Abstract
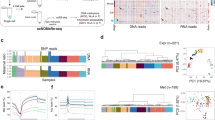
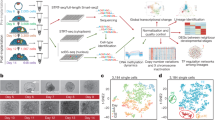
DNA methylation, chromatin states and their interrelationships represent critical epigenetic information, but these are largely unknown in human early embryos. Here, we apply single-cell chromatin overall omic-scale landscape sequencing (scCOOL-seq) to generate a genome-wide map of DNA methylation and chromatin accessibility at single-cell resolution during human preimplantation development. Unlike in mice, the chromatin of the paternal genome is already more open than that of the maternal genome at the mid-zygote stage in humans, and this state is maintained until the 4-cell stage. After fertilization, genes with high variations in DNA methylation, and those with high variations in chromatin accessibility, tend to be two different sets. Furthermore, 1,797 out of 5,155 (35%) widely open chromatin regions in promoters closed when transcription activity was inhibited, indicating a feedback mechanism between transcription and open chromatin maintenance. Our work paves the way for dissecting the complex, yet highly coordinated, epigenetic reprogramming during human preimplantation development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Burton, A. & Torres-Padilla, M. E. Chromatin dynamics in the regulation of cell fate allocation during early embryogenesis. Nat. Rev. Mol. Cell Biol. 15, 723–734 (2014).
Li, E. Chromatin modification and epigenetic reprogramming in mammalian development. Nat. Rev. Genet. 3, 662–673 (2002).
Rossant, J. & Tam, P. P. New insights into early human development: lessons for stem cell derivation and differentiation. Cell Stem Cell 20, 18–28 (2017).
Rugg-Gunn, P. J., Cox, B. J., Ralston, A. & Rossant, J. Distinct histone modifications in stem cell lines and tissue lineages from the early mouse embryo. Proc. Natl Acad. Sci. USA 107, 10783–10790 (2010).
Saitou, M., Kagiwada, S. & Kurimoto, K. Epigenetic reprogramming in mouse pre-implantation development and primordial germ cells. Development 139, 15–31 (2012).
Guo, F. et al. Single-cell multi-omics sequencing of mouse early embryos and embryonic stem cells. Cell Res. 27, 967–988 (2017).
Lu, F. et al. Establishing chromatin regulatory landscape during mouse preimplantation development. Cell 165, 1375–1388 (2016).
Wu, J. et al. The landscape of accessible chromatin in mammalian preimplantation embryos. Nature 534, 652–657 (2016).
Liu, X. et al. Distinct features of H3K4me3 and H3K27me3 chromatin domains in pre-implantation embryos. Nature 537, 558–562 (2016).
Dahl, J. & Jung, I. Broad histone H3K4me3 domains in mouse oocytes modulate maternal-to-zygotic transition. Nature 537, 548–552 (2016).
Zhang, B., Zheng, H., Huang, B. & Li, W. Allelic reprogramming of the histone modification H3K4me3 in early mammalian development. Nature 537, 553–557 (2016).
Guo, H. et al. The DNA methylation landscape of human early embryos. Nature 511, 606–610 (2014).
Smith, Z. D. et al. DNA methylation dynamics of the human preimplantation embryo. Nature 511, 611–615 (2014).
Okae, H. et al. Genome-wide analysis of DNA methylation dynamics during early human development. PLoS Genet. 10, e1004868 (2014).
Fulka, H., Mrazek, M., Tepla, O. & Fulka, J. Jr. DNA methylation pattern in human zygotes and developing embryos. Reproduction 128, 703–708 (2004).
Molaro, A. et al. Sperm methylation profiles reveal features of epigenetic inheritance and evolution in primates. Cell 146, 1029–1041 (2011).
Hatada, I. et al. Genome-wide profiling of promoter methylation in human. Oncogene 25, 3059–3064 (2006).
Fang, F., Hodges, E., Molaro, A. & Dean, M. Genomic landscape of human allele-specific DNA methylation. Proc. Natl Acad. Sci. USA 109, 7332–7337 (2012).
Zhu, P. et al. Single-cell DNA methylome sequencing of human preimplantation embryos. Nat. Genet. 50, 12–19 (2018).
Ambartsumyan, G. & Clark, A. T. Aneuploidy and early human embryo development. Hum. Mol. Genet. 17, R10–R15 (2008).
Vanneste, E. et al. Chromosome instability is common in human cleavage-stage embryos. Nat. Med. 15, 577–583 (2009).
Bolton, H. et al. Mouse model of chromosome mosaicism reveals lineage-specific depletion of aneuploid cells and normal developmental potential. Nat. Commun. 7, 11165 (2016).
Clark, S. J. et al. scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells. Nat. Commun. 9, 781 (2018).
Pott, S. Simultaneous measurement of chromatin accessibility, DNA methylation, and nucleosome phasing in single cells. eLife 6, e23203 (2017).
Taberlay, P. C. et al. Polycomb-repressed genes have permissive enhancers that initiate reprogramming. Cell 147, 1283–1294 (2011).
Kelly, T. K. et al. Genome-wide mapping of nucleosome positioning and DNA methylation within individual DNA molecules. Genome Res. 22, 2497–2506 (2012).
Nabilsi, N. H. et al. Multiplex mapping of chromatin accessibility and DNA methylation within targeted single molecules identifies epigenetic heterogeneity in neural stem cells and glioblastoma. Genome Res. 24, 329–339 (2014).
Taberlay, P. C., Statham, A. L., Kelly, T. K., Clark, S. J. & Jones, P. A. Reconfiguration of nucleosome-depleted regions at distal regulatory elements accompanies DNA methylation of enhancers and insulators in cancer. Genome Res. 24, 1421–1432 (2014).
Lay, F. D. et al. The role of DNA methylation in directing the functional organization of the cancer epigenome. Genome Res. 25, 467–477 (2015).
Miura, F., Enomoto, Y., Dairiki, R. & Ito, T. Amplification-free whole-genome bisulfite sequencing by post-bisulfite adaptor tagging. Nucleic Acids Res. 40, e136 (2012).
Smallwood, S. A. et al. Single-cell genome-wide bisulfite sequencing for assessing epigenetic heterogeneity. Nat. Methods 11, 817–820 (2014).
Guo, H. et al. DNA methylation and chromatin accessibility profiling of mouse and human fetal germ cells. Cell Res. 27, 165–183 (2017).
Tolstorukov, M. Y., Volfovsky, N., Stephens, R. M. & Park, P. J. Impact of chromatin structure on sequence variability in the human genome. Nat. Struct. Mol. Biol. 18, 510–515 (2011).
Fincher, J. A., Tyson, G. S. & Dennis, J. H. DNA-encoded chromatin structural intron boundary signals identify conserved genes with common function. Int. J. Genomics 2015, 167578 (2015).
Schwartz, S., Meshorer, E. & Ast, G. Chromatin organization marks exon–intron structure. Nat. Struct. Mol. Biol. 16, 990–995 (2009).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Yan, L. et al. Single-cell RNA-seq profiling of human preimplantation embryos and embryonic stem cells. Nat. Struct. Mol. Biol. 20, 1131–1139 (2013).
Okamoto, I. et al. Eutherian mammals use diverse strategies to initiate X-chromosome inactivation during development. Nature 472, 370–374 (2011).
Petropoulos, S. et al. Single-cell RNA-seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 165, 1012–1026 (2016).
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: from properties to genome-wide predictions. Nat. Rev. Genet. 15, 272–286 (2014).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Tsankov, A. M. et al. Transcription factor binding dynamics during human ES cell differentiation. Nature 518, 344–349 (2015).
Stadler, M. B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011).
Gao, L. et al. Chromatin accessibility landscape in human early embryos and its association with evolution. Cell 173, 248–259 (2018).
Wu, J. et al. Chromatin analysis in human early development reveals epigenetic transition during ZGA. Nature 557, 256–260 (2018).
Williams, B. R. et al. Aneuploidy affects proliferation and spontaneous immortalization in mammalian cells. Nat. Genet. 322, 703–709 (2008).
Rivera, C. M. & Ren, B. Mapping human epigenomes. Cell 155, 39–55 (2013).
Wu, J. et al. The landscape of accessible chromatin in mammalian preimplantation embryos. Nature 534, 652–657 (2016).
Mulqueen, R. M. et al. Highly scalable generation of DNA methylation profiles in single cells. Nat. Biotechnol. 36, 428–431 (2018).
Olova, N. et al. Comparison of whole-genome bisulfite sequencing library preparation strategies identifies sources of biases affecting DNA methylation data. Genome Biol. 19, 33 (2018).
Li, R., Qiao, J., Wang, L., Zhen, X. & Lu, Y. Serum progesterone concentration on day of HCG administration and IVF outcome. Reprod. Biomed. Online 16, 627–631 (2008).
Niakan, K. K., Han, J., Pedersen, R. A., Simon, C. & Pera, R. A. Human pre-implantation embryo development. Development 139, 829–841 (2012).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for bisulfite-seq applications. Bioinformatics 27, 1571–1572 (2011).
Eichten, S. R., Stuart, T., Srivastava, A., Lister, R. & Borevitz, J. O. DNA methylation profiles of diverse Brachypodium distachyon align with underlying genetic diversity. Genome Res. 26, 1520–1531 (2016).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-seq. Bioinformatics 25, 1105–1111 (2009).
Ha, G. et al. Integrative analysis of genome-wide loss of heterozygosity and monoallelic expression at nucleotide resolution reveals disrupted pathways in triple-negative breast cancer. Genome Res. 22, 1995–2007 (2012).
Statham, A. L., Taberlay, P. C., Kelly, T. K., Jones, P. A. & Clark, S. J. Genome-wide nucleosome occupancy and DNA methylation profiling of four human cell lines. Genom. Data 3, 94–96 (2015).
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013).
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Van der Auwera, G. A. et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinformatics 43, 11.10.1–11.10.33 (2013).
Li, H. Toward better understanding of artifacts in variant calling from high-coverage samples. Bioinformatics 30, 2843–2851 (2014).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007).
Burger, L., Gaidatzis, D., Schubeler, D. & Stadler, M. B. Identification of active regulatory regions from DNA methylation data. Nucleic Acids Res. 41, e155 (2013).
Acknowledgements
This work was supported by grants from the Ministry of Science and Technology of China (2017YFA0102702 and 2017YFA0103402) and the National Natural Science Foundation of China (81521002 and 31625018). F.T. and J.Q. were also supported by a grant from the Beijing Municipal Science and Technology Commission (D151100002415000). This work was also supported by the Beijing Advanced Innovation Center for Genomics at Peking University. Some of the bioinformatics analyses were conducted on the Computing Platform at the Center for Life Science.
Author information
Authors and Affiliations
Contributions
F.T. and J.Q. conceived the project. F.G., Y.G., Y.R., P.Y., L.Y., R.L., Y.L., J.L. and J.G. performed the experiments. L.L. and B.H. conducted the bioinformatics analyses. F.T., L.L. and Y.G. wrote the manuscript with help from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 CNV and quality control of scCOOL-seq samples.
(a) Summary of CNV of each chromosome in single gametes, blastomeres and hESCs. The number of cells, including euploid and aneuploid cells, in each stage are listed on the left. Twenty-three α-Amanitin treated 8-cell blastomeres are not included. (b) Barplot showing the number of single cells with CNV on each chromosome. (c) Comparison analysis of average chromatin accessibility between euploid and aneuploid cells from zygote to TE. A two-tailed Student’s t-test was used to calculate the significance between two groups based on the normalized GCH methylation, that is, the average GCH methylation of each euploid or aneuploid cell is normalized by the mean value of all euploid cells within that stage. The mean value of all euploid cells in each stage was normalized to 1. P = 0.20. ns, not significant. The sample size in this panel corresponds to the euploid and aneuploid sample sizes provided in Fig. 1a (expect for euploid 8-cell blastomeres, n = 31 cells). The center represents mean, whereas error bars represent 1*SEM. (d) Boxplot showing the mapping rate of scCOOL-seq sample across each stage. (e-f) Boxplots showing the number of WCG (ACG/TCG) (e) and GCH (GCA/GCC/GCT) sites (f) covered in a single-cell sample across each stage at 1x depth. Each box in d-f represents median and 25% and 75% quantiles, while whiskers indicate 1.5 times of IQR. (g-h) Chromatin accessibility and DNA methylation coverage of RefSeq gene promoters (g) and CGIs (h) of scCOOL-seq libraries. Only regions covering 5 GCH or 3 WCG sites were calculated. The last row named ‘Mean’ is the mean coverage of all analyzed single cells. The sample size in d-h corresponds to the euploid sample sizes provided in Fig. 1a (expect for 8-cell, n = 31 cells).
Supplementary Figure 2 Chromatin accessibility around transcription start sites.
(a) Chromatin accessibility 2 kb upstream of the TSS, within the gene body, and 2 kb downstream of the TES of pools of samples across each stage. (b) Chromatin accessibility (upper panel) and DNA methylation level (lower panel) of H3K4me3 marked (in red) and unmarked (in blue) promoters. (c) Chromatin accessibility and DNA methylation levels of promoters with different expression levels. Genes were divided into 4 groups, including highly expressed (RPKM > 10), intermediately expressed (1 < RPKM ≤ 10), and lowly expressed genes (0.1 < RPKM ≤ 1) and genes that were not expressed (RPKM ≤ 0.1). The sample size in b-c corresponds to 3 euploid hESCs. (d) GO analysis of 8,385 genes that turned into an open state after fertilization (Related to Fig. 1d). The sample size in a, d corresponds to the euploid sample sizes provided in Fig. 1a (expect for 8-cell, n = 31 cells).
Supplementary Figure 3 Chromatin accessibility and DNA methylation level around the center of NDRs and NORs.
(a) Diagram of the number of TSS NDRs (NDRs containing TSS), proximal NDRs (NDRs within 2 kb upstream and downstream of the TSS) and distal NDRs (NDRs at least 2 kb away from the TSS) called at each stage from pools of samples. The last row named ‘Total’ is the total number of NDRs called from oocytes to hESCs. Overlapping NDRs longer than 10 bp were elongated and combined as one NDR. (b) Boxplot showing the dynamics of the length of TSS NDR distributions during human preimplantation development. The number of TSS NDR in each stage are shown in Supplementary Fig. 3a. Each box represents median and 25% and 75% quantiles, while whiskers indicate 1.5 times of IQR. (c) Chromatin accessibility and DNA methylation levels around the center of proximal NDRs and distal NDRs. (d) Chromatin accessibility and DNA methylation levels around the center of NORs. The sample size in a-d corresponds to the euploid sample sizes provided in Fig. 1a (expect for 8-cell, n = 31 cells).
Supplementary Figure 4 Heterogeneity of TSS NDRs among individual blastomeres from the 8-cell to blastocyst stage.
(a) RNA expression levels of 3 classes of genes during human preimplantation development. A two-tailed Student’s t-test was used to calculate the significance between comparisons. ns, not significant. The numbers of gene promoters in 3 classes defined in each stage were shown on the left of heat map in Fig. 3e and Supplementary Fig. 4c. (b) Coefficient of variation (CV) of expression levels of homogeneously open, divergent and homogeneously closed genes among individual cells during human preimplantation development. A two-tailed Student’s t-test was used to calculate the significance between comparisons. ns, not significant. The numbers of gene promoters in 3 classes defined in each stage were shown on the left of heat map in Fig. 3e and Supplementary Fig. 4c. Each box in a-b represents median and 25% and 75% quantiles, while whiskers indicate 1.5 times of IQR. (c) Heat map showing chromatin accessibility, DNA methylation levels and enriched GO terms of homogeneously open, divergent and homogeneously closed genes from the 8-cell to blastocyst stage. “n” on the left represents the number of promoters in each class. The sample size in a-c corresponds to the euploid sample sizes provided in Fig. 1a (expect for 8-cell, n = 31 cells).
Supplementary Figure 5 Differential DNA methylation and chromatin accessibility between parental genomes within each aneuploid blastomere during human preimplantation development.
(a) The global pattern of differential DNA methylation levels (left panel) and chromatin accessibility (right panel) between parental genomes within individual aneuploid blastomeres. The green bar represents the difference between euploid oocytes and sperm. (b) The difference in DNA methylation levels and chromatin accessibility between parental genomes within individual aneuploid blastomeres in different genomic regions. The green bar represents the difference between euploid oocytes and sperm. The sample size in a-b corresponds to the aneuploid sample sizes provided in Fig. 1a.
Supplementary Figure 6 Chromatin accessibility evolutionary comparison and the linkage between DNA methylation variance and DNA methylation level of gene promoter regions during human preimplantation development.
(a) Chromatin accessibility evolutionary comparison between human and mouse preimplantation embryos was performed using t-SNE (t-Distributed Stochastic Neighbor Embedding) analysis based on 15,057 human-mouse homologous proximal NDRs longer than 100 bp (NDRs within 2 kb upstream and downstream of the TSS) called from oocytes to ESCs. (b) Density plot of the relationship between the DNA methylation variance of the promoter regions (1 kb upstream and 0.5 kb downstream of the TSS) and the DNA methylation level of the corresponding regions among individual cells in each stage. The sample size in a-b corresponds to the euploid sample sizes provided in Fig. 1a (expect for 8-cell, n = 31 cells).
Supplementary Figure 7 The linkage between DNA methylation reprogramming and chromatin remodeling of gene promoter regions during human preimplantation development.
(a) Density plot of the relationship between the DNA methylation variance of the promoter regions (1 kb upstream and 0.5 kb downstream of the TSS) and the chromatin accessibility of the corresponding regions (200 bp upstream and 100 bp downstream of the TSS) among individual cells in each stage. (b) Density plot of the relationship between the chromatin accessibility variance of the promoter regions and the DNA methylation level of the corresponding regions among individual cells in each stage. The sample size in a-b corresponds to the euploid sample sizes provided in Fig. 1a (expect for 8-cell, n = 31 cells).
Supplementary Figure 8 Chromatin accessibility of repeat elements, subfamilies and around putative TF binding sites.
(a) The chromatin accessibility (left panel) and DNA methylation level (right panel) of SVAs. (b) The chromatin accessibility (left panel) and DNA methylation level (right panel) of the subfamilies of LINEs. L1, red line; L2, blue line. (c) The chromatin accessibility (left panel) and DNA methylation level (right panel) of the subfamilies of SINEs. Alu, red line; MIR, blue line. In a-c, the center represents mean, whereas error bars represent 1*SEM. (d) Boxplot showing the chromatin accessibility of repeat elements during human preimplantation development. Each box represents median and 25% and 75% quantiles, while whiskers indicate 1.5 times of IQR. (e) Heat map showing the openness of SOX2 and NANOG putative binding sites identified in hESCs during human preimplantation development (see the legend of Fig. 7d for details.). The sample size in a-e corresponds to the euploid sample sizes provided in Fig. 1a (expect for 8-cell, n = 31 cells). (f) Venn diagram shows the overlap between human-mouse homologous de-novo predicted enhancers based on open chromatin and low-methylated regions (LMRs). De-novo predicted enhancers in human preimplantation development and hESCs are listed in the Supplementary Table 4.
Supplementary information
Supplementary Information
Supplementary Figures 1–8 and Supplementary Tables 1–4 legends
Rights and permissions
About this article
Cite this article
Li, L., Guo, F., Gao, Y. et al. Single-cell multi-omics sequencing of human early embryos. Nat Cell Biol 20, 847–858 (2018). https://doi.org/10.1038/s41556-018-0123-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0123-2
This article is cited by
-
Advances in single-cell omics and multiomics for high-resolution molecular profiling
Experimental & Molecular Medicine (2024)
-
Single-cell multi-omics profiling of human preimplantation embryos identifies cytoskeletal defects during embryonic arrest
Nature Cell Biology (2024)
-
CRISPR/Cas9 technology: applications in oocytes and early embryos
Journal of Translational Medicine (2023)
-
Whole-genome transcriptome and DNA methylation dynamics of pre-implantation embryos reveal progression of embryonic genome activation in buffaloes
Journal of Animal Science and Biotechnology (2023)
-
Methods and applications for single-cell and spatial multi-omics
Nature Reviews Genetics (2023)