Abstract
Solid malignancies have been speculated to depend on cancer stem cells (CSCs) for expansion and relapse after therapy. Here we report on quantitative analyses of lineage tracing data from primary colon cancer xenograft tissue to assess CSC functionality in a human solid malignancy. The temporally obtained clone size distribution data support a model in which stem cell function in established cancers is not intrinsically, but is entirely spatiotemporally orchestrated. Functional stem cells that drive tumour expansion predominantly reside at the tumour edge, close to cancer-associated fibroblasts. Hence, stem cell properties change in time depending on the cell location. Furthermore, although chemotherapy enriches for cells with a CSC phenotype, in this context functional stem cell properties are also fully defined by the microenvironment. To conclude, we identified osteopontin as a key cancer-associated fibroblast-produced factor that drives in situ clonogenicity in colon cancer.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Vermeulen, L. et al. Defining stem cell dynamics in models of intestinal tumor initiation. Science 342, 995–998 (2013).
Kozar, S. et al. Continuous clonal labeling reveals small numbers of functional stem cells in intestinal crypts and adenomas. Cell Stem Cell 13, 626–633 (2013).
Ritsma, L. et al. Intestinal crypt homeostasis revealed at single-stem-cell level by in vivo live imaging. Nature 507, 362–365 (2014).
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Buczacki, S. J. et al. Intestinal label-retaining cells are secretory precursors expressing Lgr5. Nature 495, 65–69 (2013).
Vermeulen, L. & Snippert, H. J. Stem cell dynamics in homeostasis and cancer of the intestine. Nat. Rev. Cancer 14, 468–480 (2014).
Chen, J. et al. A restricted cell population propagates glioblastoma growth after chemotherapy. Nature 488, 522–526 (2012).
Schepers, A. G. et al. Lineage tracing reveals Lgr5+ stem cell activity in mouse intestinal adenomas. Science 337, 730–735 (2012).
Driessens, G., Beck, B., Caauwe, A., Simons, B. D. & Blanpain, C. Defining the mode of tumour growth by clonal analysis. Nature 488, 527–530 (2012).
Nakanishi, Y. et al. Dclk1 distinguishes between tumor and normal stem cells in the intestine. Nat. Genet. 45, 98–103 (2013).
Vermeulen, L. et al. Single-cell cloning of colon cancer stem cells reveals a multi-lineage differentiation capacity. Proc. Natl Acad. Sci. USA 105, 13427–13432 (2008).
Todaro, M. et al. Colon cancer stem cells dictate tumor growth and resist cell death by production of interleukin-4. Cell Stem Cell 1, 389–402 (2007).
O’Brien, C. A., Pollett, A., Gallinger, S. & Dick, J. E. A human colon cancer cell capable of initiating tumour growth in immunodeficient mice. Nature 445, 106–110 (2007).
Ricci-Vitiani, L. et al. Identification and expansion of human colon-cancer-initiating cells. Nature 445, 111–115 (2007).
Shimokawa, M. et al. Visualization and targeting of LGR5+ human colon cancer stem cells. Nature 545, 187–192 (2017).
Li, H. et al. Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors. Nat. Genet. 49, 708–718 (2017).
Lamprecht, S. et al. Multicolor lineage tracing reveals clonal architecture and dynamics in colon cancer. Nat. Commun. 8, 1406 (2017).
Vermeulen, L. et al. Wnt activity defines colon cancer stem cells and is regulated by the microenvironment. Nat. Cell Biol. 12, 468–476 (2010).
Lotti, F. et al. Chemotherapy activates cancer-associated fibroblasts to maintain colorectal cancer-initiating cells by IL-17A. J Exp. Med. 210, 2851–2872 (2013).
Todaro, M. et al. CD44v6 is a marker of constitutive and reprogrammed cancer stem cells driving colon cancer metastasis. Cell Stem Cell 14, 342–356 (2014).
De Sousa e Melo, F. et al. A distinct role for Lgr5+ stem cells in primary and metastatic colon cancer. Nature 543, 676–680 (2017).
Rodriguez, E. et al. Versatile and enhanced tumour modelling in mice via somatic cell transduction. J. Pathol. 232, 449–457 (2014).
Rulands, S. et al. Universality of clone dynamics during tissue development. Nat. Phys. 14, 469–474 (2018).
Klein, A. M., Nakagawa, T., Ichikawa, R., Yoshida, S. & Simons, B. D. Mouse germ line stem cells undergo rapid and stochastic turnover. Cell Stem Cell 7, 214–224 (2010).
Waclaw, B. et al. A spatial model predicts that dispersal and cell turnover limit intratumour heterogeneity. Nature 525, 261–264 (2015).
Gong, P., Wang, Y., Liu, G., Zhang, J. & Wang, Z. New insight into Ki67 expression at the invasive front in breast cancer. PLoS ONE 8, e54912 (2013).
Horst, D. et al. Differential WNT activity in colorectal cancer confers limited tumorigenic potential and is regulated by MAPK signaling. Cancer Res. 72, 1547–1556 (2012).
De Sousa E Melo, F. et al. Poor-prognosis colon cancer is defined by a molecularly distinct subtype and develops from serrated precursor lesions. Nat. Med. 19, 614–618 (2013).
Merlos-Suárez, A. et al. The intestinal stem cell signature identifies colorectal cancer stem cells and predicts disease relapse. Cell Stem Cell 8, 511–524 (2011).
Bao, S. et al. Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature 444, 756–760 (2006).
Humphries, A. et al. Lineage tracing reveals multipotent stem cells maintain human adenomas and the pattern of clonal expansion in tumor evolution. Proc. Natl Acad. Sci. USA 110, E2490–2499 (2013).
Marusyk, A. et al. Spatial proximity to fibroblasts impacts molecular features and therapeutic sensitivity of breast cancer cells influencing clinical outcomes. Cancer Res. 76, 6495–6506 (2016).
Guinney, J. et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 21, 1350–1356 (2015).
Hu, Y. & Smyth, G. K. ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009).
Susaki, E. A. et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell 157, 726–739 (2014).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA–seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Harris, A. L. Hypoxia—a key regulatory factor in tumour growth. Nat. Rev. Cancer 2, 38–47 (2002).
Kemper, K. et al. BRAF(V600E) kinase domain duplication identified in therapy-refractory melanoma patient-derived xenografts. Cell Rep. 16, 263–277 (2016).
Dobin, A. et al. STAR: ultrafast universal RNA–seq aligner. Bioinformatics 29, 15–21 (2013).
Acknowledgements
This work is supported by the Academic Medical Center (Amsterdam), The New York Stem Cell Foundation, Cancer Research UK, and grants from KWF (UVA2011-4969, UVA2014-7245 and 10529), the Maurits en Anna de Kock Stichting (2015-2), Worldwide Cancer Research (14-1164), the Maag Lever Darm Stichting (MLDS-CDG 14-03), the European Research Council (ERG-StG 638193) and ZonMw (Vidi 016.156.308) to L.V. L.V. is a New York Stem Cell Foundation–Robertson Investigator.
Author information
Authors and Affiliations
Contributions
K.J.L., S.C.L., X.R.R., L.E.N., F.C.L., M.C.L., M.v.d.H., S.M.v.N., A.v.O., N.L., F.d.S.E.M., J.O., P.V., G.H. and L.K. performed experiments. D.M.M., E.M. and L.V. developed the quantitative models. K.J.L., D.M.M., S.C.L., R.S.A., T.v.d.B., E.M., M.F.B. and L.V. analysed data. S.K.L. and G.S. contributed reagents. K.J.L., D.M.M., M.F.B. and L.V. conceived and designed research. D.J.W. and J.P.M. advised on the work and commented critically on the manuscript. K.J.L., D.M.M., M.F.B. and L.V. wrote the manuscript. L.V. directed the research. All authors approved the content of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Subcutaneous and orthotopically grafted xenograft models are well differentiated.
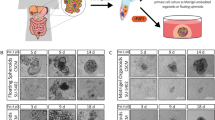
(a) Hematoxylin and Eosin (H&E) staining of Co100 subcutaneous (s.c) xenograft. Scale bar, 500 μm. Representative of 5 independent tumours. (b) Immunofluorescent stainings of s.c. Co100 xenografts for mucin-2 (MUC2), cytokeratin 20 (CK20), intestine alkaline phosphatase (IAP), alpha smooth muscle actin (aSMA), lysozyme EC, alpha defensin-5 (aDef-5), all stainings (yellow), nuclear stain, Hoechst (blue). Scale bars, 50 μm (all panels). Representative of 5 independent tumours. (c and d) H&E, Alcian blue (c) and immunofluorescent F-actin (green) (d) staining for s.c. tumour models (Co100, CC09 and HCT-15), nuclear stain, Hoechst (blue). Scale bars, 50 μm (c) or 100 μm (d) Representative of 5 independent tumours. (e) H&E staining of Co100 and HCT-15 orthotopically grafted tumours (cecal wall). Scale bars, 500 μm. Representative of 3 independent tumours. (f) Orthotopically growing Co100 tumours do express differentiation markers as indicated (yellow). Nuclear stain, Hoechst (blue). Scale bars, 100 μm. Representative of 3 independent tumours
Supplementary Figure 2 In vitro and in vivo validation of LV-indLS2 system.
(a) Single cell cloned cultures of LV-indLS2 transduced cell lines were treated with increasing 4-OH-Tamoxifen (TAM) concentrations, mStrawberry expression was measured 7 days post induction by flow cytometry. (b) mStrawberry expression is randomly distributed amongst stem-like (TOP-GFPhigh) or more differentiated (TOP-GFPlow) Co100.G7 cells directly after induction. (c) mStrawberry labeling is stable in time and does not influence clonal fitness. Single cell cloned LV-indLS2 transduced cultures of respectively Co100, HCT-15 and CC09 cells were treated with 4-OH-Tamoxifen or were cultured untreated as a control, mStrawberry expression was measured at day 7 and day 42 post-treatment by flow cytometry. (a-c) Data is represented as mean ± S.D., n = 3 independent experiments. n.s., not significant, two-sided Student’s t-test. (d) Confocal images of subcutaneous Co100 tumour xenografts, 7 days post induction with the indicated 4-OH-Tamoxifen concentrations. mStrawberry (red), nuclear staining, Hoechst (blue). Scale bars, 100 μm. Representatives of 3 tumours per group. (e) Percentage of mStrawberry+ cells is dose-dependent, as determined by flow cytometry (n = 4 tumours per condition). (f, g) Confocal images of tumour xenograft sections were used for automated fluorescence image (FI) detection of mStrawberry+ cells (f) and the percentage of single cell clones 7 days after induction was determined (g) (n = 10 tumour sections per condition). A 4-OH-Tamoxifen dose of 0.05 mg/mouse resulted in the induction of the optimal number of clones without evidence for clone collision; the induction of too many clones results in overestimation of clone sizes. (e-g) Data is represented as mean ± S.D. (h) Quantification of the distances between cells from different clones 1 week after clonal induction with the indicated 4-OH-Tamoxifen concentrations. The red dashed lines indicate the maximum distance between cells within a clone
Supplementary Figure 3 Marker free lineage tracing and model inference for additional xenograft models.
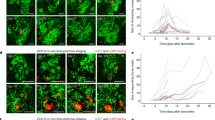
(a, b) Images of labelled clones in Co100 tumours either orthotopically (a) or subcutaneously (b) grafted in NSG mice. Scale bars, 250 μm. (c, d) Representative images of labelled clones in HCT-15 (c) and CC09 (d) xenografts at indicated time points. mStrawberry (red), F-Actin (green), nuclear stain, Hoechst (blue). Scale bars, 250 μm. (e-h) Sectional clone size distribution over time in orthotopic Co100 (e), subcutaneous Co100 (NSG mice) (f), HCT-15 (g) and CC09 (h) xenografts. Depicted are the experimentally measured fractions of growing clones within the indicated bins (black dots) and the model predicted clone size distributions using the optimal fit parameters (dashed line). Data is represented as mean ± S.E.M., grey shade represents 95% confidence interval of prediction. Source data for Supplementary Figure 3 can be found in Supplementary Table 1
Supplementary Figure 4 Proliferation is associated with stromal cells but not with vasculature or hypoxia.
(a) Images of human primary colorectal cancer tissue. Ki67+ cells, (yellow), myofibroblasts (aSMA, green). Nuclear stain, Hoechst (blue). Scale bars, 0.5 mm (left) and 100 μm (right), representative of 10 independent tumours. (b) Percentage of Ki67+ cells is significantly higher in the edge region (E) of human primary colon cancer tissue compared to centre (C). (c) Average distance of either Ki67 negative cells or proliferating (Ki67+) tumour cells to the nearest aSMA+ fibroblast in human primary colon cancer tissue. (b, c) Data is represented as mean ± S.D., n = 10 tumours. (Two-tailed Wilcoxon matched pairs signed rank test). (d-e) Cleaved Caspase-3 (green) (d) or CD31 (yellow) (e) expression in edge (upper) and centre areas (lower) in a Co100 tumour section. Nuclear staining, Hoechst (blue). Scale bars, 100 μm (d) or 200 μm (e), representative of 5 tumours. (f) Immunohistochemical staining for HIF-1a in the edge (upper) and centre (lower) of a Co100 tumour. Scale bars, 100 μm, representative of 3 tumours. (g) Gene set enrichment analysis on RNA expression profiles of cells located at the edge and centre of the xenografts indicates an enrichment for DNA replication and cell cycle gene sets (Molecular Signatures Database entries M1017 and M5468) in the edge region, but not for hypoxia associated genes43, n = 2 tumours per cell line. (h) Immunofluorescence staining for aSMA (green) and Ki67 (yellow), in HCT-15 (upper panel) and CC09 (lower panel) xenografts. Nuclear stain, Hoechst (blue). Scale bars, 100 μm. Right images are magnifications of the box in the left images. (i) Average distance of either all or proliferating (Ki67+) tumour cells to the nearest aSMA+ fibroblast in HCT-15 and CC09 (n = 20.000 cells from 6 tumours per cell line) xenografts. (j) Average distance of either all or apoptotic (CC3+) tumour cells to the nearest aSMA+ fibroblast in Co100 tumours treated with Oxaliplatin-5FU (n = 20.000 cells from 4 tumours). (i, j) Data is represented as mean ± S.E.M., distances were compared using paired two-tailed Student’s t-test
Supplementary Figure 5 Growth of individual tumour xenografts is best fitted using a surface growth model.
(a) Tumour growth of individual subcutaneous Co100, HCT-15 and CC09 tumour xenografts was measured in time (black dots). Solid and dashed lines represent respectively model fits of exponential volume and surface growth. Both models are fitted with the initial volume constrained between 0 and the total volume of injected cells (0.1 mm3). (b) Goodness of fit measured by R2 (R-squared), comparing the surface and exponential volume growth models indicate a better fit of the surface model for almost all tumours. R2 values smaller than zero are rounded up to zero. (c) The Akaike information criterion (AIC) similarly indicated surface growth as the best model to describe the data (the best model has the lowest AIC value). (a-c) N = 29 (Co100), 25 (HCT-15) and 23 (CC09) tumours
Supplementary Figure 6 Osteopontin is a fibroblast secreted factor that enhances clonogenicity.
(a) Co100 cells were adherently seeded as single cells, with or without human or mouse primary intestinal fibroblasts conditioned medium. 3 days after seeding, cells were stained for F-Actin (green) and nuclear stain Hoechst (blue). Scale bars, 100 μm, representative of 10 images per condition. (b) Quantification of clone sizes of data shown in panel (a), n = 10 images per condition. (c) Proliferation of Co100 cells growing in a mixture of reduced growth medium and fibroblast conditioned medium, 3 days after seeding, n = 3 replicates, data is represented as mean ± S.D. (d) Analysis of murine gene expression, specifically encoding secreted proteins, obtained from RNAseq data of 3 different xenograft models, revealed Spp1, encoding osteopontin (OPN) as the most abundantly expressed. (e) In vitro treatment of Co100 cells with recombinant human OPN (500 ng/ml) increased proliferation, n = 3 independent experiments. (f, g) Analysis of OPN expression of Co100.OPN cells on mRNA (n = 1 experiment with 3 technical replicates) (f) and secreted protein levels (n = 2 independent experiments) (g) by respectively qPCR and ELISA. (wt, wildtype Co100, neg, OPN- Co100 population, OPN, OPN+ Co100 population) (e-g) Data is represented as mean ± S.D. (h) Relative clone frequency (colour) per clone size (columns) in time (rows) for Co100.OPN tumours. Number of clones and tumours (between parentheses) are depicted. (i) Goodness of fit (inverse and normalized least-squares distance) as a function of A and h on expanding clones (clone size > 1 cell) in Co100.OPN xenografts. Source data are shown in Supplementary Table 1. (j) Ki67 (yellow) expression in Co100.OPN xenografts, aSMA (green), nuclear stain, Hoechst (blue). Scale bars, 500 μm, representative of 8 tumours per group. Quantification of Ki67 (k) and aSMA (l) expression in Co100 (white bars) (aSMA as shown in Fig. 5h) and Co100.OPN (red bars) xenografts. (n = 8 (Co100) and 12 (Co100.OPN) for Ki67 and n = 10 (Co100) and 12 (Co100.OPN) for aSMA . Two-tailed Student’s t-test, to compare edge to centre a paired two-tailed Student’s t-test was used. Data is represented as mean ± S.D
Supplementary Figure 7 Example figure FACS gating strategy.
Cells were selected in FSC/SSC dot plot to remove debris, single cells were gated using the FSC-H/FSC-W dot plot. GFP+, mStrawberry+ or PE+ cells were gated and compared with a control sample with no detectable fluorochrome expression
Supplementary information
Supplementary Information
Supplementary Figures 1–7, Supplementary Table legends and Supplementary Note 1
Supplementary Table 1
Statistics Source Data. All clone sizes (in cells) in Co100, CC09 and HCT-15 xenografts at the indicated time points. Data is used for Figures 1, 2, 6 and 7 and Supplementary Figures 3 and 6
Supplementary Table 2
Differentially expressed genes between centre and edge region of Co100, CC09 and HCT-15 xenografts. Listed are all genes that are significantly differentially expressed (non-multiple testing corrected p < 0.05) between the centre and edge region of all three xenograft models together. Two replicates were included for each cell line. All RNA sequencing data are deposited in the GEO database under GSE95499
Rights and permissions
About this article
Cite this article
Lenos, K.J., Miedema, D.M., Lodestijn, S.C. et al. Stem cell functionality is microenvironmentally defined during tumour expansion and therapy response in colon cancer. Nat Cell Biol 20, 1193–1202 (2018). https://doi.org/10.1038/s41556-018-0179-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0179-z
This article is cited by
-
Compressive stress triggers fibroblasts spreading over cancer cells to generate carcinoma in situ organization
Communications Biology (2024)
-
Cancer associated fibroblasts serve as an ovarian cancer stem cell niche through noncanonical Wnt5a signaling
npj Precision Oncology (2024)
-
Apoptotic tumor cell-derived microparticles loading Napabucasin inhibit CSCs and synergistic immune therapy
Journal of Nanobiotechnology (2023)
-
Tumor matrix stiffness provides fertile soil for cancer stem cells
Cancer Cell International (2023)
-
Using picoliter droplet deposition to track clonal competition in adherent and organoid cancer cell cultures
Scientific Reports (2023)