Abstract
The maintenance of pluripotency requires coordinated expression of a set of essential genes. Using our recently established haploid human pluripotent stem cells (hPSCs), we generated a genome-wide loss-of-function library targeting 18,166 protein-coding genes to define the essential genes in hPSCs. With this we could allude to an intrinsic bias of essentiality across cellular compartments, uncover two opposing roles for tumour suppressor genes and link autosomal-recessive disorders with growth-retardation phenotypes to early embryogenesis. hPSC-enriched essential genes mainly encode transcription factors and proteins related to cell-cycle and DNA-repair, revealing that a quarter of the nuclear factors are essential for normal growth. Our screen also led to the identification of growth-restricting genes whose loss of function provides a growth advantage to hPSCs, highlighting the role of the P53–mTOR pathway in this context. Overall, we have constructed an atlas of essential and growth-restricting genes in hPSCs, revealing key aspects of cellular essentiality and providing a reference for future studies on human pluripotency.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Wutz, A. Haploid mouse embryonic stem cells: rapid genetic screening and germline transmission. Annu. Rev. Cell Dev. Biol. 30, 705–722 (2014).
Yilmaz, A., Peretz, M., Sagi, I. & Benvenisty, N. Haploid human embryonic stem cells: half the genome, double the value. Cell Stem Cell 19, 569–572 (2016).
Sagi, I. & Benvenisty, N. Haploidy in humans: an evolutionary and developmental perspective. Dev. Cell 41, 581–589 (2017).
Tarkowski, A. K., Witkowska, A. & Nowicka, J. Experimental parthenogenesis in the mouse. Nature 226, 162–165 (1970).
Leeb, M. & Wutz, A. Derivation of haploid embryonic stem cells from mouse embryos. Nature 479, 131–134 (2011).
Elling, U. et al. Forward and reverse genetics through derivation of haploid mouse embryonic stem cells. Cell Stem Cell 9, 563–574 (2011).
Yang, H. et al. Generation of genetically modified mice by oocyte injection of androgenetic haploid embryonic stem cells. Cell 149, 605–617 (2012).
Li, W. et al. Androgenetic haploid embryonic stem cells produce live transgenic mice. Nature 490, 407–411 (2012).
Li, W. et al. Genetic modification and screening in rat using haploid embryonic stem cells. Cell Stem Cell 14, 404–414 (2014).
Yang, H. et al. Generation of haploid embryonic stem cells from Macaca fascicularis monkey parthenotes. Cell Res. 23, 1187–1200 (2013).
Sagi, I. et al. Derivation and differentiation of haploid human embryonic stem cells. Nature 532, 107–111 (2016).
Carette, J. E. et al. Haploid genetic screens in human cells identify host factors used by pathogens. Science 326, 1231–1235 (2009).
Wang, T. et al. Identification and characterization of essential genes in the human genome. Science 350, 1096–1101 (2015).
Blomen, V. A. et al. Gene essentiality and synthetic lethality in haploid human cells. Science 350, 1092–1096 (2015).
Shalem, O. et al. Genome-scale CRISPR–Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Hart, T. et al. High-resolution CRISPR screens reveal fitness genes and genotype-specific cancer liabilities. Cell 163, 1515–1526 (2015).
Hart, T. et al. Evaluation and design of genome-wide CRISPR/SpCas9 knockout screens. Genes Genomes Genet. 7, 2719–2727 (2017).
OMIM—Online Mendelian inheritance in man (John Hopkins University School of Medicine, accessed 26 April 2017); https://www.omim.org/
Raya, Á. et al. Disease-corrected haematopoietic progenitors from Fanconi anaemia induced pluripotent stem cells. Nature 460, 53–59 (2009).
Tulpule, A. et al. Knockdown of Fanconi anemia genes in human embryonic stem cells reveals early developmental defects in the hematopoietic lineage. Blood 115, 3453–3462 (2010).
Henske, E. P., Jóźwiak, S., Kingswood, J. C., Sampson, J. R. & Thiele, E. A. Tuberous sclerosis complex. Nat. Rev. Dis. Prim. 2, 16035 (2016).
Walker, E. J. et al. Monoallelic expression determines oncogenic progression and outcome in benign and malignant brain tumors. Cancer Res. 72, 636–644 (2012).
Liu, J. et al. The oncogene c-Jun impedes somatic cell reprogramming. Nat. Cell Biol. 17, 856–867 (2015).
De Los Angeles, A. et al. Hallmarks of pluripotency. Nature 525, 469–478 (2015).
GTEx Consortium. The genotype-tissue expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Ravasi, T. et al. An atlas of combinatorial transcriptional regulation in mouse and man. Cell 140, 744–752 (2010).
Zhang, J. et al. Sall4 modulates embryonic stem cell pluripotency and early embryonic development by the transcriptional regulation of Pou5f1. Nat. Cell Biol. 8, 1114–1123 (2006).
Merkle, C. J., Karnitz, L. M., Henry-Sanchez, J. T. & Chen, J. Cloning and characterization of hCTF18, hCTF8, and hDCC1: human homologs of a Saccharomyces cerevisiae complex involved in sister chromatid cohesion establishment. J. Biol. Chem. 278, 30051–30056 (2003).
Low, S. C., Harney, J. W. & Berry, M. J. Cloning and functional characterization of human selenophosphate synthetase, an essential component of selenoprotein synthesis. J. Biol. Chem. 270, 21659–21664 (1995).
Lenz, M. et al. Epigenetic biomarker to support classification into pluripotent and non-pluripotent cells. Sci. Rep. 5, 8973 (2015).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Guertin, D. A. & Sabatini, D. M. The pharmacology of mTOR inhibition. Sci. Signal. 2, pe24 (2009).
Fiorenzano, A. et al. Cripto is essential to capture mouse epiblast stem cell and human embryonic stem cell pluripotency. Nat. Commun. 7, 12589 (2016).
Avior, Y., Sagi, I. & Benvenisty, N. Pluripotent stem cells in disease modelling and drug discovery. Nat. Rev. Mol. Cell Biol. 17, 170–182 (2016).
Sen, S., Takahashi, R., Rani, S., Freireich, E. J. & Stass, S. A. Expression of differentially phosphorylated Rb and mutant p53 proteins in myeloid leukemia cell lines. Leuk. Res. 17, 639–647 (1993).
ATCC. ATCC cell lines by gene mutation (American Type Culture Collection, Manassas, 2018); http://bit.ly/2GApeiM
Faderl, S. et al. The biology of chronic myeloid leukemia. N. Engl. J. Med. 341, 164–172 (1999).
Kim, J., Chu, J., Shen, X., Wang, J. & Orkin, S. H. An extended transcriptional network for pluripotency of embryonic stem cells. Cell 132, 1049–1061 (2008).
Assou, S. et al. A meta-analysis of human embryonic stem cells transcriptome integrated into a web-based expression atlas. Stem Cells 25, 961–973 (2007).
Masui, S. et al. Rex1/Zfp42 is dispensable for pluripotency in mouse ES cells. BMC Dev. Biol. 8, 45 (2008).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Merkle, F. T. et al. Human pluripotent stem cells recurrently acquire and expand dominant negative P53 mutations. Nature 545, 229–233 (2017).
Zhou, J. et al. mTOR supports long-term self-renewal and suppresses mesoderm and endoderm activities of human embryonic stem cells. Proc. Natl Acad. Sci. USA 106, 7840–7845 (2009).
Eiges, R. et al. Establishment of human embryonic stem cell-transfected clones carrying a marker for undifferentiated cells. Curr. Biol. 11, 514–518 (2001).
Sagi, I., Egli, D. & Benvenisty, N. Identification and propagation of haploid human pluripotent stem cells. Nat. Protoc. 11, 2274–2286 (2016).
Wang, T., Lander, E. S. & Sabatini, D. M. Viral packaging and cell culture for CRISPR-based screens. Cold Spring Harb. Protoc. http://doi.org/gcw7js (2016).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Binder, J. X. et al. COMPARTMENTS: unification and visualization of protein subcellular localization evidence. Database 2014, bau012 (2014).
Acknowledgements
The authors thank E. Meshorer and all members of The Azrieli Center for Stem Cells and Genetic Research for their input and critical reading of the manuscript. The authors also thank O. Yanuka, T. Golan-Lev and A. Petcho for assistance with tissue culture. This work was partially supported by the US–Israel Binational Science Foundation (grant no. 2015089), by the Israel Science Foundation (grant no. 494/17) and by the Azrieli Foundation. A.Y. is supported by the Lady Davis Postdoctoral Fellowship. I.S. is supported by the Adams Fellowship Program of the Israel Academy of Sciences and Humanities, and N.B. is the Herbert Cohn Chair in Cancer Research.
Author information
Authors and Affiliations
Contributions
A.Y., M.P. and N.B. designed the experiments, interpreted the data and wrote the manuscript, with input from all authors. A.Y. performed the experiments and analysed the data. M.P. performed the bioinformatics analyses. A.A. assisted with the bioinformatics analyses. I.S. assisted with the characterization of haploid hESCs. N.B. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Characterization of a genome-wide CRISPR-Cas9 screen in haploid hPSCs.
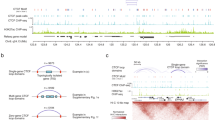
a, Analysis of the abundance of sgRNAs within the haploid hESC population at multiple time points after the co-delivery of sgRNAs and Cas9. Shown is a correlation matrix for all time points with correlation coefficients on the right diagonal and the scatter plots on the left diagonal of the matrix (n=181,131 gRNAs). b, CRISPR score formula. c, Scatter plot of CRISPR scores between two replicate experiments (n=18,166 genes, r = Pearson’s correlation coefficient). d, Chromosomal distribution of growth-restricting genes. Shown are growth-restricting genes (blue lines) and all other genes targeted in the library (grey lines).
Supplementary Figure 2 Comparison of essential genes across various studies.
a, Venn diagram showing the overlap of cell-essential genes identified in the current study in hESCs and in near-haploid KBM7 leukemic cells by Wang et al. 13 and by Blomen et al.14 and in several immortalized and cancer lines from different tissues of origin by Hart et al.16 (n=15,899 genes). b, PCA plot demonstrating the separation of genes in different studies. While the current study and Wang et al. study13 used the same sgRNA library, Blomen et al.14 used a transposon-mediated gene-trap mutant library and Hart et al.16 used two independent sgRNA libraries. c, Analysis of first percentile CRISPR scores across autosomes of near-haploid KBM7 cells (in which chromosome 8 is diploid), three diploid cell lines used in Wang et al. study6 and haploid hESCs from the current study. Chromosome 8 is highlighted in red. Dashed lines indicate the threshold corresponding to p-value of 0.025 on either side of the distribution of first percentile CRISPR scores (two-tailed Z-test, n=22 chromosomes, *P=0.02). d, -log10 P-values for chromosome 8 in the distribution of the first percentile CRISPR scores across autosomes in different cell lines. In near-haploid KBM7 cells, the first percentile CRISPR score for the diploid chromosome 8 is significantly different from those of the other autosomes (marked by asterisk), suggesting more efficient loss-of-function analysis in haploid chromosomes (two-tailed Z-test, n=22 chromosomes, *P=0.02).
Supplementary Figure 3 Analysis of cell-essential genes for their cellular compartmentalization, disease association and tumor formation.
a, Pie charts for each cellular compartment in hESCs demonstrating both enriched and depleted genes. b, Fraction of genes that are essential and expressed in hESCs within each cellular compartment c, Fraction of essential genes within the total number of genes in each cellular compartment in KBM7 cells. d, Distribution of CRISPR scores of genes associated with AR human disorders with a growth retardation phenotype in hESCs (green curve) and in leukemic KBM7 cells (purple curve). e-f, Volcano plots representing Q-value and CRISPR score of the canonical oncogenes (e) and tumor suppressor genes (f) that did not show significant values (see Fig. 2e and f, KS-test, n=20 gRNAs, two biological replicates of 10 independent gRNAs per gene). g, Heatmaps showing the comparison of CRISPR scores of three groups of genes between hESCs and the cancer cell lines: Growth-restricting tumor suppressors in hESCs that lost this feature of growth-restriction in cancer cell lines (left heatmap), essential oncogenes in hESCs that lost their essentiality in cancer cell lines (middle heatmap) and nonessential oncogenes in hESCs that gained essentiality in cancer cell lines (right heatmap). Light blue and light red boxes designate significant FDR values (shown below the CRISPR score values, positive and negative values indicate enriched and depleted gRNAs, respectively) for growth-restricting and essential genes, respectively (KS-test, n=20 gRNAs).
Supplementary Figure 4 Analysis of the hESC-essential genes.
a, Scatter plot demonstrating CRISPR scores of hESC-enriched transcription factors in hESCs and KBM7 cells; in green: essential genes in KBM7 cells and hESCs, in red: essential genes only in hESCs. b, Mean ± s.e.m. of gRNA reads/gene over time in culture for validated genes (SALL4, DSCC1, VRTN, SEPHS1) along with their expression (normalized FPKM from RNA sequencing data25) in hPSCs and 27 somatic cell types. Lymphocytes and fibroblasts are transformed cell lines. CRISPR scores and P-values appear at the bottom (KS-test, n=20 gRNAs). c, Relative transcript levels (analysed by qRT–PCR) of targeted hESC-essential genes by siRNA knockdown as compared to a non-targeting siRNA control in diploid hESCs. Cells were harvested 24 hours after transfection of siRNAs for SALL4, 48 hours for VRTN and 72 hours for DSCC1 and SEPHS1 (n=3 biological replicates, mean ± s.e.m values). d, Relative wild-type transcript levels (analysed by qRT–PCR) of targeted hESC-essential genes in sgRNA-knockout lines 3 days after the delivery of sgRNAs and Cas9. Control lines received only Cas9 in the absence of a sgRNA. Values represent the averages of two biological replicate experiments with three technical replicates for each. e, Cell viability assay in hESC and KBM7 sgRNA-knockout lines for PIK3CA and PDIA4, two genes expressed in both cell types, 4 days after the delivery of sgRNAs and Cas9 (n=3 biological replicates for KBM7 cells, n=6 biological replicates for hESC cells, mean ± s.e.m values). f, Relative wild-type transcript levels (analysed by qRT–PCR) of targeted genes in sgRNA-knockout lines of KBM7 cells, 3 days after the delivery of sgRNAs and Cas9. Values represent the average of three technical replicates. g, Relative wild-type transcript levels (analysed by qRT–PCR) of targeted hESC-essential genes in sgRNA-knockout lines 3 days after the delivery of sgRNAs and Cas9. Values represent the average of two biological replicate experiments with three technical replicates for each. When applicable, source data are provided in Supplementary Table 4.
Supplementary Figure 5 Schematics illustrating distinct P53 target pathways.
Highlighted in red are significantly enriched growth-restricting genes that were found among the top 50 genes of this group in the haploid hESC screening. Enrichment fold-changes of TP53 target genes are shown on the right. Numbers indicate different P53 target pathways, red and black bars show high and low-ranking growth-restricting genes, respectively.
Supplementary Figure 6 Selective sensitivity of hESCs to IGF1-mTOR pathway.
a-d, Growth curves of IGF1-treated (red) vs. control (blue) diploid hESCs grown in mTESR1 (a), conditioned medium with 12% KSR (standard) (b), conditioned medium with 6% KSR (c), conditioned medium with 3% KSR (d) (n=4 biologically independent samples). Mean ± s.e.m. are presented and unpaired two-tailed t-test was applied (PmTeSR1=0.01, P12% KSR=2.7e-4, P6% KSR=8.4e-4, P3% KSR=3.6e-4, *P < 0.05; ***P < 0.001). e-g, Gating strategy for flow cytometry analysis of apoptotic cells (e, Annexin-V staining), cell-cycle profile (f, Propidium Iodide staining) and TRA-1-60+ pluripotent cells (g, TRA-1-60-staining). A minimum of 30,000 cells were analysed for each sample. h-i, Western blots demonstrating phosphorylated levels of mTORC1 target, 4E-BP1 and mTORC2 target AKT (top panels) upon treatment of hESCs with rapamycin and Torin 1 for short- (h) and long-term (i) treatments. To enable incubation with three primary antibodies separately, nitrocellulose membrane was cut where the dashed lines were indicated. Quantification of the western blots are shown in the bottom panels (n=3 biological replicates, mean ± s.d. values). As shown, rapamycin inhibits the activity of mTORC1 but not that of mTORC2 in hESCs in both short- and long-term treatments, whereas Torin 1 inhibits both complexes. j and k, Growth curves of rapamycin- or Torin 1-treated diploid hESC lines, REX1-GFP WA09 (j) and CSES9 (k), over the course of four days (n=3 biological replicates, mean ± s.e.m values). Shown are the values normalized to the untreated controls of the corresponding time points. l, Gene ontology analysis for the significantly down-regulated transcripts after four days of rapamycin treatment in hESCs (CSES9, blue) and in human foreskin fibroblasts (pink) (n=3 biological replicates). When applicable, source data are provided in Supplementary Table 4.
Supplementary Figure 7 Uncropped membranes.
Uncropped membranes used in the Supplementary Figure 6h-i. Sample IDs as follows: Gel1: 1) untreated (short term), biological replicate 1; 2) untreated (short term), biological replicate 2; 3) untreated (short term), biological replicate 3; 4) Rapamycin (1h), biological replicate 1; 5) Rapamycin (1h), biological replicate 2; 6) Rapamycin (1h), biological replicate 3; 7) Rapamycin (3h), biological replicate 1; 8) Rapamycin (3h), biological replicate 2; 9) Rapamycin (3h), biological replicate 3; 10) Torin 1 (1h), biological replicate 1; 11) Torin 1 (1h), biological replicate 2; 12) Torin 1 (1h), biological replicate 3. Gel2: 13) Untreated (long term), biological replicate 1; 14) Untreated (long term), biological replicate 2; 15) Untreated (long term), biological replicate 3; 16) Rapamycin (2 days), biological replicate 1; 17) Rapamycin (2 days), biological replicate 2; 18) Rapamycin (2 days), biological replicate 3.
Supplementary information
Supplementary Information
Supplementary Figures 1–7 and Supplementary Table legends
Supplementary Table 1
CRISPR scores and significance values of all genes in the loss-of-function mutant library of haploid hESCs.
Supplementary Table 2
List of essential genes linked to autosomal recessive disorders with growth retardation phenotypes.
Supplementary Table 3
List of hESC-essentialome genes.
Supplementary Table 4
Statistics source data.
Rights and permissions
About this article
Cite this article
Yilmaz, A., Peretz, M., Aharony, A. et al. Defining essential genes for human pluripotent stem cells by CRISPR–Cas9 screening in haploid cells. Nat Cell Biol 20, 610–619 (2018). https://doi.org/10.1038/s41556-018-0088-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0088-1