Abstract
The mammary gland is composed of basal cells and luminal cells. It is generally believed that the mammary gland arises from embryonic multipotent progenitors, but it remains unclear when lineage restriction occurs and what mechanisms are responsible for the switch from multipotency to unipotency during its morphogenesis. Here, we perform multicolour lineage tracing and assess the fate of single progenitors, and demonstrate the existence of a developmental switch from multipotency to unipotency during embryonic mammary gland development. Molecular profiling and single cell RNA-seq revealed that embryonic multipotent progenitors express a unique hybrid basal and luminal signature and the factors associated with the different lineages. Sustained p63 expression in embryonic multipotent progenitors promotes unipotent basal cell fate and was sufficient to reprogram adult luminal cells into basal cells by promoting an intermediate hybrid multipotent-like state. Altogether, this study identifies the timing and the mechanisms mediating early lineage segregation of multipotent progenitors during mammary gland development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Change history
17 July 2018
In the version of this Article originally published, ref. 52 was incorrectly only attributed to its corresponding author, Fre, S., and an older title was used. The correct citation should have been: Lilja, A. M. et al. Clonal analysis of Notch1-expressing cells reveals the existence of unipotent stem cells that retain long-term plasticity in the embryonic mammary gland. Nat. Cell Biol. https://doi.org/10.1038/s41556-018-0108-1 (2018)’. This has now been amended in all online versions of the Article.
References
Watson, C. J. & Khaled, W. T. Mammary development in the embryo and adult: a journey of morphogenesis and commitment. Development 135, 995–1003 (2008).
de Visser, K. E. et al. Developmental stage-specific contribution of LGR5+ cells to basal and luminal epithelial lineages in the postnatal mammary gland. J. Pathol. 228, 300–309 (2012).
Lafkas, D. et al. Notch3 marks clonogenic mammary luminal progenitor cells in vivo. J. Cell Biol. 203, 47–56 (2013).
Prater, M. D. et al. Mammary stem cells have myoepithelial cell properties. Nat. Cell Biol. 16, 942–950 (2014).
Rodilla, V. et al. Luminal progenitors restrict their lineage potential during mammary gland development. PLoS Biol. 13, e1002069 (2015).
Tao, L., van Bragt, M. P., Laudadio, E. & Li, Z. Lineage tracing of mammary epithelial cells using cell-type-specific cre-expressing adenoviruses. Stem Cell Rep. 2, 770–779 (2014).
van Amerongen, R., Bowman, A. N. & Nusse, R. Developmental stage and time dictate the fate of Wnt/beta-catenin-responsive stem cells in the mammary gland. Cell Stem Cell 11, 387–400 (2012).
Van Keymeulen, A. et al. Distinct stem cells contribute to mammary gland development and maintenance. Nature 479, 189–193 (2011).
Wang, C., Christin, J. R., Oktay, M. H. & Guo, W. Lineage-biased stem cells maintain estrogen-receptor-positive and -negative mouse mammary luminal lineages. Cell Rep. 18, 2825–2835 (2017).
Wuidart, A. et al. Quantitative lineage tracing strategies to resolve multipotency in tissue-specific stem cells. Genes Dev. 30, 1261–1277 (2016).
Scheele, C. L. et al. Identity and dynamics of mammary stem cells during branching morphogenesis. Nature 542, 313–317 (2017).
Davis, F. M. et al. Single-cell lineage tracing in the mammary gland reveals stochastic clonal dispersion of stem/progenitor cell progeny. Nat. Commun. 7, 13053 (2016).
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010).
Lescroart, F. et al. Early lineage restriction in temporally distinct populations of Mesp1 progenitors during mammalian heart development. Nat. Cell Biol. 16, 829–840 (2014).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Trejo, C. L., Luna, G., Dravis, C., Spike, B. T. & Wahl, G. M. Lgr5 is a marker for fetal mammary stem cells, but is not essential for stem cell activity or tumorigenesis. NPJ Breast Cancer 3, 16 (2017).
Van Keymeulen, A. et al. Reactivation of multipotency by oncogenic PIK3CA induces breast tumour heterogeneity. Nature 525, 119–123 (2015).
Wansbury, O. et al. Transcriptome analysis of embryonic mammary cells reveals insights into mammary lineage establishment. Breast Cancer Res. 13, R79 (2011).
Biggs, L. C. & Mikkola, M. L. Early inductive events in ectodermal appendage morphogenesis. Semin. Cell Dev. Biol. 25–26, 11–21 (2014).
Boras-Granic, K., Chang, H., Grosschedl, R. & Hamel, P. A. Lef1 is required for the transition of Wnt signaling from mesenchymal to epithelial cells in the mouse embryonic mammary gland. Dev. Biol. 295, 219–231 (2006).
Chu, E. Y. et al. Canonical WNT signaling promotes mammary placode development and is essential for initiation of mammary gland morphogenesis. Development 131, 4819–4829 (2004).
Hiremath, M. & Wysolmerski, J. Parathyroid hormone-related protein specifies the mammary mesenchyme and regulates embryonic mammary development. J. Mammary Gland Biol. Neoplasia 18, 171–177 (2013).
Howard, B. A. & Lu, P. Stromal regulation of embryonic and postnatal mammary epithelial development and differentiation. Semin. Cell Dev. Biol. 25–26, 43–51 (2014).
Wysolmerski, J. J., McCaughern-Carucci, J. F., Daifotis, A. G., Broadus, A. E. & Philbrick, W. M. Overexpression of parathyroid hormone-related protein or parathyroid hormone in transgenic mice impairs branching morphogenesis during mammary gland development. Development 121, 3539–3547 (1995).
Raafat, A. et al. Expression of Notch receptors, ligands, and target genes during development of the mouse mammary gland. J. Cell Physiol. 226, 1940–1952 (2011).
Sale, S., Lafkas, D. & Artavanis-Tsakonas, S. Notch2 genetic fate mapping reveals two previously unrecognized mammary epithelial lineages. Nat. Cell Biol. 15, 451–460 (2013).
Robinson, G. W. Cooperation of signalling pathways in embryonic mammary gland development. Nat. Rev. Genet 8, 963–972 (2007).
Mills, A. A. et al. p63 is a p53 homologue required for limb and epidermal morphogenesis. Nature 398, 708–713 (1999).
Yang, A. et al. p63 is essential for regenerative proliferation in limb, craniofacial and epithelial development. Nature 398, 714–718 (1999).
Forster, N. Basal cell signaling by p63 controls luminal progenitor function and lactation via NRG1. Dev. Cell 28, 147-160 (2014).
Dravis, C. et al. Sox10 regulates stem/progenitor and mesenchymal cell states in mammary epithelial cells. Cell Rep. 12, 2035–2048 (2015).
Ye, X. et al. Distinct EMT programs control normal mammary stem cells and tumour-initiating cells. Nature 525, 256–260 (2015).
Choi, Y. S., Chakrabarti, R., Escamilla-Hernandez, R. & Sinha, S. Elf5 conditional knockout mice reveal its role as a master regulator in mammary alveolar development: failure of Stat5 activation and functional differentiation in the absence of Elf5. Dev. Biol. 329, 227–241 (2009).
Oakes, S. R., Hilton, H. N. & Ormandy, C. J. The alveolar switch: coordinating the proliferative cues and cell fate decisions that drive the formation of lobuloalveoli from ductal epithelium. Breast Cancer Res 8, 207 (2006).
Bernardo, G. M. et al. FOXA1 is an essential determinant of ERalpha expression and mammary ductal morphogenesis. Development 137, 2045–2054 (2010).
Balko, J. M. et al. The receptor tyrosine kinase ErbB3 maintains the balance between luminal and basal breast epithelium. Proc. Natl Acad. Sci. USA 109, 221–226 (2012).
Kendrick, H. et al. Transcriptome analysis of mammary epithelial subpopulations identifies novel determinants of lineage commitment and cell fate. BMC Genomics 9, 591 (2008).
Spike, B. T. et al. A mammary stem cell population identified and characterized in late embryogenesis reveals similarities to human breast cancer. Cell Stem Cell 10, 183–197 (2012).
Van Keymeulen, A. et al. Lineage-restricted mammary stem cells sustain the development, homeostasis, and regeneration of the estrogen receptor positive lineage. Cell Rep. 20, 1525–1532 (2017).
Kiselev, V. Y. et al. SC3: consensus clustering of single-cell RNA-seq data. Nat. Methods 14, 483–486 (2017).
Ibusuki, M. et al. Midkine in plasma as a novel breast cancer marker. Cancer Sci. 100, 1735–1739 (2009).
Kuang, X. Y. et al. Stathmin and phospho-stathmin protein signature is associated with survival outcomes of breast cancer patients. Oncotarget 6, 22227–22238 (2015).
Prochazkova, I. et al. Targeted proteomics driven verification of biomarker candidates associated with breast cancer aggressiveness. Biochim. Biophys. Acta 1865, 488–498 (2017).
Saal, L. H. et al. Poor prognosis in carcinoma is associated with a gene expression signature of aberrant PTEN tumor suppressor pathway activity. Proc. Natl Acad. Sci. USA 104, 7564–7569 (2007).
Shepherd, J. H. et al. The SOX11 transcription factor is a critical regulator of basal-like breast cancer growth, invasion, and basal-like gene expression. Oncotarget 7, 13106–13121 (2016).
Zvelebil, M. et al. Embryonic mammary signature subsets are activated in Brca1−/− and basal-like breast cancers. Breast Cancer Res. 15, R25 (2013).
Fernandez-Garcia, B. et al. Expression and prognostic significance of fibronectin and matrix metalloproteases in breast cancer metastasis. Histopathology 64, 512–522 (2014).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Malhotra, G. K. et al. The role of Sox9 in mouse mammary gland development and maintenance of mammary stem and luminal progenitor cells. BMC Dev. Biol. 14, 47 (2014).
Yalcin-Ozuysal, O. et al. Antagonistic roles of Notch and p63 in controlling mammary epithelial cell fates. Cell Death Differ. 17, 1600–1612 (2010).
Latil, M. et al. Cell-type-specific chromatin states differentially prime squamous cell carcinoma tumor-initiating cells for epithelial to mesenchymal transition. Cell Stem Cell 20, 191–204 (2017).
Lilja, A. M. et al. Clonal analysis of Notch1-expressing cells reveals the existence of unipotent stem cells that retain long-term plasticity in the embryonic mammary gland. Nat. Cell Biol. https://doi.org/10.1038/s41556-018-0108-1 (2018).
Signoretti, S. et al. p63 is a prostate basal cell marker and is required for prostate development. Am. J. Pathol. 157, 1769–1775 (2000).
Kurita, T., Medina, R. T., Mills, A. A. & Cunha, G. R. Role of p63 and basal cells in the prostate. Development 131, 4955–4964 (2004).
van Bokhoven, H. & McKeon, F. Mutations in the p53 homolog p63: allele-specific developmental syndromes in humans. Trends Mol. Med 8, 133–139 (2002).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Nguyen, H., Rendl, M. & Fuchs, E. Tcf3 governs stem cell features and represses cell fate determination in skin. Cell 127, 171–183 (2006).
Perl, A. K., Wert, S. E., Nagy, A., Lobe, C. G. & Whitsett, J. A. Early restriction of peripheral and proximal cell lineages during formation of the lung. Proc. Natl Acad. Sci. USA 99, 10482–10487 (2002).
Gonzalez-Roca, E. et al. Accurate expression profiling of very small cell populations. PLoS One 5, e14418 (2010).
Gautier, L., Cope, L., Bolstad, B. M. & Irizarry, R. A. affy—analysis of Affymetrix GeneChip data at the probe level. Bioinformatics 20, 307–315 (2004).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
Leek, J. T., Johnson, W. E., Parker, H. S., Jaffe, A. E. & Storey, J. D. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28, 882–883 (2012).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
McCarthy, D. J., Campbell, K. R., Lun, A. T. & Wills, Q. F. Scater: pre-processing, quality control, normalization and visualization of single-cell RNA-seq data in R. Bioinformatics 33, 1179–1186 (2017).
Acknowledgements
The authors acknowledge the animal house facility from ULB (Erasme campus). Sequencing was performed at the Brussels Interuniversity Genomics High Throughput core (www.brightcore.be) and the Genomics Core Leuven. The authors thank N. Dedoncker for help with single-cell RNA-seq library construction. C.B. is an investigator with WELBIO, A.W. is supported by a FNRS fellowship. M.F. is supported by a Télévie fellowship. A.V.K. is Maître de Recherches of the FNRS. A.S., D.B. and T.V. are supported by KU Leuven (SymBioSys, PFV/10/016), Stichting Tegen Kanker (2015-143) and FWO (postdoctoral fellow number 12W7318N, [PEGASUS]² Marie Skłodowska-Curie fellow number 12O5617N). The authors thank colleagues who provided reagents mentioned in the text, and J.-M. Vanderwinden for help with confocal imaging. This work was supported by the FNRS, a research grant from the Fondation Contre le Cancer, the ULB fondation, the Fond Gaston Ithier, the Télévie, the foundation Bettencourt Schueller, the foundation Baillet Latour, and the European Research Council (EXPAND).
Author information
Authors and Affiliations
Contributions
A.W. and C.B. designed the experiments and performed data analysis. A.W., S.M., M.F., A.C. and A.V.K performed the biological experiments. A.B. performed GSEA analysis. A.S. and T.V. performed single-cell RNA-seq data analysis and provided the related figures and methods. D.B. and T.V. performed single-cell RNA-seq processing and sequencing, and provided the related methods, A.D. provided technical support. C.D. provided technical support for cell sorting. A.W., A.S., C.B. and A.V.K. prepared the figures. C.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
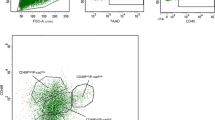
Supplementary Figure 1 Gating strategy for flow cytometry analysis.
a-g, Unicellular suspension of skin and mammary bud cells from Lgr5-IRES-GFP E14 embryos stained for Lin (CD31, CD45, CD140a) in APC and CD49f in PE were gated as shown in a to eliminate debris, doublets were discarded with gate shown in b followed by gate showed in c, the living cells were gated by DAPI dye exclusion as shown in d, the non-epithelial Lin positive cells were discarded in e. The CD49f Hi cells were gated as shown in f and the GFP + cells were gated as shown in g. h-o, Unicellular suspension of mammary cells from adult K8rtTA/TetOCre/ΔNp63-IRES-GFP, induced at P30 and analyzed at P45, stained for Lin (CD31, CD45, CD140a) in PE, CD24 in PECy7 and CD29 in APC, were gated as shown in h to eliminate debris, doublets were discarded with gates shown in i followed by gate shown in j, the living cells were gated by DAPI dye exclusion as shown in k, the non-epithelial Lin positive cells were discarded in L and the GFP + cells were gated as shown in m. CD24 and CD29 expression was studied in Lin- cells (n) or in YFP + cells (o). The CD24 + CD29Lo gate corresponds to luminal cells (LC), while CD24 + CD29Hi gate corresponds to basal cells (BC). The stromal population corresponds to the cells labelled due to the leakiness of the Tet-O-Cre, as described previously in reference 14.
Supplementary Figure 2 Transcriptional profiling of EMPs reveals their hybrid basal and luminal gene expression signature.
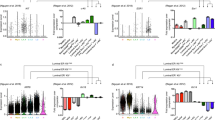
a, Graph of enrichment score of the top functional annotation clusters for genes overrepresented in EMPs compared to LCs. Their ranking is shown in parentheses. b, Graph representing mRNA expression measured by microarray analysis of upregulated genes in FACS-isolated BCs and Lgr5 cells (fold over LC), showing the genes of the axon guidance cluster enriched in EMPs and BCs. c, d, Gene ontology (GO) analysis of genes upregulated > 1.5-fold in both LCs and Lgr5 cells compared to BCs. Histogram represents enrichment score (c) and Benjamini corrected p-value (in log10 base) (d) of the top functional annotation clusters for genes overrepresented in EMPs compared to BCs. Their ranking is shown in parentheses. e, Graph representing mRNA expression measured by microarray analysis of upregulated genes in FACS-isolated LCs and Lgr5 cells (fold over BC), showing the enrichment of cell cycle related genes in EMPs and LCs. a,c,d, are derived from the list of genes upregulated in the comparison of the mean of n = 3 independent microarrays samples for Lgr5 and the mean of n = 2 independent microarrays samples for LCs and BCs. b,e, are representing the fold change of the mean of n = 3 independent microarrays for Lgr5 and the mean of n = 2 independent microarrays for LCs and BCs.
Supplementary Figure 3 Unsupervised clustering and gene ontology of scRNA-seq data set.
a, Unsupervised clustering using SC3 of LC (n = 73) and BC (n = 45) using clustering parameters k = 2. Heatmaps of the top 15 marker genes for each cluster and their corresponding normalized expression are displayed (AUC > 0.8 and Wilcoxon signed rank test FDR adjusted p-value < 0.01). Columns represent single cells, colour-coded by their respective lineage. UND (undetermined significance, n = 7) represents few FACS isolated CD29HiCD24 + with LC gene signature. b, c, Gene ontology analysis of EMP scRNA-seq. Histogram representing enrichment score (b) and Benjamini corrected p-value (in log10 base) (c) of the top functional annotation clusters for genes overrepresented in EMPs (n = 68). Their ranking is shown in parentheses. d, scRNAseq Sox10 expression represented on PCA plot (n = 193 cells): PCA was performed on the top 500 most variable genes in the scRNAseq data, every dot represents a single cell. Colouring represents the normalized expression of Sox10.
Supplementary Figure 4 Expression of luminal and basal markers in EMPs.
Marker genes are obtained from scRNAseq data using SC3 on the adult cell lineages (n = 118) with k = 2 and filtering marker genes with AUC > 0.8 and Wilcoxon signed rank test FDR adjusted p-value < 0.01, and only showing genes which are expressed in more than 50% of EMPs and 50% of either BCs or LCs. The heatmap colouring represents the proportion of cells with > 0 expression for that gene for each cell type (LC, BC, EMP; n = 73, n = 45, n = 68 respectively).
Supplementary Figure 5 Single cell RNA-seq using lower stringency quality control criteria.
Analysis of scRNAseq data of cells with > 2000 genes detected (n = 261) and including 1 row of the sorting plate with aberrant transcriptional profiles. a, Unsupervised clustering using SC3 on EMPs (n = 87), adult BC (n = 76) and LCs (n = 98) using clustering parameters k = 4. Heatmaps of the top 15 marker genes for each cluster and their corresponding normalized expression are displayed (AUC > 0.8 and Wilcoxon signed rank test FDR adjusted p-value < 0.01). Columns represent single cells, colour-coded by their respective lineage. b,c, Dimensionality reduction using t-Distributed Stochastic Neighbor Embedding (b) and Principal Component Analysis (c), every dot (n = 261) represents one cell with the colour representing either cell-type or the assigned SC3 cluster represented in a respectively. d, Scatter plot with the X-axis representing the proportion of BC-specific marker genes detected by SC3 (n = 53 cells) and the Y-axis LC-specific marker genes (n = 47 cells). Marker genes were selected to be expressed in at least 75% of the respective cell type and in less than 25% of the opposite cell type. The proportion of expressed markers is computed as the fraction of markers with > 0 expression over the total number of markers. Every dot (n = 261) represents one cell and are colour-coded according to cell type. Aberrant BC cells with low number of genes detected and aberrant LC/BC cells stemming from 1 row in the plate show a pseudo-hybrid signature and don’t cluster with their respective cell types.
Supplementary Figure 6 Unsupervised clustering of EMPs.
a, Heatmap showing scRNAseq expression within Lgr5-GFP + E14 subclusters including Lgr5 + EMP (n = 68) and stromal mesenchymal cells (n = 11). Marker genes are obtained from the scRNAseq data comparing LC (n = 73) to BC cells (n = 45) using SC3 with clustering parameter k = 2 and selecting marker genes with an AUC > 0.8 and a Wilcoxon signed rank test FDR adjusted p-value < 0.01. Expression of epithelial markers Krt14, Krt5 and Krt8 is also shown. The heatmap colouring represents the normalized expression in the scRNAseq where rows represent the top 27 marker genes for each cluster and columns represent EMP cells. b-e, Scatter plots depicting the correlation between the proportion of BC-specific (n = 53 cells) (b) and LC specific markers detected (n = 47 cells)(d) and total number of genes detected before (c) and after correction (e). Each dot (n = 193) represents a single cell, and colour represents cell type. Correction was performed using a robust linear model for each cell type using the rlm function in R, depicted for each population as a straight line with 95% confidence interval.
Supplementary Figure 7 Immunofluorescence analysis of LC reprograming following ΔNp63 expression.
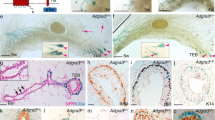
a-d, Immunofluorescence analysis K8, K14 and p63 (a), K8, K14 and GFP (b, c) and K14, GFP and Foxa1 (d) in WT (a) and in K8rtta/TetOCre/DNp63-IRES-GFP mice 2 weeks following the expression of p63-IRES-GFP in LCs (b-d) (6 mice analysed). Arrowheads point to hybrid GFP + cells, coexpressing luminal and basal markers. Arrow points to GFP + K14 + K8- cell at the basal membrane. e, Venn diagram showing the important and statistically significant overlap between the genes upregulated by 2 fold in BCs compared to LCs (adult basal signature) and the genes upregulated by p63 in LCs (p63 LC signature). The gene lists were derived by comparing the means of RNAseq data (n = 2 for p63, WT LC and WT BC). Enrichment p value was calculated using the hypergeometric test performed with R software without adjustment, to test if these 2 data sets of 802 and 2860 genes have a significantly higher overlap (295 genes) than 2 data sets of the same size chosen randomly.
Supplementary information
Supplementary Information
Supplementary Figures 1–7 and Supplementary Table legends
Supplementary Table 1
Statistics source data
Rights and permissions
About this article
Cite this article
Wuidart, A., Sifrim, A., Fioramonti, M. et al. Early lineage segregation of multipotent embryonic mammary gland progenitors. Nat Cell Biol 20, 666–676 (2018). https://doi.org/10.1038/s41556-018-0095-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0095-2
This article is cited by
-
Mechanosensitive TRPV4 channel guides maturation and organization of the bilayered mammary epithelium
Scientific Reports (2024)
-
Lineage plasticity enables low-ER luminal tumors to evolve and gain basal-like traits
Breast Cancer Research (2023)
-
p63: a crucial player in epithelial stemness regulation
Oncogene (2023)
-
Reconstruction of dynamic mammary mini gland in vitro for normal physiology and oncogenesis
Nature Methods (2023)
-
Reactivation of embryonic genetic programs in tissue regeneration and disease
Nature Genetics (2023)