Abstract
The intestinal epithelium harbours remarkable self-renewal capacity that is driven by Lgr5+ intestinal stem cells (ISCs) at the crypt base. However, the molecular mechanism controlling Lgr5+ ISC stemness is incompletely understood. We show that a Gata6 long noncoding RNA (lncGata6) is highly expressed in ISCs. LncGata6 knockout or conditional knockout in ISCs impairs the stemness of ISCs and epithelial regeneration. Mechanistically, lncGata6 recruits the NURF complex onto the Ehf promoter to induce its transcription, which promotes the expression of Lgr4/5 to enhance Wnt signalling activation. Moreover, the human orthologue lncGATA6 is highly expressed in the cancer stem cells of colorectal cancer and promotes tumour initiation and progression. Antisense oligonucleotides against lncGATA6 exhibit strong therapeutic efficacy on colorectal cancer. Thus, targeting lncGATA6 will have potential clinical applications in colorectal cancer treatment as an ideal therapeutic target.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Microarray data that support the findings of this study have been deposited in the Gene Expression Omnibus (GEO) under accession code GSE107959. Source data have been provided as Supplementary Table 4. All other data supporting the findings of this study are available from the corresponding author on reasonable request.
References
Huch, M. et al. In vitro expansion of single Lgr5+ liver stem cells induced by Wnt-driven regeneration. Nature 494, 247–250 (2013).
Barker, N. Adult intestinal stem cells: critical drivers of epithelial homeostasis and regeneration. Nat. Rev. Mol. Cell Biol. 15, 19–33 (2014).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Van der Flier, L. G. et al. Transcription factor achaete scute-like 2 controls intestinal stem cell fate. Cell 136, 903–912 (2009).
Van der Flier, L. G., Haegebarth, A., Stange, D. E., Van de Wetering, M. & Clevers, H. OLFM4 is a robust marker for stem cells in human intestine and marks a subset of colorectal cancer cells. Gastroenterology 137, 15–17 (2009).
Sato, T. et al. Single Lgr5 stem cells build crypt–villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009).
Clevers, H., Loh, K. M. & Nusse, R. Stem cell signaling. An integral program for tissue renewal and regeneration: Wnt signaling and stem cell control. Science 346, 1248012 (2014).
Ohlstein, B. & Spradling, A. Multipotent Drosophila intestinal stem cells specify daughter cell fates by differential notch signaling. Science 315, 988–992 (2007).
Clevers, H. The intestinal crypt, a prototype stem cell compartment. Cell 154, 274–284 (2013).
De Lau, W. et al. Lgr5 homologues associate with Wnt receptors and mediate R-spondin signalling. Nature 476, 293–297 (2011).
Batista, P. J. & Chang, H. Y. Long noncoding RNAs: cellular address codes in development and disease. Cell 152, 1298–1307 (2013).
Quinn, J. J. & Chang, H. Y. Unique features of long non-coding RNA biogenesis and function. Nat. Rev. Genet. 17, 47–62 (2016).
Wang, Y. et al. The long noncoding RNA lncTCF7 promotes self-renewal of human liver cancer stem cells through activation of Wnt signaling. Cell Stem Cell 16, 413–425 (2015).
Zhu, P. et al. lnc-beta-Catm elicits EZH2-dependent beta-catenin stabilization and sustains liver CSC self-renewal. Nat. Struct. Mol. Biol. 23, 631–639 (2016).
Zhu, P. et al. LncBRM initiates YAP1 signalling activation to drive self-renewal of liver cancer stem cells. Nat. Commun. 7, 13608 (2016).
Liu, S. J. et al. CRISPRi-based genome-scale identification of functional long noncoding RNA loci in human cells. Science 355, aah7111 (2017).
Platt, R. J. et al. CRISPR-Cas9 knockin mice for genome editing and cancer modeling. Cell 159, 440–455 (2014).
Metcalfe, C., Kljavin, N. M., Ybarra, R. & de Sauvage, F. J. Lgr5+ stem cells are indispensable for radiation-induced intestinal regeneration. Cell Stem Cell 14, 149–159 (2014).
McHugh, C. A. et al. The Xist lncRNA interacts directly with SHARP to silence transcription through HDAC3. Nature 521, 232–236 (2015).
Furuyama, K. et al. Continuous cell supply from a Sox9-expressing progenitor zone in adult liver, exocrine pancreas and intestine. Nat. Genet. 43, 34–52 (2011).
Cheasley, D. et al. Myb controls intestinal stem cell genes and self-renewal. Stem Cells 29, 2042–2050 (2011).
Yan, K. S. et al. Non-equivalence of Wnt and R-spondin ligands during Lgr5+ intestinal stem-cell self-renewal. Nature 545, 238–242 (2017).
Nassar, D. & Blanpain, C. Cancer stem cells: basic concepts and therapeutic implications. Annu. Rev. Pathol. 11, 47–76 (2016).
Barker, N. et al. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature 457, 608–611 (2009).
Melo, F. D. E. et al. A distinct role for Lgr5+ stem cells in primary and metastatic colon cancer. Nature 543, 676–680 (2017).
Shimokawa, M. et al. Visualization and targeting of LGR5+ human colon cancer stem cells. Nature 545, 187–192 (2017).
Katayama, S. et al. Antisense transcription in the mammalian transcriptome. Science 309, 1564–1566 (2005).
Anderson, K. M. et al. Transcription of the non-coding RNA upperhand controls Hand2 expression and heart development. Nature 539, 433–436 (2016).
Luo, S. et al. Divergent lncRNAs regulate gene expression and lineage differentiation in pluripotent cells. Cell Stem Cell 18, 637–652 (2016).
Liu, B. et al. Long noncoding RNA lncKdm2b is required for ILC3 maintenance by initiation of Zfp292 expression. Nat. Immunol. 18, 499–508 (2017).
Whissell, G. et al. The transcription factor GATA6 enables self-renewal of colon adenoma stem cells by repressing BMP gene expression. Nat. Cell Biol. 16, 695–707 (2014).
Kaneko, S. et al. Interactions between JARID2 and noncoding RNAs regulate PRC2 recruitment to chromatin. Mol. Cell 53, 290–300 (2014).
Klattenhoff, C. A. et al. Braveheart, a long noncoding RNA required for cardiovascular lineage commitment. Cell 152, 570–583 (2013).
Xia, P. Y. et al. WASH is required for the differentiation commitment of hematopoietic stem cells in a c-Myc-dependent manner. J. Exp. Med. 211, 2119–2134 (2014).
Zhu, P. P. et al. ZIC2-dependent OCT4 activation drives self-renewal of human liver cancer stem cells. J. Clin. Invest. 125, 3795–3808 (2015).
Schepers, A. G. et al. Lineage tracing reveals Lgr5+ stem cell activity in mouse intestinal adenomas. Science 337, 730–735 (2012).
Junttila, M. R. et al. Targeting LGR5+ cells with an antibody–drug conjugate for the treatment of colon cancer. Sci. Transl. Med. 7, 314ra186 (2015).
Hao, H. X. et al. ZNRF3 promotes Wnt receptor turnover in an R-spondin-sensitive manner. Nature 485, 195–200 (2012).
Storm, E. E. et al. Targeting PTPRK-RSPO3 colon tumours promotes differentiation and loss of stem-cell function. Nature 529, 97–100 (2016).
Zhu, X. X. et al. An efficient genotyping method for genome-modified animals and human cells generated with CRISPR/Cas9 system. Sci. Rep. 4, 6420 (2014).
Yoshimi, K. et al. ssODN-mediated knock-in with CRISPR-Cas for large genomic regions in zygotes. Nat. Commun. 7, 10431 (2016).
Miyoshi, H. & Stappenbeck, T. S. In vitro expansion and genetic modification of gastrointestinal stem cells in spheroid culture. Nat. Protoc. 8, 2471–2482 (2013).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Fujii, M., Matano, M., Nanki, K. & Sato, T. Efficient genetic engineering of human intestinal organoids using electroporation. Nat. Protoc. 10, 1474–1485 (2015).
Zhu, P. P. et al. C8orf4 negatively regulates self-renewal of liver cancer stem cells via suppression of NOTCH2 signalling. Nat. Commun. 6, 7122 (2015).
Acknowledgements
We thank X. Shi, Y. Xu and Y. Teng for technical support. We thank J. Li (Cnkingbio Company Ltd, Beijing, China) for technical support. This work was supported by the National Natural Science Foundation of China (91640203, 31530093, 81330047, 31771638, 81672897, 81672956, 81472413, 81572433, 81772646 and 31601189), Strategic Priority Research Programs of the Chinese Academy of Sciences (XDA12020219 and XDB19030203), Beijing Natural Science Foundation (7181006, 7162125) and the Postdoctoral Innovative Talent Support Program to P.Z.
Author information
Authors and Affiliations
Contributions
P.Z. designed and performed experiments, analysed data and wrote the paper; J.W. and Y.W. performed experiments and analysed data; X.Z., S.M. and D.F. generated genome-modified mice; T.L., Luyun He, L.Y., X.Q., Y.D., C.L. and Lei He performed some experiments; B.L., B.Y., S.W. and J.W. analysed data; X.W., Lei He and W.R. provided CRC samples; Y.T. designed the study and analysed data; Z.F. initiated the study, organized, designed and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Biological characteristics of lncGata6.
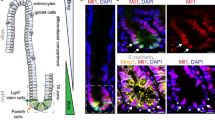
a, LncRNAs were depleted through CRISPRi strategy and examined by Realtime PCR (upper panel) and Northern blot (lower panel). n = 3 biologically independent samples. b, LncGata6 expression levels were detected by realtime PCR. n = 8 mice were examined. c, d, LncGata6 overexpressing ISCs cells were generated via lentivirus carrying lncGata6 infection (c), followed by organoid formation assay (d). n = 6 mice for each group. e, f, LncGata6 silenced ISCs were generated via lentivirus (e), followed by organoid formation assay (f). n = 6 mice for each group. g, Schematic annotation of lncGata6 genomic locus on mouse chromosome 18. LncGata6 orthologs were analyzed in different species. Three isoforms were annotated (a, b, c). h, Expression levels of isoforms a, b and c in ISCs were examined through realtime PCR and isoform a (lncGata6) was identified as the major transcript. n = 4 independent experiments per group. i, qRT-PCR analysis of lncGata6 expression levels in adult mouse tissues (n = 4). j, Full length lncGata6 of mouse intestines was cloned by 3’RACE and 5’RACE. k, LncGata6 was translated by in vitro translation analysis. l, Coding capacity of lncGata6 was predicted by CPC and CPAT. m, Fractionation of mouse organoid lysates followed by qRT-PCR (left panel). Fractionation controls were analyzed by immunoblotting (right panel). EEA1, endosome antigen 1; H3, histone H3; N: Nuclear fraction; C: Cytoplasmic fraction. n = 4 independent experiments. n, LncGata6RFP mice were generated by CRISPR/Cas9 approach through inserting IRES-RFP cassette into the end of lncGata6 allele immediately after the exon4. o, Intestinal crypts isolated from Lgr5GFP;lncGatat6RFP mice were used for GFP/RFP detection by FACS. p, Indicated intestinal sections from Lgr5GFP-CreERT2 mice were stained with digoxin-labeled lncGata6 probes, followed by digoxin antibody and Alexa-594 conjugated secondary antibody. Scale bar, 50 μm. q, Lgr5GFP-CreERT2 mice were crossed with lncGata6RFP mice, and RFP+ cells were enriched using FACS for organoid formation. Scale bar, 300 μm. All data are representative of at least three independent experiments. In all panels, data are shown as mean ± s.d. ** P < 0.01, *** P < 0.001, by unpaired Student’s t-test. Raw data and P values are shown in Supplementary Table 4.
Supplementary Figure 2 Generation of lncGata6 KO, lncGata6Stop and lncGata6flox mice.
a, b LncGata6 KO mice were generated by CRISPR/Cas9 technology (a) and tested by agarose gels (b). c, LncGata6 deletion was detected by qRT-PCR analysis. n = 6 mice, 3 pairs of primers were used. d, Gata6 expression levels were examined by Western blot. e, Gata6 expression in lncGata6-/- ISCs and non-ISCs was examined by realtime PCR. n = 3 independent experiments per group. f, FACS analysis of ISCs in lncGata6-/- mice. n = 6 for each group. g, mRNA expression levels of the indicated ISC markers were examined by realtime PCR. n = 5 for each group. h, i, Ascl2 (h) and Lysozyme (i) were detected in lncGata6+/+ and lncGata6-/- intestines. Paneth cells and CBC cells were denoted by brown and black arrowheads, respectively. n = 20 fields were observed. Scale bars, 50 μm. j, Schematic representation of generation of lncGata6flox mice. Two loxPs were inserted into lncGata6 allele flanking at the exon 2-4 of lncGata6 gene. k, lncGata6flox/flox mice were identified by agarose gels. l, Lgr5GFP-CreERT2;lncGata6flox/flox mice were treated with tamoxifen (TAM) for lncGata6 knockout. m, Intestinal whole-mount staining of β-gal for LRlacZ mice. Typical jejunum sections of P65 were shown. n = 7 mice were sacrificed at indicated time points. For each mouse, crypt-villus unit (blue plot) numbers were counted within 1 cm small intestine. Scale bars, 300 mm. n, Cell cycle and quiescence of lncGata6-/- ISCs from LRlacZ;lncGata6-/- mice were examined by Hoechst/Pyronin Y FACS. n = 6 mice per group. o, p, Proliferation (o) and apoptosis (p) of LRlacZ;lncGata6-/- ISCs were examined by Ki67 and TUNEL staining, respectively. For o, n = 6 mice per group. For p, n = 5 mice per group. q, SV40 poly(A) (STOP) module was inserted into the promoter of lncGata6 through CRISPR/Cas9 approach. r, Gata6 expression in LncGata6Stop mice was examined by Western blot (lower panel). t, Gata6 expression levels in LncGata6Stop ISCs and non-ISCs were examined by realtime PCR. n = 3 independent experiments per group. u, Representative H&E staining images of intestines from two-month-old WT and lncGata6Stop mice (n = 6 per group). Scale bar, 50 μm. v, ISCs from lncGata6WT and lncGata6 Stop mice were visualized (left panel) and calculated (right panel). n = 200 fields for ISC calculation. Scale bar, 50 μm. w. Intestinal organoid formation assay of lncGATA6 depleted human intestinal crypt cells. Scale bar, 300 μm. n = 3 independent experiments. In all panels, data are shown as mean ± s.d. * P < 0.05, ** P < 0.01, *** P < 0.001, ns, not significant, by unpaired Student’s t-test. Raw data and P values are shown in Supplementary Table 4. All data are representative of at least three independent experiments.
Supplementary Figure 3 Identification of interacted proteins of lncGata6 in intestinal crypts.
a, The indicated genes located near lncGata6 locus ( < 1 megabase) were examined in lncGata6-/- and lncGata6Stop mice. n = 3 independent experiments per group. Primers were listed in Supplementary Table 3. b, For probe screening, imbricate antisense DNA probes against lncGata6 transcript were designed and incubated with mouse crypt cell lysates, followed by RNase H treatment, and then detected for lncGata6 integrity by Northern blot. c, d, Captured peptide sequences of MS-MS profiles for Bptf (c) and Snf2l (d), corresponding peptide sequences are listed on the top of corresponding graphs. e, CRISPRi strategy was used to delete Bptf, Snf2l or Rbbp4 in crypt cells for RNA immunoprecipitation (RIP). LncGata6 enrichment was examined by realtime PCR. n = 3 independent experiments. f, Immunofluorescence staining of intestinal organoids. Intestinal organoids were collected for immunofluorescence with indicated antibodies and lncGata6 probes. Scale bar, 100 μm. g, Prediction structure of the exon 4 of lncGata6. Prediction of the exon 4 region of lncGata6 structure was based on minimum free energy (MFE). WT and mutant lncGata6 were shown in upper and lower panels, respectively. HR: hairpin loop region. h, Schematic representation of generation of lncGata6Mut mice. Mutation of lncGata6 at HR3 region in LncGata6Mut mice was confirmed by DNA sequencing. i, PCR products from LncGata6Mut and LncGata6WT intestinal tissues were digested by BamHI, confirming the mutation of lncGata6 (CCGTCC- > TGGATCC). j, ISCs were isolated from Lgr5GFP-CreERT2;lncGata6Mut and Lgr5GFP-CreERT2;lncGata6WT mice, followed by organoid formation assay. Typical images were shown in left panel and organoid formation ratios were shown in right panel. n = 3 independent experiments per group. k, LncGata6Mut ISCs were detected by FACS (left panel), and GFP+ cells were visualized by microscope (right panel). Scale bars, 10 μm. Four lncGata6Mut mice were used for ISC enrichment, and ISCs were pooled together for transcriptome analysis. In all panels, data are shown as mean ± s.d. * P < 0.05, ** P < 0.01, *** P < 0.001, ns, not significant, by unpaired Student’s t-test. Raw data and P values are shown in Supplementary Table 4. Data represent at least three independent experiments.
Supplementary Figure 4 LncGata6 and the NURF complex promote Ehf expression leading to Lgr4/Lgr5 promoter activation.
a, Gene ontology analysis of downregulated genes in lncGata6Mut ISCs versus WT ISCs. b, LRCas9 mice were infected with AAV sgRNA, and GFP+ cells were isolated five days later for Western blots with indicated antibodies. c, Indicated genes were silenced in ISCs through shRNA, followed by organoid formation. d, Ehf was rescued in Ehf silenced ISCs for organoid formation detection. ShRNA sequences were listed in supplementary Table 2. e, Ehf was rescued in lncGata6 knockout ISCs, and self-renewal of ISCs was examined by organoid formation. Scale bar, 300 μm. f, LncGata6 Chromatin Isolation by RNA purification (ChIRP) was performed and indicated eluates were examined for enrichment of Ehf promoter. g, ChIP assays against Bptf, Snf2l and Rbbp4 were performed, followed by realtime PCR for enrichment of the indicated regions of Ehf promoter. h, Indicated regions of Ehf promoter were cloned into pGL3 luciferase reporter plasmid for promoter activation detection. oelncG6, overexpressing lncGata6. i, Schematic representation of generation of Bptfflox mice. The second exon (E2) was deleted with CRISPR/Cas9 technology. j, Bptfflox/flox mice were identified by agarose electrophoresis. k, Bptf was analyzed in Bptf knockout ISCs by Western blot. l, Schematic representation of Ehf knockout mice. DNA sequencing results were shown in lower panel. TSS, transcription start site. m, Ehf knockout efficiency in intestines were detected by Western blot. n, GFP+ ISCs (gated by green) and GFP- non-ISCs (gated by black) were isolated from Ehf-/- mice through FACS (left panel), and GFP signals were visualized by confocal microscopy (right panel). Scale bars, 10 μm. o, p, Ehf ChIP assay was performed with Ehf−/− and WT intestinal cells, and enrichment of indicated regions on Lgr4 (o) and Lgr5 (p) promoters was examined by realtime PCR. q, Indicated organoids were established and expression levels of Lgr4 and Lgr5 were examined by Western blot. For f, n = 4 independent experiments per group. For c-e, g, h, o, p, n = 3 independent experiments per group, data are shown as mean ± s.d. * P < 0.05, ** P < 0.01, *** P < 0.001, by unpaired Student’s t-test. Raw data and P values are listed in Supplementary Table 4.
Supplementary Figure 5 Ehf knockout abrogates the stemness of ISCs.
a, Ehf expression levels in the indicated murine tissues were examined by Western blot. Gapdh served as a loading control. b, Ehf in situ hybridization was performed in intestine tissues. AP-red was used for detection. c, Typical images of intestines from 2-month-old Ehf+/+ and Ehf–/– mice. For each group, n = 200 fields from six mice were examined (mean ± s.d.). d, Ehf+/+ and Ehf–/– intestines were stained with Olfm4. Representative images were shown in left panels and average cell numbers were shown in right panel (mean ± s.d.). n = 20 fields were observed. e, Ascl2 in situ hybridization was performed in Ehf+/+ and Ehf–/– intestines. f, Lysozyme was stained in Ehf+/+ and Ehf–/– intestines for observation of Paneth cells and CBC cells. n = 20 fields were observed and data were shown as mean ± s.d. g, h, Multiple color staining was performed using HRP/AP detection kits (GBI labs). Ehf+/+ and Ehf–/– intestines and the indicated antibodies were used for multiple color staining. AP-red, AP-blue and DAB were used for observation (g). The indicated cell numbers (n = 19 fields) in Ehf+/+ and Ehf–/– intestines were shown as mean ± s.d. (h). VCU, villus-crypt unit. i, Indicated KO LRlacZ ISCs (1×104) and WT LRYFP ISCs (1×104) were mixed for passages of organoid formation, and established organoids were analyzed using FACS. Scale bars in all panels, 50 μm. YFP+ cells and β-gal+ cells are daughter cells of LRYFP ISCs and indicated knockout ISCs, respectively. *** P < 0.001, by unpaired Student’s t-test. Raw data and P values are listed in Supplementary Table 4. Data represent four independent experiments.
Supplementary Figure 6 LncGata6 is implicated in CRC oncogenesis.
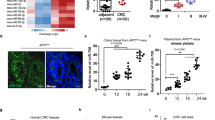
a, A colon cancer tissue array was used for lncGata6 in situ staining. Typical images were shown in left panel. LncGata6high cell ratios and lncGata6 photon intensity were shown in right panel. Scale bars, 50 μm. Sample numbers (n) were shown in right panel. b, Colon cancer samples were divided into two groups according to lncGATA6 expression levels, and Kaplan–Meier survival analysis was performed. c, BPTF was detected in colon tumors by immunohistochemistry. Representative results are shown in left panels and photon intensity is shown in right panel. Scale bars, 50 μm. Sample numbers (n) are shown. d, e, Colon cancer samples were divided into two groups according to BPTF expression levels (d) or BPTF/lncGATA6 together (e), followed by Kaplan–Meier survival analysis. f, Colorectal cancer primary cells were infected with lncGATA6 silenced or control lentivirus (pSiCoR-GFP), and proliferation was examined by Ki67 staining. n = 10 fields were observed for each treated cells. Scale bars, 50 μm. g, LncGATA6 silenced cells were generated with lentivirus (pSiCoR-puro), and cell proliferation was detected with CFSE staining and FACS. h, LncGATA6 silenced and control cells were used for oncosphere formation, and impaired sphere formation was found upon lncGATA6 depletion. Scale bars, 500 μm. i, 1 × 106 lncGATA6 silenced or control cells were injected into B-NSG mice for tumor growth. One month later, tumors were obtained and tumor weight was detected. n = 5 mice per group. j, 1 × 106 lncGATA6 silenced or control cells were injected into B-NSG mice, and the indicated tissues were obtained three months later. Human α-satellite DNA and pSiCoR DNA were detected by realtime PCR. n = 3 independent experiments per group. k, 1 × 106 lncGATA6 depleted cells were used for tumor formation, and CD31 was detected by FACS. In all panels, data are shown as mean ± s.d. * P < 0.05, ** P < 0.01, *** P < 0.001, by unpaired Student’s t-test. Raw data and P values are listed in Supplementary Table 4. All functional experiments were repeated at least three times.
Supplementary Figure 7 LncGATA6 is highly expressed in colorectal cancer stem cells.
a, b, Decreased ISCs in lncGata6 knockout colons were detected by microscopy (a) and FACS analysis (b). Scale bars, 50 μm. n = 6 mice per group and data were shown as mean ± s.d. c, Schematic representation for generation of LGR5P2A-GFP or lncGATA6IRES-GFP CRC cells by inserting GFP into the LGR5 or lncGATA6 alleles with CRISPR/Cas9 technology. d, LGR5P2A-GFP (left) and lncGATA6IRES-GFP (right) clones were examined by DNA electrophoresis. Two PCR products were observed because of heterozygous knock-in. e, f, GFP signals for LGR5 (e) or lncGATA6 (f) in CCOs were assayed by FACS. LGR5P2A-GFP and lncGATA6IRES-GFP correctly targeted CCOs were used for FACS analysis and the ratios of LGR5+ (e) or lncGATA6+ (f) ISCs in CCOs were shown. Five CRC samples were used for generation of LGR5GFP and lncGATA6GFP cells and results of 3 CRC samples were shown. g, GFP+ stem cells (S) and GFP- non-stem cells (N) were sorted by FACS, and total RNAs were extracted for lncGATA6 detection by Northern blot. h, Wnt/β-catenin activation in the indicated cells was analyzed with TOPFLash luciferase assay. n = 7 mice per group and data were shown as mean ± s.d. i, Schematic representation for generation of LGR5iCR or lncGATA6iCR CRC cells by inserting iCaspase9-P2A-RFP into LGR5 or lncGATA6 alleles via CRISPR/Cas9 technology. j, LGR5iCR (left) and lncGATA6CR (right) clones were generated and examined by DNA electrophoresis. k, RFP+ LGR5iCR or lncGATA6iCR CRC cells were enriched by FACS. LGR5iCR and lncGATA6iCR correctly targeted CCOs were used for FACS analysis and the ratios of LGR5+ and lncGATA6+ CRCs in CCOs were shown. l, LGR5iCR or lncGATA6CR tumors were established, and dimerizer was administrated when tumors reached to 400 mm3. At the indicated time points, dimerizer treated and non-treated tumors were obtained and examined for RFP expression by FACS. m, CRC primary cells were infected with Caspase9-GFP lentivirus, GFP+ cells were analyzed by FACS. In all panels, data are shown as mean ± s.d. ** P < 0.01, *** P < 0.001, by unpaired Student’s t-test. Raw data and P values are listed in Supplementary Table 4. Data represent three independent experiments.
Supplementary Figure 8 Full blots of figures.
The red sections indicate blot results shown in the indicated figures.
Supplementary information
Supplementary Information
Supplementary Figures 1–8 and Supplementary Table legends.
Supplementary Table 1
Sequences for sgRNAs used in this work.
Supplementary Table 2
Sequences for shRNAs used in this work.
Supplementary Table 3
Sequences for primers used in this work.
Supplementary Table 4
Statistics source data.
Rights and permissions
About this article
Cite this article
Zhu, P., Wu, J., Wang, Y. et al. LncGata6 maintains stemness of intestinal stem cells and promotes intestinal tumorigenesis. Nat Cell Biol 20, 1134–1144 (2018). https://doi.org/10.1038/s41556-018-0194-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0194-0