Abstract
Adhesion to the extracellular matrix persists during mitosis in most cell types. However, while classical adhesion complexes, such as focal adhesions, do and must disassemble to enable mitotic rounding, the mechanisms of residual mitotic cell–extracellular matrix adhesion remain undefined. Here, we identify ‘reticular adhesions’, a class of adhesion complex that is mediated by integrin αvβ5, formed during interphase, and preserved at cell–extracellular matrix attachment sites throughout cell division. Consistent with this role, integrin β5 depletion perturbs mitosis and disrupts spatial memory transmission between cell generations. Reticular adhesions are morphologically and dynamically distinct from classical focal adhesions. Mass spectrometry defines their unique composition, enriched in phosphatidylinositol-4,5-bisphosphate (PtdIns(4,5)P2)-binding proteins but lacking virtually all consensus adhesome components. Indeed, reticular adhesions are promoted by PtdIns(4,5)P2, and form independently of talin and F-actin. The distinct characteristics of reticular adhesions provide a solution to the problem of maintaining cell–extracellular matrix attachment during mitotic rounding and division.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
Data availability
MS data have been deposited at the ProteomeXchange Consortium via the PRIDE partner repository with primary accession codes PXD008645 and PXD008680. Source data for Fig. 4a–d and Supplementary Fig. 4a,b are provided as Supplementary Table 2. All other quantitative data are presented in Supplementary Table 1, while sample images from screening analyses are provided via the GitHub repository described under ‘Code availability’. All additional data supporting the findings of this study are available from the corresponding authors on reasonable request.
References
Lock, J. G., Wehrle-Haller, B. & Strömblad, S. Cell-matrix adhesion complexes: master control machinery of cell migration. Semin. Cancer Biol. 18, 65–76 (2008).
Hynes, R. O. Integrins: bidirectional, allosteric signaling machines. Cell 110, 673–687 (2002).
Berrier, A. L. & Yamada, K. M. Cell-matrix adhesion. J. Cell. Physiol. 213, 565–573 (2007).
Horton, E. R. et al. Definition of a consensus integrin adhesome and its dynamics during adhesion complex assembly and disassembly. Nat. Cell Biol. 17, 1577–1587 (2015).
Tadokoro, S. et al. Talin binding to integrin β tails: a final common step in integrin. Science 302, 103–106 (2008).
Liu, J. et al. Talin determines the nanoscale architecture of focal adhesions. Proc. Natl Acad. Sci. USA 112, E4864–E4873 (2015).
Gardel, M. L., Schneider, I. C., Aratyn-Schaus, Y. & Waterman, C. M. Mechanical integration of actin and adhesion dynamics in cell migration. Annu. Rev. Cell Dev. Biol. 26, 315–333 (2010).
Jime, A. et al. Experimental and theoretical study of mitotic spindle orientation. Nature 447, 493–496 (2007).
Théry, M. et al. The extracellular matrix guides the orientation of the cell division axis. Nat. Cell Biol. 7, 947–953 (2005).
Théry, M., Jiménez-Dalmaroni, A., Racine, V., Bornens, M. & Jülicher, F. Experimental and theoretical study of mitotic spindle orientation. Nature 447, 493–496 (2007).
Théry, M. & Bornens, M. Cell shape and cell division. Curr. Opin. Cell Biol. 18, 648–657 (2006).
Akanuma, T., Chen, C., Sato, T., Merks, R. M. H. & Sato, T. N. Memory of cell shape biases stochastic fate decision-making despite mitotic rounding. Nat. Commun. 7, 11963 (2016).
Maddox, A. S. & Burridge, K. RhoA is required for cortical retraction and rigidity during mitotic cell rounding. J. Cell Biol. 160, 255–265 (2003).
Lancaster, O. et al. Mitotic rounding alters cell geometry to ensure efficient bipolar spindle formation. Dev. Cell 25, 270–283 (2013).
Dao, V. T., Dupuy, A. G., Gavet, O., Caron, E. & de Gunzburg, J. Dynamic changes in Rap1 activity are required for cell retraction and spreading during mitosis. J. Cell Sci. 122, 2996–3004 (2009).
Petridou, N. I. & Skourides, P. A. A ligand-independent integrin β1 mechanosensory complex guides spindle orientation. Nat. Commun. 7, 10899 (2016).
LaFlamme, S. E., Nieves, B., Colello, D. & Reverte, C. G. Integrins as regulators of the mitotic machinery. Curr. Opin. Cell Biol. 20, 576–582 (2008).
Ramkumar, N. & Baum, B. Coupling changes in cell shape to chromosome segregation. Nat. Rev. Mol. Cell Biol. 17, 511–521 (2016).
Winograd-Katz, S. E., Fässler, R., Geiger, B. & Legate, K. R. The integrin adhesome: from genes and proteins to human disease. Nat. Rev. Mol. Cell Biol. 15, 273–288 (2014).
Hernández-Varas, P., Berge, U., Lock, J. G. & Strömblad, S. A plastic relationship between vinculin-mediated tension and adhesion complex area defines adhesion size and lifetime. Nat. Commun. 6, 7524 (2015).
Lock, J. G. et al. Plasticity in the macromolecular-scale causal networks of cell migration. PLoS ONE 9, e90593 (2014).
Ballestrem, C., Hinz, B., Imhof, B. A. & Wehrle-Haller, B. Marching at the front and dragging behind: differential αVβ3-integrin turnover regulates focal adhesion behavior. J. Cell Biol. 155, 1319–1332 (2001).
Besser, A. & Safran, S. A. Force-induced adsorption and anisotropic growth of focal adhesions. Biophys. J. 90, 3469–3484 (2006).
Yu, C. H. et al. Integrin-matrix clusters form podosome-like adhesions in the absence of traction forces. Cell Rep. 5, 1456–1468 (2013).
Mohl, C., Kirchgessner, N., Schafer, C., Hoffmann, B. & Merkel, R. Quantitative mapping of averaged focal adhesion dynamics in migrating cells by shape normalization. J. Cell Sci. 125, 155–165 (2012).
Spiess, M. et al. Active and inactive β1 integrins segregate into distinct nanoclusters in focal adhesions. J. Cell Biol. 217, 1929–1940 (2018).
Zhang, X. et al. Talin depletion reveals independence of initial cell spreading from integrin activation and traction. Nat. Cell Biol. 10, 1062–1068 (2008).
Cluzel, C. et al. The mechanisms and dynamics of αvβ3 integrin clustering in living cells. J. Cell Biol. 171, 383–392 (2005).
Saltel, F. et al. New PI(4,5)P2- and membrane proximal integrin-binding motifs in the talin head control 3-integrin clustering. J. Cell Biol. 187, 715–731 (2009).
Mellacheruvu, D. et al. The CRAPome: a contaminant repository for affinity purification-mass spectrometry data. Nat. Methods 10, 730–736 (2013).
Toyoshima, F. & Nishida, E. Integrin-mediated adhesion orients the spindle parallel to the substratum in an EB1- and myosin X-dependent manner. EMBO J. 26, 1487–1498 (2007).
Mali, P., Wirtz, D. & Searson, P. C. Interplay of RhoA and motility in the programmed spreading of daughter cells postmitosis. Biophys. J. 99, 3526–3534 (2010).
Held, M. et al. CellCognition: time-resolved phenotype annotation in high-throughput live cell imaging. Nat. Methods 7, 747–754 (2010).
Maupin, P. & Pollard, T. D. Improved preservation and staining of HeLa cell actin filaments, clathrin-coated membranes, and other cytoplasmic structures by tannic acid-glutaraldehyde-saponin fixation. J. Cell Biol. 96, 51–62 (1983).
Aszodi, A., Hunziker, E. B., Brakebusch, C. & Fässler, R. β1 integrins regulate chondrocyte rotation, G1 progression, and cytokinesis. Genes Dev. 17, 2465–2479 (2003).
Thullberg, M., Gad, A., Le Guyader, S. & Strömblad, S. Oncogenic H-Ras V12 promotes anchorage-independent cytokinesis. Proc. Natl Acad. Sci. USA 104, 20338–20343 (2007).
Reverte, C. G., Benware, A., Jones, C. W. & LaFlamme, S. E. Perturbing integrin function inhibits microtubule growth from centrosomes, spindle assembly, and cytokinesis. J. Cell Biol. 174, 491–497 (2006).
Cramer, L. P. & Mitchison, T. J. Investigation of the mechanism of retraction of the cell margin and rearward flow of nodules during mitotic cell rounding. Mol. Biol. Cell 8, 109–119 (1997).
Riddelle, K. S., Hopkinson, S. B. & Jones, J. C. Hemidesmosomes in the epithelial cell line 804G: their fate during wound closure, mitosis and drug induced reorganization of the cytoskeleton. J. Cell Sci. 103, 475–490 (1992).
Baker, J. & Garrod, D. Epithelial cells retain junctions during mitosis. J. Cell Sci. 104, 415–425 (1993).
Dix, C. L. et al. The role of mitotic cell-substrate adhesion re-modeling in animal cell division. Dev. Cell 45, 132–145 (2018).
Lane, N. E. et al. Mice lacking the integrin β5 subunit have accelerated osteoclast maturation and increased activity in the estrogen-deficient state. J. Bone Miner. Res. 20, 58–66 (2005).
Coudert, A. E. et al. Differentially expressed genes in autosomal dominant osteopetrosis type II osteoclasts reveal known and novel pathways for osteoclast biology. Lab. Invest. 94, 275–285 (2014).
Duperret, E. K., Dahal, A. & Ridky, T. W. Focal-adhesion-independent integrin-v regulation of FAK and c-Myc is necessary for 3D skin formation and tumor invasion. J. Cell Sci. 128, 3997–4013 (2015).
Wayner, E. A., Orlando, R. A. & Cheresh, D. A. Integrins αvβ3 and αvβ5 contribute to cell attachment to vitronectin but differentially distribute on the cell surface. J. Cell Biol. 113, 919–929 (1991).
Klapholz, B. & Brown, N. H. Talin—the master of integrin adhesions. J. Cell Sci. 130, 2435–2446 (2017).
Calderwood, D. A. et al. Integrin β cytoplasmic domain interactions with phosphotyrosine-binding domains: a structural prototype for diversity in integrin signaling. Proc. Natl Acad. Sci. USA 100, 2272–2277 (2003).
Leyton-Puig, D. et al. Flat clathrin lattices are dynamic actin-controlled hubs for clathrin-mediated endocytosis and signalling of specific receptors. Nat. Commun. 8, 16068 (2017).
Elkhatib, N. et al. Tubular clathrin/AP-2 lattices pinch collagen fibers to support 3D cell migration. Science 356, eaal4713 (2017).
De Deyne, P. G. et al. The vitronectin receptor associates with clathrin-coated membrane domains via the cytoplasmic domain of its β5 subunit. J. Cell Sci. 111, 2729–2740 (1998).
Vassilopoulos, S. et al. Actin scaffolding by clathrin heavy chain is required for skeletal muscle sarcomere organization. J. Cell Biol. 205, 377–393 (2014).
Papoulas, O., Hays, T. S. & Sisson, J. C. The golgin lava lamp mediates dynein-based Golgi movements during Drosophila cellularization. Nat. Cell Biol. 7, 612–618 (2005).
Akisaka, T., Yoshida, H., Suzuki, R., Shimizu, K. & Takama, K. Clathrin sheets on the protoplasmic surface of ventral membranes of osteoclasts in culture. J. Electron Microscop. 52, 535–543 (2003).
Saffarian, S., Cocucci, E. & Kirchhausen, T. Distinct dynamics of endocytic clathrin-coated pits and coated plaques. PLoS Biol. 7, e1000191 (2009).
Nicol, A. & Nermut, M. A new type of substratum adhesion structure in NRK cells revealed by correlated interference reflection and electron microscopy. Eur. J. Cell Biol. 43, 348–357 (1987).
Akisaka, T., Yoshida, H., Suzuki, R. & Takama, K. Adhesion structures and their cytoskeleton-membrane interactions at podosomes of osteoclasts in culture. Cell Tissue Res. 331, 625–641 (2008).
Smilenov, L., Forsberg, E., Zeligman, I., Sparrman, M. & Johansson, S. Separation of fibronectin from a plasma gelatinase using immobilized metal affinity chromatography. FEBS Lett. 302, 227–230 (1992).
Yatohgo, T., Izumi, M., Kashiwagi, H. & Hayashi, M. Novel purification of vitronectin from human plasma by heparin affinity chromatography. Cell Struct. Funct. 13, 281–292 (1988).
Kiss, A. et al. Non-monotonic cellular responses to heterogeneity in talin protein expression-level. Integr. Biol. 7, 1171–1185 (2015).
Li, Z. et al. Integrin-mediated cell attachment induces a PAK4-dependent feedback loop regulating cell adhesion through modified integrin αvβ5 clustering and turnover. Mol. Biol. Cell. 21, 3317–3329 (2010).
Bates, M., Huang, B., Dempsey, G. T. & Zhuang, X. Multicolor super-resolution imaging with photo-switchable fluorescent probes. Science 317, 1749–1753 (2007).
Shafqat-Abbasi, H. et al. An analysis toolbox to explore mesenchymal migration heterogeneity reveals adaptive switching between distinct modes. eLife 5, e11384 (2016).
Ester, M., Kriegel, H.-P., Sander, J. & Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. Proceedings of 2nd International Conference on Knowledge Discovery and Data Mining KDD-96, 226–231 (1996).
Schindelin, J. et al. Fiji: an open source platform for biological image analysis. Nat. Methods 9, 676–682 (2012).
Szklarczyk, D. et al. STRING v10: protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Wählby, C., Sintorn, I. M., Erlandsson, F., Borgefors, G. & Bengtsson, E. Combining intensity, edge and shape information for 2D and 3D segmentation of cell nuclei in tissue sections. J. Microsc. 215, 67–76 (2004).
Yebra, M. et al. Requirement of receptor-bound urokinase-type plasminogen activator for integrin αvβ5-directed cell migration. J. Biol. Chem. 271, 29393–29399 (1996).
Strömblad, S. et al. Loss of p53 compensates for αv-integrin function in retinal neovascularization. J. Biol. Chem. 277, 13371–13374 (2002).
Acknowledgements
The authors thank D. Critchley (University of Leicester) for providing talin-1-null embryonic stem cells and Z. Li for assistance with the integrin β5/β3-EGFP construct. This work was supported by grants to S.S. from the EU-FP7-Systems Microscopy Network of Excellence (HEALTH-F4-2010-258068), EU-H2020-Multimot (EU grant agreement no. 634107), the Strategic Research Foundation of Sweden (SB16-0046), the Swedish Research Council (521-2012-3180) and the Swedish Cancer Society, and by a grant to M.J.H. from Cancer Research UK (grant no. 13329/A21671). Imaging was in part performed at the live cell imaging facility and Nikon Center of Excellence at the Department of Biosciences and Nutrition, Karolinska Institutet, supported by grants from the Knut and Alice Wallenberg Foundation, the Swedish Research Council, the Centre for Innovative Medicine and the Jonasson donation to the School of Technology and Health, Royal Institute of Technology, Sweden. The authors also acknowledge support from the Bioimaging and Bio-MS core facilities at the University of Manchester. The mass spectrometers and microscopes used in this study were purchased with grants from BBSRC, Wellcome Trust and the University of Manchester Strategic Fund. The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. J.A.A. and X.G. contributed equally to this paper.
Author information
Authors and Affiliations
Contributions
J.G.L., M.C.J., J.A.A., M.L., M.J.H. and S.S. conceived the project and devised experiments. J.G.L. and J.A.A. performed live cell imaging and related image analyses. J.G.L., M.C.J., J.A.A., X.G. and H.O. performed fixed cell imaging. X.G. performed the siRNA screening for PIP regulators. M.C.J. and J.A.A. undertook experiments relating to integrin β5 RNAi and MS analyses. A.O. performed STORM imaging and related image analyses. H.O. and S.G. performed immunoblotting. J.L. performed image analyses, statistical analyses and data visualization. J.G.L., M.C.J., J.A.A., X.G., M.L., M.J.H. and S.S. contributed to writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 β5-positive, talin- or F-actin-negative structures found in multiple cell lines.
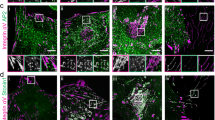
(a) A549 lung cancer cells and A375 melanoma cells plated on glass coverslips for 72 h. Confocal images of immuno-fluorescently labeled integrins αVβ3 (LM609), β1 (9EG7) and αVβ5 (15F11). (b) U2OS cells plated on glass coverslips for 24 and 48 h. Confocal images of immuno-fluorescently labeled integrin β5 and vinculin. (c,d,e) The indicated human-, mouse- and hamster-derived cancer cell lines (c), immortalized diploid cell lines (d) and primary cells (e) were found to contain central β5-positive, talin- and F-actin-negative structures equivalent to those first observed in U2OS cells. Confocal images of cells replated for 3 h on 10 µg/ml vitronectin and immuno-labeled for integrin β5 and either talin or F-actin, as indicated. All images representative of results from at least n = 3 biologically independent experiments. Scale bars: 10 µm
Supplementary Figure 2 β5-positive reticular adhesions lack consensus adhesome components.
(a-m) Confocal images of U2OS cells plated for 3 h on vitronectin and immuno-labeled against integrin β5 and: the alpha V (αV) subunit of the αVβ5 heterodimer (a); consensus adhesome components [paxilin (b), FAK (c), zyxin (d), kindlin 2 (e)]; integrin β5-binding partners [CSK (genetically encoded GFP-CSK) (f), ICAP1 (g), DOK1 (h)]; phospho-tyrosine (i); and cytoskeletal proteins [F-actin (j), cytokeratin (k), beta (β)-tubulin (l), vimentin (m)]. (n) Confocal image of U2OS cells plated for 3 h on laminin (ECM ligand not bound by αVβ5) and immuno-labeled for integrin β5 and vinculin. (o) Confocal images of an unpermeabilised U2OS cell expressing β5-2GFP, immuno-labeled for β5. All images representative of results from at least n = 3 biologically independent experiments. Scale bars: 10 µm. Boxed areas shown at higher magnification in lower right corners
Supplementary Figure 3 Validation of reticular adhesion independence from F-actin and talin.
(a, upper) Confocal images of U2OS cells attached to VN-coated surfaces and labelled for integrin β5, vinculin and F-actin, following 30 min in suspension and 3 h replating in the presence of DMSO, 20 µM CytoD, or 1 µM latrunculin A (LatA). (a, lower) Representative confocal images of U2OS cells attached to vitronectin (VN)-coated surfaces and labeled for integrin β5, vinculin and F-actin, following 3 h replating, with final 1 h in presence of DMSO or 20 µM CytoD. (b) Images comparing attachment of CS1WT or CS1β5 cells to vitronectin in the presence of RAD or RGD peptides and/or CytoD. Zoomed panels demonstrate image-based identification of attached cells using automated cell counting in NIS Elements. Representative of n = 6 biologically independent experiments. (c) Representative immunoblot of talin-2 (68e7 antibody) and tubulin following siRNA treatment with control (CTRL) oligonucleotides or one of three alternative talin-2-targeting oligonucleotides. Unprocessed blots in Supplementary Fig 9. (d) Confocal images of U2OS cells expressing RFP-talin head or RFP-talin rod domains plated on glass coverslips for 48 hours and immunolabeled for integrin β5 and vinculin. Images in A, C-D representative of results from at least n = 3 biologically independent experiments. Scale bars: A, Bvi-viii = 10 µm; Bi-ii and Bix-x = 1000 µm; Biii-v = 100 µm; D = 5 µm
Supplementary Figure 4 Validation of reticular adhesion proteins identified by mass spectrometry.
(a,b) Gene-ontology analysis of U2OS steady state adhesion proteins (DMSO treated; focal and reticular adhesions) showing terms from Biological Process (a) and KEGG pathway analysis (b) significantly enriched over whole cell proteome (based on data from n = 3 biologically independent mass spec experiments). P-values were derived from EASE scores calculated using a modified Fishers Exact test with Holm-Bonferroni correction from multiple tests using the DAVID annotation system. (c-g) Confocal images of U2OS β5 V cells attached to vitronectin (VN)-coated surfaces and immuno-labeled against EPS15L1 (c), HIP1R (d), or WASL (e), Intersectin-1 (f), and ARP3 (g). All images representative of results from at least n = 3 biologically independent experiments. Scale bars: C-E = 10 µm; F-G = 5 µm
Supplementary Figure 5 Quantitative imaging of siRNA- or Arp2/3 inhibitor-treated cells.
(a) Representative images illustrating segmentation of U2OS cells based on DAPI (nuclei) and F-actin staining and subsequent segmentation of mCherry-vinculin and integrin β5-2GFP fluorescence (shown for an individual cell; cropped). Images representative of n = 2 biologically independent experiments. Scale bars 50 µm (10 µm in cropped images). (b) Representative images of U2OS cells stained with DAPI (nuclei) and phalloidin, and immunolabelled for integrin β5 and vinculin following treatment with Arp2/3 inhibitor (CK-666) or inactive analogue control (CK-689). Scale bars = 50 µm. (c) Boxplots summarizing single cell quantification (from images as shown in B; n = 1019 cells analysed, averaging 509 + /- 23 (stdev) per condition) of reticular to focal adhesion integrin β5 ratios following treatment with Arp2/3 inhibitor (CK-666) or inactive analogue control (CK-689). Boxplot centre and box edges indicate median and 25th or 75th percentiles, respectively, while whiskers indicate the median + /- 1.5*IQR (inter-quartile range) or the most extreme observations within these limits. Boxplot notches approximate 95% confidence intervals (CI; see methods for details). P-values reflect two-sided unpaired Mann Whitney U testing. Images and data in B-C derived from 3 biologically independent experiments. Source data for C is available in Supplementary Table 1
Supplementary Figure 6 Secondary siRNA validation and effects of PI4,5P2 modulation.
(a) Single cell quantification (n = 8070 cells analysed, averaging 224 + /- 63 (stdev) per condition) of reticular to focal adhesion integrin β5 ratios for 4 independent siRNAs targeting controls or PIP regulators. Boxplot centre and box edges indicate median and 25th or 75th percentiles, respectively, while whiskers indicate the median + /- 1.5*IQR (inter-quartile range) or the most extreme observations within these limits. Boxplot notches approximate 95% confidence intervals (see methods for details). P-values reflect two-sided unpaired Mann Whitney U testing with Holm-Bonferroni correction from multiple tests. Data derived from 2 biologically independent experiments. Source data is available in Supplementary Table 1 (b) Representative images of U2OS cells stained with DAPI (nuclei) and deep red cell mask membrane dye and showing integrin β5-2GFP and mCherry-vinculin fluorescence following treatment with DMSO, 10 mM Neomycin or 25 μM LY294002. Images in B representative of n = 3 biologically independent experiments. Scale bars = 50 µm
Supplementary Figure 7 Mitotic reticular adhesions lack canonical adhesome components.
(a-e) Confocal images of mitotic U2OS cells (yellow dashed outlines) plated on VN and labeled for integrin β5 and: integrin β3 subunit (a); and consensus adhesome components [talin (b), vinculin (c), paxillin (d), zyxin (e)]. (f-g) Confocal images of mitotic U2OS cells plated on 10μg/ml laminin (f) or fibronectin (g) immuno-labeled against integrin β5 and integrin β1 subunits. Images in A-K representative of at least n = 3 biologically independent experiments. (h-k) Alternate views of actin-positive retraction filaments (phalloidin-labeled) formed during mitosis and terminating at integrin β5-2GFP-positive adhesion complexes (K cropped from yellow box in J; see Supplementary Movie 9). Images in H-K representative of n = 2 biologically independent experiments. Scale bars:10 µm except in K; 5 µm
Supplementary Figure 8 Disruption of HeLa cell mitosis by integrin β5 siRNA.
(a) Hela cells transfected with control siRNA showing normal cell division (see Supplementary Movie 10). (b) Three example microscopy time series representative of different effects of depleting β5 on cell division: i) cell remains round for an extended time before eventually dividing, often with incomplete cytokinesis (see Supplementary Movie 11). ii) cell repeatedly rounds up and respreads without dividing (see Supplementary Movie 12). iii) cell appears to divide but before cytokinesis merges back into a single bi-nucleate cell (see Supplementary Movie 13. (c) Hela cell treated with β5 siRNA and rescued with β5-EGFP (see Supplementary Movie 14). (a-c) Images were taken at 10 min intervals. Arrows point to mitotic cells of interest. Images representative of n = 3 biologically independent experiments. (d) Representative immunoblot of endogenous and β5-EGFP expression in control and knockdown cells. Unprocessed blot in Supplementary Fig 9. Scale bars = 50 µm
Supplementary Figure 9 Unprocessed images of all gels and blots.
(a) Unprocessed blots from figure SF3C. Film from talin2 blot was accidentally cut at the bottom because the background was too low to see where the membrane ended. (b) Unprocessed blot from figure SF8D
Supplementary information
Supplementary Information
Supplementary Figures 1–9, Supplementary Table and Supplementary Video legends.
Supplementary Table 1
Statistics source data.
Supplementary Table 2
Proteomic characterisation of reticular adhesions.
Supplementary Table 3
Direct and indirect PI45P2 interactors.
Supplementary Table 4
Details of antibodies used.
Supplementary Table 5
PIP regulator siRNA screen oligonucleotide sequences.
Supplementary Video 1
Reticular adhesion formation, maturation and turnover.
Supplementary Video 2
Reticular adhesions do not recruit vinculin during their lifetime.
Supplementary Video 3
Comparison of reticular and focal adhesion dynamics.
Supplementary Video 4
FRAP comparison of integrin-β5 2GFP.
Supplementary Video 5
Reticular adhesion formation during cell-ECM attachment in the presence and absence of F-actin.
Supplementary Video 6
Reticular adhesions persist throughout cell division when classical focal adhesions disassemble.
Supplementary Video 7
Mitotic retraction fibres attach to ECM at reticular adhesion sites.
Supplementary Video 8
Post-mitotic daughter cells re-spread through retraction fibres tethered to the ECM by reticular adhesions, thereby recovering the pre-mitotic footprint.
Supplementary Video 9
Mitotic retraction fibres contain dense F-actin labelling while reticular adhesion sites have only weak F-actin labelling.
Supplementary Video 10
Individual centrally located reticular adhesion complexed persist throughout mitosis.
Supplementary Video 11
Cell division in HeLa cells after synchronisation.
Supplementary Video 12
Defects in cell division after integrin-β5 knockdown.
Supplementary Video 13
Defects in cell division after integrin-β5 knockdown.
Supplementary Video 14
Defects in cell division after integrin-β5 knockdown.
Supplementary Video 15
Rescue of cell division defects by wild type integrin-β5.
Rights and permissions
About this article
Cite this article
Lock, J.G., Jones, M.C., Askari, J.A. et al. Reticular adhesions are a distinct class of cell-matrix adhesions that mediate attachment during mitosis. Nat Cell Biol 20, 1290–1302 (2018). https://doi.org/10.1038/s41556-018-0220-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0220-2
This article is cited by
-
Canonical and non-canonical integrin-based adhesions dynamically interconvert
Nature Communications (2024)
-
Talin2 and KANK2 functionally interact to regulate microtubule dynamics, paclitaxel sensitivity and cell migration in the MDA-MB-435S melanoma cell line
Cellular & Molecular Biology Letters (2023)
-
Curved adhesions mediate cell attachment to soft matrix fibres in three dimensions
Nature Cell Biology (2023)
-
In mitosis integrins reduce adhesion to extracellular matrix and strengthen adhesion to adjacent cells
Nature Communications (2023)
-
Organization, dynamics and mechanoregulation of integrin-mediated cell–ECM adhesions
Nature Reviews Molecular Cell Biology (2023)