cancer stem cells and their chromatin. Curr. Opin. Genet. Dev. 36, 8–15 (2016).
You, J. S. & Jones, P. A. Cancer genetics and epigenetics: two sides of the same coin? Cancer Cell. 22, 9–20 (2012).
Mohammad, F. et al. EZH2 is a potential therapeutic target for H3K27M-mutant pediatric gliomas. Nat. Med. 23, 483–492 (2017).
Piunti, A. et al. Therapeutic targeting of polycomb and BET bromodomain proteins in diffuse intrinsic pontine gliomas. Nat. Med. 23, 493–500 (2017).
Dawson, M. A. et al. Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia. Nature 478, 529–533 (2011).
Rathert, P. et al. Transcriptional plasticity promotes primary and acquired resistance to BET inhibition. Nature 525, 543–547 (2015).
Zuber, J. et al. RNAi screen identifies Brd4 as a therapeutic target in acute myeloid leukaemia. Nature 478, 524–528 (2011).
Filippakopoulos, P. & Knapp, S. Targeting bromodomains: epigenetic readers of lysine acetylation. Nat. Rev. Drug Discov. 13, 337–356 (2014).
Schuster-Bockler, B. & Lehner, B. Chromatin organization is a major influence on regional mutation rates in human cancer cells. Nature 488, 504–507 (2012).
Polak, P. et al. Reduced local mutation density in regulatory DNA of cancer genomes is linked to DNA repair. Nat. Biotechnol. 32, 71–75 (2014).
Polak, P. et al. Cell-of-origin chromatin organization shapes the mutational landscape of cancer. Nature 518, 360–364 (2015).
Woo, Y. H. & Li, W. H. DNA replication timing and selection shape the landscape of nucleotide variation in cancer genomes. Nat. Commun. 3, 1004 (2012).
Supek, F. & Lehner, B. Differential DNA mismatch repair underlies mutation rate variation across the human genome. Nature 521, 81–84 (2015).
Liu, L., De, S. & Michor, F. DNA replication timing and higher-order nuclear organization determine single-nucleotide substitution patterns in cancer genomes. Nat. Commun. 4, 1502 (2013).
Wen, B., Wu, H., Shinkai, Y., Irizarry, R. A. & Feinberg, A. P. Large histone H3 lysine 9 dimethylated chromatin blocks distinguish differentiated from embryonic stem cells. Nat. Genet. 41, 246–250 (2009).
Sampath, S. C. et al. Methylation of a histone mimic within the histone methyltransferase G9a regulates protein complex assembly. Mol. Cell 27, 596–608 (2007).
Schaefer, A. et al. Control of cognition and adaptive behavior by the GLP/G9a epigenetic suppressor complex. Neuron 64, 678–691 (2009).
Chen, X. et al. G9a/GLP-dependent histone H3K9me2 patterning during human hematopoietic stem cell lineage commitment. Gene Dev. 26, 2499–2511 (2012).
Mozzetta, C. et al. The histone H3 lysine 9 methyltransferases G9a and GLP regulate polycomb repressive complex 2-mediated gene silencing. Mol. Cell 53, 277–289 (2014).
Lehnertz, B. et al. The methyltransferase G9a regulates HoxA9-dependent transcription in AML. Gene Dev. 28, 317–327 (2014).
Dong, C. et al. Loss of FBP1 by Snail-mediated repression provides metabolic advantages in basal-like breast cancer. Cancer Cell 23, 316–331 (2013).
Hua, K. T. et al. The H3K9 methyltransferase G9a is a marker of aggressive ovarian cancer that promotes peritoneal metastasis. Mol. Cancer 13, 189 (2014).
Dong, C. et al. G9a interacts with Snail and is critical for Snail-mediated E-cadherin repression in human breast cancer. J. Clin. Invest. 122, 1469–1486 (2012).
Chen, M. W. et al. H3K9 histone methyltransferase G9a promotes lung cancer invasion and metastasis by silencing the cell adhesion molecule Ep-CAM. Cancer Res. 70, 7830–7840 (2010).
Liu, S. et al. G9a is essential for EMT-mediated metastasis and maintenance of cancer stem cell-like characters in head and neck squamous cell carcinoma. Oncotarget 6, 6887–6901 (2015).
Latil, M. et al. Cell-type-specific chromatin states differentially prime squamous cell carcinoma tumor-initiating cells for epithelial to mesenchymal transition. Cell Stem Cell 20, 191–204 (2017).
Malanchi, I. et al. Interactions between cancer stem cells and their niche govern metastatic colonization. Nature 481, 85–89 (2011).
Lapouge, G. et al. Identifying the cellular origin of squamous skin tumors. Proc. Natl Acad. Sci. USA 108, 7431–7436 (2011).
Schober, M. & Fuchs, E. Tumor-initiating stem cells of squamous cell carcinomas and their control by TGF-beta and integrin/focal adhesion kinase (FAK) signaling. Proc. Natl Acad. Sci. USA 108, 10544–10549 (2011).
Nassar, D., Latil, M., Boeckx, B., Lambrechts, D. & Blanpain, C. Genomic landscape of carcinogen-induced and genetically induced mouse skin squamous cell carcinoma. Nat. Med. 21, 946–954 (2015).
McCreery, M. Q. et al. Evolution of metastasis revealed by mutational landscapes of chemically induced skin cancers. Nat. Med. 21, 1514–1520 (2015).
Pickering, C. R. et al. Mutational landscape of aggressive cutaneous squamous cell carcinoma. Clin. Cancer Res. 20, 6582–6592 (2014).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Alexandrov, L. B., Nik-Zainal, S., Wedge, D. C., Campbell, P. J. & Stratton, M. R. Deciphering signatures of mutational processes operative in human cancer. Cell Rep. 3, 246–259 (2013).
Hoang, M. L. et al. Mutational signature of aristolochic acid exposure as revealed by whole-exome sequencing. Sci. Transl Med. 5, 197ra102 (2013).
Schmeiser, H. H., Bieler, C. A., Wiessler, M., van Ypersele de Strihou, C. & Cosyns, J. P. Detection of DNA adducts formed by aristolochic acid in renal tissue from patients with Chinese herbs nephropathy. Cancer Res. 56, 2025–2028 (1996).
Chakravarti, D., Pelling, J. C., Cavalieri, E. L. & Rogan, E. G. Relating aromatic hydrocarbon-induced DNA adducts and c-H-ras mutations in mouse skin papillomas: the role of apurinic sites. Proc. Natl Acad. Sci. USA 92, 10422–10426 (1995).
Alexandrov, L. B. et al. Clock-like mutational processes in human somatic cells. Nat. Genet. 47, 1402–1407 (2015).
Kim, J. et al. Somatic ERCC2 mutations are associated with a distinct genomic signature in urothelial tumors. Nat. Genet. 48, 600–606 (2016).
Supek, F. & Lehner, B. Clustered mutation signatures reveal that error-prone DNA repair targets mutations to active genes. Cell 170, 534–547 (2017).
Zheng, C. L. et al. Transcription restores DNA repair to heterochromatin, determining regional mutation rates in cancer genomes. Cell Rep. 9, 1228–1234 (2014).
Li, F. et al. The histone mark H3K36me3 regulates human DNA mismatch repair through its interaction with MutSalpha. Cell 153, 590–600 (2013).
Boumahdi, S. et al. SOX2 controls tumour initiation and cancer stem-cell functions in squamous-cell carcinoma. Nature 511, 246–250 (2014).
Huang, Y., Gu, L. & Li, G. M. H3K36me3-mediated mismatch repair preferentially protects actively transcribed genes from mutation. J. Biol. Chem. 293, 7811–7823 (2018).
Linardopoulos, S. et al. Deletion and altered regulation of p16INK4a and p15INK4b in undifferentiated mouse skin tumors. Cancer Res. 55, 5168–5172 (1995).
Rinaldi, L. et al. Loss of Dnmt3a and Dnmt3b does not affect epidermal homeostasis but promotes squamous transformation through PPAR-gamma.eLife 6, e21697 (2017).
Abel, E. L., Angel, J. M., Kiguchi, K. & DiGiovanni, J. Multi-stage chemical carcinogenesis in mouse skin: fundamentals and applications. Nat. Protoc. 4, 1350–1362 (2009).
Jensen, K. B., Driskell, R. R. & Watt, F. M. Assaying proliferation and differentiation capacity of stem cells using disaggregated adult mouse epidermis. Nat. Protoc. 5, 898–911 (2010).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Sugiura, M. et al. Induced pluripotent stem cell generation-associated point mutations arise during the initial stages of the conversion of these cells. Stem Cell Reports 2, 52–63 (2014).
Bouaoun, L. et al. TP53 variations in human cancers: new lessons from the IARC TP53 database and genomics data. Hum. Mutat. 37, 865–876 (2016).
Gonzalez-Roca, E. et al. Accurate expression profiling of very small cell populations. PLoS ONE 5, e14418 (2010).
Acknowledgements
The S.A.B. Lab is supported by the European Research Council (ERC), the Government of Cataluña (via a SGR grant), the Government of Spain (MINECO), and the Fundación Botín and Banco Santander through Santander Universities. A.A. was supported by a Marie-Curie Actions Cofound postdoctoral fellowship, and A.S. was supported by a MINECO grant. F.S. is funded by the Ramon y Cajal fellowship of MINECO and by the ERC grant HYPER-INSIGHT. The IRB Barcelona is the recipient of a Severo Ochoa Award of Excellence from MINECO (Government of Spain). The authors thank V. Raker for manuscript editing.
Author information
Authors and Affiliations
Contributions
A.A. and S.A.B. designed the experiments, and A.A. carried out the majority of the experiments. A.S. and F.S. performed the computational analyses, and U.U.-U. performed the immunohistological characterization and immunoblotting of some of the samples. I.P.-R. assisted with immunoblotting, and N.P. carried out most of the histological characterization of the samples. L.S.-B. performed the histone mark characterization and real-time quantitative PCR assays. B.L. assisted in designing the first WES experiment, and A.C. and M.M. performed some of the DMBA/TPA experiments with tamoxifen-inducible G9acKO mice and dKO mice. S.A.B., A.A. and F.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Loss of G9a does not alter overall levels of H3K4me1, H3K4me3, H3K27me3 or H3K27Ac.
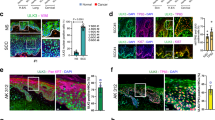
(a) Schematic of generation of G9acKO mice. (b) Immunofluorescence for the indicated histone marks (green) on WT and G9acKO skin. Nuclei are counterstained with DAPI (blue). White dashed line marks the epidermis. Yellow dashed line indicates the dermal papilla. Scale bar, 100 µm. Images shown are representative of n = 4 (WT) and n = 4 (G9acKO) independent animals. (c) Representative immunoblots for the indicated histone marks on WT or G9acKO epidermal cells. (d) Quantification of (c). Data shown are mean ± s.e.m. of individual animals. WT, n = 5 independent animals for H3K4me1 and H3K27me3, n = 4 independent animals for H3K4me3, n = 3 for H3K27Ac. G9acKO, n = 6 independent animals for H3K4me1 and H3K27me3, n = 5 independent animals for H3K4me3, n = 4 for H3K27Ac. Two-sided Student´s t-test. (e) Sorting strategy for isolation of HFSCs and EpSCs. (f) Location of differentially open ATAC-seq regions of WT or G9acKO EpSCs or HFSCs annotated by genomic feature. Data shown are mean ± s.e.m. of individual animals. WT, n = 3 independent animals; G9acKO, n = 3 independent animals. Two-sided Fisher´s exact test with FDR < 0.05 to correct for multiple testing. (g) Location of differentially open ATAC-seq regions of WT or G9acKO EpSCs or HFSCs with respect to the closest TSS. Data shown are mean ± s.e.m. of individual animals. WT, n = 3 independent animals; G9acKO, n = 3 independent animals. Two-sided Fisher´s exact test with FDR < 0.05 to correct for multiple testing. EpSCs, epidermal stem cells; HFSCs, hair follicle stem cells; TSS, transcription start site. See Supplementary Table 8 for source data
Supplementary Figure 2 G9a is dispensable for epidermal development and maintenance.
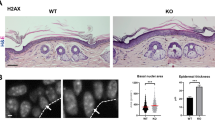
(a) Heatmap showing unsupervised hierarchical clustering of top 1000 expressed genes between WT and G9acKO HFSCs or EpSCs isolated from homeostatic epidermis. (b) Venn diagrams indicating genes commonly down- or up-regulated in G9acKO HFSCs or EpSCs as compared to WT. (c,d) Representative FACS plots showing frequency of CD45-positive cells within WT or G9acKO epidermis, with quantification. Data shown are mean ± s.e.m. WT, n = 5 independent animals; G9acKO, n = 4 independent animals. Two-sided Student´s t-test. (e,f) Immunostaining for CD3 of WT and G9acKO skin to assess infiltration of lymphocytes, with quantification. Scale bar, 50 µm. Data shown are mean ± s.e.m. WT, n = 5 independent animals; G9acKO, n = 5 independent animals. Two-sided Student´s t-test. (g) Haematoxylin and eosin staining on WT or G9acKO skin at 15 or 90 weeks of age. Scale bar, 50 µm. (h) Quantification of interfollicular epidermis thickness of WT and G9acKO skin at 15 or 90 weeks of age. Data shown are mean ± s.e.m. WT, n = 3 independent animals; G9acKO, n = 3 independent animals. Data points indicate fields of view. Two-sided Student´s t-test. (i) Distance between hair follicles of WT and G9acKO skin at 15 or 90 weeks of age. Data shown are mean ± s.e.m. WT, n = 3 independent animals; G9acKO, n = 3 independent animals. Data points indicate fields of view. Two-sided Student´s t-test. (j,k) Immunostaining for Ki67 of WT and G9acKO skin to assess proliferation rate, with quantification of positive cells within hair follicles or interfollicular epidermis. Scale bar, 50 µm. Data shown are mean ± s.e.m. WT, n = 3 independent animals; G9acKO, n = 3 independent animals. Data points indicate fields of view. Two-sided Student´s t-test. (l,m) Representative FACS plots showing frequency of HFSCs and EpSCs populations within WT or G9acKO epidermis with quantification. Data shown are mean ± s.e.m. WT, n = 13 independent animals; G9acKO, n = 13 independent animals. Two-sided Student´s t-test. EpSCs, epidermal stem cells; HFSCs, hair follicle stem cells. See Supplementary Table 8 for source data
Supplementary Figure 3 Ablation of G9a decreases tumour burden due to decreased proliferation and increased apoptosis in pre-neoplastic epidermis.
(a) Representative images of two WT or two G9acKO mice. WT, n = 8; G9acKO, n = 18 animals were monitored. WT animals had to be sacrificed at 6 months after treatment. G9acKO animals are shown through time until 12 months of follow-up. (b) Sorting strategy for tumour cells. (c) Strategy for isolation of pre-neoplastic epidermis. Epidermis is FACS-sorted to isolate HFSCs and EpSCs after six DMBA and six TPA treatments, before macroscopic tumours were visible. (d) Heatmap showing unsupervised hierarchical clustering of top 1000 expressed genes between WT and G9acKO pre-neoplastic epidermis. (e) Venn diagrams indicating genes commonly down- or up-regulated in G9acKO pre-neoplastic HFSCs and EpSCs as compared to WT. (f) Venn diagram indicating genes up-regulated in hair follicle and EpSCs from both homeostatic and pre-neoplastic epidermis. (g-h) Immunostaining for Ki67 of WT and G9acKO pre-neoplastic epidermis, with quantification. Scale bar, 50 µm. Data shown are mean ± s.e.m. WT, n = 4 individual animals; G9acKO, n = 4 individual animals. Data points indicate fields of view. Two-sided Student´s t-test. (i-j) Immunostaining for cleaved caspase 3 of WT and G9acKO pre-neoplastic epidermis with quantification. Scale bar, 50 µm. Data shown are mean ± s.e.m. WT, n = 4 individual animals; G9acKO, n = 4 individual animals. Data points indicate fields of view. Two-sided Student´s t-test. EpSCs, epidermal stem cells; HFSCs, hair follicle stem cells. See Supplementary Table 8 for source data
Supplementary Figure 4 Whole exome and whole genome sequencing of WT and G9aKO squamous cell carcinomas.
(a) Haematoxylin and eosin staining of all nine WT and eight G9acKO SCCs used for whole exome sequencing. Scale bar, 100 µm. (b) Coverage of Ehmt2 (G9a) in WT and G9acKO SCCs by whole exome sequencing data. Red box highlights exons 25 and 26, which are deleted in G9acKO epidermis. (c) Frequency of different types of SNVs found in WT and G9acKO DMBA/TPA-induced SCCs by tumour. (d) Individual 96-element substitution profiles from whole exome sequencing of WT and G9acKO SCCs. (e) Coverage of Ehmt2 (G9a) in WT and G9acKO SCCs by WGS data. Red box highlights exons 25 and 26, which are deleted in G9acKO epidermis. (f) Number of SNVs per tumour by genotype. Data shown are mean ± s.e.m. WT, n = 3 independent SCCs; G9acKO, n = 3 independent SCCs. (g) Frequency of different types of single nucleotide substitutions found in WES versus WGS data of tumour #20. (h) Individual 96-element substitution profiles from whole genome sequencing of WT and G9acKO SCCs. SCC, squamous cell carcinoma; SNV, single nucleotide variant; WGS, whole genome sequencing; WES, whole exome sequencing. See Supplementary Table 8 for source data
Supplementary Figure 5 Genomic distribution of SNVs is largely unaltered in G9acKO SCCs.
(a) Heatmap showing unsupervised hierarchical clustering of top 1000 expressed genes between WT and G9acKO tumour progenitors. WT, n = 6 independent SCCs; G9acKO, n = 7 independent SCCs. (b, c) Enrichment in SNV densities in genomic regions with high levels of DHS, H3K36me3, H3K9me3 and RepliChip signal (b) or controlling for epidermal ATAC-seq data from WT or G9acKO epidermis (c). Enrichment is adjusted for the shown variables and trinucleotide content in the genomic regions. Data for A > T transversions (the predominant substitution in DMBA/TPA-induced SCCs) or not A > T transversions are shown. Log enrichments are coefficients from negative binomial regression and their 95% CI, shown in base 2. Enrichments are relative to the lowest genomic bin of each feature (bin 0, below-baseline signal, see methods). (d) Mutation rate of Trp53 locus in not altered in G9acKO SCCs after controlling for replication time. Log enrichments are coefficients from negative binomial regression and their 95% CI, shown in base 2. Enrichments are relative to the genomic bin 0 (see below), which therefore has log2 enrichment = 0 by definition. Bin 3 is the entire Trp53 gene plus flanking 1 Mb (+/- 500 kb to each side); bin 2 is the entire Trp53 gene plus flanking 5 Mb, excluding bin 3; bin 1 is the entire Trp53 gene plus flanking 10 Mb, excluding bin 3 and bin 2; bin 0 is the remainder of the mm9 genome, excluding bins 3, 2 and 1. Trp53 exons + 5nt neighbouring exons were excluded from all analyses in this plot. Genome 37 did not have sufficient mutations within the examined area to perform the regression analysis. Log enrichments for b, c and d are coefficients from negative binomial regression and their 95% CI, shown in base 2. Enrichments are relative to the lowest genomic bin of each feature (bin 0, below-baseline signal, see methods), which thus by definition has log enrichment = 0 and is not shown on plots. DHS, DNase hypersensitivity; SCC, squamous cell carcinoma; SNV, single nucleotide variant
Supplementary Figure 6 Co-ablation of p53 and G9a in the epidermis rescues G9acKO homeostatic epidermis phenotypes.
(a) Haematoxylin and eosin staining on WT, dKO or p53cKO skin at 15 weeks of age. Scale bar, 50 µm. (b) Quantification of epidermal thickness of skin of the indicated genotypes at 15 weeks of age. Data shown are mean ± s.e.m. of individual animals. WT, n = 7 independent animals; G9acKO, n = 5 independent animals; p53cKO, n = 4 independent animals. Two-sided Student´s t-test. (c) Distance between hair follicles of WT, dKO or p53cKO skin at 15 weeks of age. Data shown are mean ± s.e.m. of individual animals. WT, n = 7 independent animals; G9acKO, n = 5 independent animals; p53cKO = 3 independent animals. Two-sided Student´s t-test. (d, e) Immunostaining for Ki67 of WT, dKO and p53cKO skin to assess proliferation rate, with quantification of positive cells within the interfollicular epidermis. Scale bar, 50 µm. Data shown are mean ± s.e.m. of individual animals. WT, n = 3 independent animals; G9acKO, n = 3 independent animals. Two-sided Student´s t-test. (f-g) Representative FACS plots showing frequency of HFSC and EpSC populations within WT, dKO or p53cKO epidermis with quantification. Data shown are mean ± s.e.m. of individual animals. WT, n = 14; cKO, n = 11 independent animals, p53cKO independent animals, n = 4 independent animals. Two-sided Student´s t-test. See Supplementary Table 8 for source data
Supplementary Figure 7 Uncropped immunoblots from figures.
The red boxes highlight the section of the immunoblots depicted in the figures
Supplementary information
Supplementary Information
Supplementary Figures 1–7, and Supplementary Table titles and legends.
Supplementary Table 1
Analysed whole-genome expression 2 data of homeostatic epidermis.
Supplementary Table 2
Analysed whole-genome expression data of pre-neoplastic epidermis.
Supplementary Table 3
Analysed whole-genome expression data of tumour progenitors.
Supplementary Table 4
GAGE pathway analysis of tumour progenitors.
Supplementary Table 5
Data matrix showing the mutational status of known cutaneous SCC-driver genes.
Supplementary Table 6
Specific amino acid substitutions within p53 in G9acKO tumours.
Supplementary Table 7
Specifications of commercially available antibodies used in this study.
Supplementary Table 8
Statistics source data in this paper.
Rights and permissions
About this article
Cite this article
Avgustinova, A., Symeonidi, A., Castellanos, A. et al. Loss of G9a preserves mutation patterns but increases chromatin accessibility, genomic instability and aggressiveness in skin tumours. Nat Cell Biol 20, 1400–1409 (2018). https://doi.org/10.1038/s41556-018-0233-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0233-x
This article is cited by
-
Tissue memory relies on stem cell priming in distal undamaged areas
Nature Cell Biology (2023)
-
TP53-dependent toxicity of CRISPR/Cas9 cuts is differential across genomic loci and can confound genetic screening
Nature Communications (2022)
-
Resistance to chemical carcinogenesis induction via a dampened inflammatory response in naked mole-rats
Communications Biology (2022)
-
Inhibition of histone methyltransferase G9a attenuates liver cancer initiation by sensitizing DNA-damaged hepatocytes to p53-induced apoptosis
Cell Death & Disease (2021)
-
Epigenetics and beyond: targeting writers of protein lysine methylation to treat disease
Nature Reviews Drug Discovery (2021)