Abstract
For cancer cells to survive during extracellular matrix (ECM) detachment, they must inhibit anoikis and rectify metabolic deficiencies that cause non-apoptotic cell death. Previous studies in ECM-detached cells have linked non-apoptotic cell death to reactive oxygen species (ROS) generation, although the mechanistic underpinnings of this link remain poorly defined. Here, we uncover a role for receptor-interacting protein kinase 1 (RIPK1) in the modulation of ROS and cell viability during ECM detachment. We find that RIPK1 activation during ECM detachment results in mitophagy induction through a mechanism dependent on the mitochondrial phosphatase PGAM5. As a consequence of mitophagy, ECM-detached cells experience diminished NADPH production in the mitochondria, and the subsequent elevation in ROS levels leads to non-apoptotic death. Furthermore, we find that antagonizing RIPK1/PGAM5 enhances tumour formation in vivo. Thus, RIPK1-mediated induction of mitophagy may be an efficacious target for therapeutics aimed at eliminating ECM-detached cancer cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Frisch, S. M. & Francis, H. Disruption of epithelial cell-matrix interactions induces apoptosis. J. Cell. Biol. 124, 619–626 (1994).
Debnath, J. et al. The role of apoptosis in creating and maintaining luminal space within normal and oncogene-expressing mammary acini. Cell 111, 29–40 (2002).
Schafer, Z. T. et al. Antioxidant and oncogene rescue of metabolic defects caused by loss of matrix attachment. Nature 461, 109–113 (2009).
Buchheit, C. L., Rayavarapu, R. R. & Schafer, Z. T. The regulation of cancer cell death and metabolism by extracellular matrix attachment. Semin. Cell. Dev. Biol. 23, 402–411 (2012).
Buchheit, C. L., Weigel, K. J. & Schafer, Z. T. Cancer cell survival during detachment from the ECM: multiple barriers to tumour progression. Nat. Rev. Cancer 14, 632–641 (2014).
Buchheit, C. L., Angarola, B. L., Steiner, A., Weigel, K. J. & Schafer, Z. T. Anoikis evasion in inflammatory breast cancer cells is mediated by Bim-EL sequestration. Cell. Death Differ. 22, 1275–1286 (2015).
Rayavarapu, R. R. et al. The role of multicellular aggregation in the survival of ErbB2-positive breast cancer cells during extracellular matrix detachment. J. Biol. Chem. 290, 8722–8733 (2015).
Weigel, K. J. et al. CAF-secreted IGFBPs regulate breast cancer cell anoikis. Mol. Cancer Res. 12, 855–866 (2014).
Palorini, R. et al. Protein kinase A activation promotes cancer cell resistance to glucose starvation and anoikis. PLoS Genet. 12, e1005931 (2016).
Pedanou, V. E. et al. The histone H3K9 demethylase KDM3A promotes anoikis by transcriptionally activating pro-apoptotic genes BNIP3 and BNIP3L. eLife 5, e16844 (2016).
Jeon, S. M., Chandel, N. S. & Hay, N. AMPK regulates NADPH homeostasis to promote tumour cell survival during energy stress. Nature 485, 661–665 (2012).
Davison, C. A. et al. Antioxidant enzymes mediate survival of breast cancer cells deprived of extracellular matrix. Cancer Res. 73, 3704–3715 (2013).
Mason, J. A. et al. Oncogenic Ras differentially regulates metabolism and anoikis in extracellular matrix-detached cells. Cell. Death Differ. 23, 1271–1282 (2016).
Cho, Y. S. et al. Phosphorylation-driven assembly of the RIP1–RIP3 complex regulates programmed necrosis and virus-induced inflammation. Cell 137, 1112–1123 (2009).
Degterev, A. et al. Chemical inhibitor of nonapoptotic cell death with therapeutic potential for ischemic brain injury. Nat. Chem. Biol. 1, 112–119 (2005).
Hitomi, J. et al. Identification of a molecular signaling network that regulates a cellular necrotic cell death pathway. Cell 135, 1311–1323 (2008).
Sun, L. et al. Mixed lineage kinase domain-like protein mediates necrosis signaling downstream of RIP3 kinase. Cell 148, 213–227 (2012).
Vanden Berghe, T., Linkermann, A., Jouan-Lanhouet, S., Walczak, H. & Vandenabeele, P. Regulated necrosis: the expanding network of non-apoptotic cell death pathways. Nat. Rev. Mol. Cell Biol. 15, 135–147 (2014).
Fulda, S. Regulation of necroptosis signaling and cell death by reactive oxygen species. Biol. Chem. 397, 657–660 (2016).
Kim, Y. S., Morgan, M. J., Choksi, S. & Liu, Z. G. TNF-induced activation of the Nox1 NADPH oxidase and its role in the induction of necrotic cell death. Mol. Cell 26, 675–687 (2007).
Zhang, D. W. et al. RIP3, an energy metabolism regulator that switches TNF-induced cell death from apoptosis to necrosis. Science 325, 332–336 (2009).
Zhang, Y. et al. RIP1 autophosphorylation is promoted by mitochondrial ROS and is essential for RIP3 recruitment into necrosome. Nat. Commun. 8, 14329 (2017).
Degterev, A. et al. Identification of RIP1 kinase as a specific cellular target of necrostatins. Nat. Chem. Biol. 4, 313–321 (2008).
Moquin, D. M., McQuade, T. & Chan, F. K. CYLD deubiquitinates RIP1 in the TNFα-induced necrosome to facilitate kinase activation and programmed necrosis. PLoS ONE 8, e76841 (2013).
Jono, H. et al. NF-κB is essential for induction of CYLD, the negative regulator of NF-κB: evidence for a novel inducible autoregulatory feedback pathway. J. Biol. Chem. 279, 36171–36174 (2004).
Chen, N. & Debnath, J. IκB kinase complex (IKK) triggers detachment-induced autophagy in mammary epithelial cells independently of the PI3K–AKT–MTORC1 pathway. Autophagy 9, 1214–1227 (2013).
Tait, S. W. & Green, D. R. Mitochondria and cell death: outer membrane permeabilization and beyond. Nat. Rev. Mol. Cell Biol. 11, 621–632 (2010).
Ashkenazi, A. & Salvesen, G. Regulated cell death: signaling and mechanisms. Annu. Rev. Cell Dev. Biol. 30, 337–356 (2014).
Weinlich, R., Oberst, A., Beere, H. M. & Green, D. R. Necroptosis in development, inflammation and disease. Nat. Rev. Mol. Cell Biol. 18, 127–136 (2017).
Wang, H. et al. Mixed lineage kinase domain-like protein MLKL causes necrotic membrane disruption upon phosphorylation by RIP3. Mol. Cell 54, 133–146 (2014).
Overholtzer, M. et al. A nonapoptotic cell death process, entosis, that occurs by cell-in-cell invasion. Cell 131, 966–979 (2007).
Wang, Z., Jiang, H., Chen, S., Du, F. & Wang, X. The mitochondrial phosphatase PGAM5 functions at the convergence point of multiple necrotic death pathways. Cell 148, 228–243 (2012).
Kang, Y. J. et al. Regulation of NKT cell-mediated immune responses to tumours and liver inflammation by mitochondrial PGAM5-Drp1 signalling. Nat. Commun. 6, 8371 (2015).
Chen, G. et al. A regulatory signaling loop comprising the PGAM5 phosphatase and CK2 controls receptor-mediated mitophagy. Mol. Cell. 54, 362–377 (2014).
Lu, W. et al. Genetic deficiency of the mitochondrial protein PGAM5 causes a Parkinson’s-like movement disorder. Nat. Commun. 5, 4930 (2014).
Lu, W. et al. Mitochondrial protein PGAM5 regulates mitophagic protection against cell necroptosis. PLoS ONE 11, e0147792 (2016).
Jin, S. M. et al. Mitochondrial membrane potential regulates PINK1 import and proteolytic destabilization by PARL. J. Cell. Biol. 191, 933–942 (2010).
Lazarou, M., Jin, S. M., Kane, L. A. & Youle, R. J. Role of PINK1 binding to the TOM complex and alternate intracellular membranes in recruitment and activation of the E3 ligase Parkin. Dev. Cell. 22, 320–333 (2012).
Lazarou, M. et al. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 524, 309–314 (2015).
Vives-Bauza, C. et al. PINK1-dependent recruitment of Parkin to mitochondria in mitophagy. Proc. Natl Acad. Sci. USA 107, 378–383 (2010).
Nakatogawa, H., Suzuki, K., Kamada, Y. & Ohsumi, Y. Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat. Rev. Mol. Cell Biol. 10, 458–467 (2009).
Youle, R. J. & Narendra, D. P. Mechanisms of mitophagy. Nat. Rev. Mol. Cell Biol. 12, 9–14 (2011).
Basit, F. et al. Mitochondrial complex I inhibition triggers a mitophagy-dependent ROS increase leading to necroptosis and ferroptosis in melanoma cells. Cell. Death Dis. 8, e2716 (2017).
Jiang, L. et al. Reductive carboxylation supports redox homeostasis during anchorage-independent growth. Nature 532, 255–258 (2016).
Nutt, L. K. et al. Metabolic regulation of oocyte cell death through the CaMKII-mediated phosphorylation of caspase-2. Cell 123, 89–103 (2005).
Davison, C. A. et al. Multimodal optical, X-ray CT, and SPECT imaging of a mouse model of breast cancer lung metastasis. Curr. Mol. Med. 13, 368–376 (2013).
Uhlen, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Gyorffy, B., Surowiak, P., Budczies, J. & Lanczky, A. Online survival analysis software to assess the prognostic value of biomarkers using transcriptomic data in non-small-cell lung cancer. PLoS ONE 8, e82241 (2013).
Lanczky, A. et al. miRpower: a web-tool to validate survival-associated miRNAs utilizing expression data from 2178 breast cancer patients. Breast Cancer Res. Treat. 160, 439–446 (2016).
Hsu, H., Huang, J., Shu, H. B., Baichwal, V. & Goeddel, D. V. TNF-dependent recruitment of the protein kinase RIP to the TNF receptor-1 signaling complex. Immunity 4, 387–396 (1996).
Lin, J. et al. RIPK1 counteracts ZBP1-mediated necroptosis to inhibit inflammation. Nature 540, 124–128 (2016).
Takahashi, N. et al. RIPK1 ensures intestinal homeostasis by protecting the epithelium against apoptosis. Nature 513, 95–99 (2014).
Green, D. R. & Levine, B. To be or not to be? How selective autophagy and cell death govern cell fate. Cell 157, 65–75 (2014).
Egan, D. F. et al. Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science 331, 456–461 (2011).
Bernardini, J. P., Lazarou, M. & Dewson, G. Parkin and mitophagy in cancer. Oncogene 36, 1315–1327 (2017).
Chourasia, A. H. et al. Mitophagy defects arising from BNip3 loss promote mammary tumor progression to metastasis. EMBO Rep. 16, 1145–1163 (2015).
Gupta, A. et al. PARK2 depletion connects energy and oxidative stress to PI3K/Akt activation via PTEN S-nitrosylation. Mol. Cell 65, 999–1013 (2017).
Acknowledgements
We thank Veronica Schafer, all current and past Schafer lab members, James Clancy, Crislyn D’Souza-Schorey, H. Clay Conner, Athanasia Panopoulos, Ian Guldner, Siyuan Zhang, Rebecca Wingert and Paul Huber for helpful comments, experimental assistance and valuable discussion. We thank Jeffrey Hawk, Janet Hawk and Veronica Wende for helpful comments and support. We also thank Leta Nutt (St. Jude Children’s Research Hospital) for the Bax(−/−)/Bak(−/−) MEFs, Mary Ann McDowell (Notre Dame) for assistance with immunofluorescence, Kevin T. Vaughan (Notre Dame) for help with soft agar and William Archer (Notre Dame) for guidance with immunofluorescence. This work was supported by a Lee National Denim Day Research Scholar Grant (RSG-14-145-01-CSM) from the American Cancer Society (to Z.T.S.), a research grant from the Phi Beta Psi National Project (to Z.T.S.), a National Science Foundation Graduate Research Fellowship grant DGE 1313583 (to M.A.H.), the Coleman Foundation (Chicago, IL), and funds from Ron and Rosemarie Malanga.
Author information
Authors and Affiliations
Contributions
M.A.H., C.L.G., P.F., J.A.M., K.J.W., M.A.T., L.S., S.S., J.Z. and S.H. conducted experiments, analysed data and interpreted results. C.L., S.E.K., J.C.H. and M.O. assisted with the imaging and quantitation of necrotic cells and provided the ATG5 knockout MCF-10A cells. L.J. and R.J.D. provided H460 cells with IDH1 or IDH2 deficiencies and assisted with analysis and interpretation of related experiments. S.C. and W.M.L. carried out the in vivo studies in immunocompromised mice. M.A.H. and Z.T.S. wrote the manuscript with feedback from all other authors. Z.T.S. was responsible for conception/design of the project and overall study supervision.
Corresponding author
Ethics declarations
Competing interests
R.J.D. is an advisor for Agios Pharmaceuticals. All other authors declare no conflicts of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 NF-κB promotes CYLD transcription during ECM-detachment and overexpression of RIPK1 lowers cell viability.
(a) MCF-10A EV cells were grown in either Att or Det conditions for 24h. Lysates were immunoblotted as noted. (b) Human mammary epithelial cells (HMEC) were grown in either Att or Det conditions for 24h. Lysates were immunoblotted as noted. (c) 10A-Bcl2 cells were grown in either Att or Det conditions for 24h. Lysates were immunoblotted as noted. (d) 10A-Bcl2 cells were grown in Att or Det for 24h. Relative expression of CYLD was measured by qRT-PCR. n=3 independent biological samples. (e, f) 10A-Bcl2 cells were grown in Att or Det for 24h treated with either DMSO or Bay-117082 (10μM). Relative expression of CYLD was measured by qRT-PCR. n=3 independent biological samples (e). Lysates were immunoblotted as noted (f). (g) HMEC cells were grown in Att or Det conditions treated with DMSO or Nec1 (10μM) for 24h. Cell viability was measured using AlamarBlue. n=3 independent biological samples. (h) 10A-Bcl2 cells transfected with empty vector (EV) or RIPK1 wildtype (RIPK1 WT) were grown in Att for 24h. Lysates were immunoblotted as noted. (i) Cell viability was measured using AlamarBlue following transfection of Att 10A-Bcl2 cells after being treated with either DMSO or Nec1 (10μM) for 24h. n=3 independent biological samples. (j) Left panel, Bax/Bak null (Bax(-/-)/Bak(-/-)) MEFs transfected with empty vector (EV), RIPK1 wildtype (RIPK1 WT), or RIPK1 kinase mutant (RIPK1 K45A) were grown in Att for 24h. Lysates were immunoblotted as noted. (k) Cell viability was measured using AlamarBlue following transfection after being grown in Att for 24h. n=3 independent biological samples. (l) BT474, T47D, and MCF7 cells were grown in either Att or Det conditions for 24h. Lysates were immunoblotted as noted. (m) HeLa, HCT116, and H460 cells were grown in either Att or Det conditions for 24h. Lysates were immunoblotted as noted. All results are presented as means +/- SEM and statistical significance was calculated using a Student’s two tailed t-test. p<0.05 is statistically significant. All western blotting experiments were independently repeated a minimum of three times with similar results. Unprocessed original scans of blots are available in Supplementary Fig. 9.
Supplementary Figure 2 Caspase activation is not impacted by Nec1-mediated RIPK1 inhibition.
(a-f) BT474 (a), T47D (b), MCF7 (c), HeLa (d), HCT116 (e), and H460 (f) cells were grown in Att and Det after being treated with DMSO or Nec1 (10μM) for 48h. Caspase-3/7 activation was measured 48h after plating in Att and Det using the CaspaseGlo 3/7 assay. Results are presented as means +/- SEM and statistical significance was calculated using a Student’s two tailed t-test. Caspase activation is represented as a ratio of Det value over Att value. n=3 independent biological samples for each cell line. N.S., not significant. (g) 10A-Bcl2 cells were grown in soft agar and imaged each day for three days. Cells were scored as viable or necrotic as indicated. Results are presented as percentage of necrotic cells +/- SD. n=12 independent biological samples. (h) 10A-Bcl2 cells were grown in soft agar and imaged each day for three days. Cells were scored as viable or entotic as indicated. Results are presented as percentage of entotic cells +/- SD. n=12 independent biological replicates.
Supplementary Figure 3 RIPK3 nor MLKL are required for ECM-detachment induced RIPK1 signalling.
(a, b) MCF-10A pLKO (a,b), shRIPK3 (a) and shMLKL (b) cells were grown in Att and Det for 48h. Caspase-3/7 activation was measured 48h after plating in Att and Det using the CaspaseGlo 3/7 assay. Caspase activation is represented as a ratio of Det value over Att value. n=3 independent biological samples. (c) Immunoblotting against RIPK3 and β actin was used to confirm the RIPK3 knockdown (shRIPK3) in H460 cells grown in Det for 24h. Experiments were performed a minimum of three independent times with similar results. (d-f) Cell viability (d), caspase activation (e), and anchorage-independent growth in soft agar (f) was measured in Att and Det H460 pLKO and shRIPK3 cells at 24h. Viability was measured using AlamarBlue, caspase activation using CaspaseGlo 3/7 assay, and after 7 days, images of soft agar growth taken following INT-violet staining. Colony count was determined using ImageJ. n=6 independent biological replicates for soft agar and n=3 independent biological samples for viability and caspase measurements. (g) Immunoblotting against MLKL and β actin was used to confirm the MLKL knockdown (shMLKL) in H460 cells grown in Det for 24h. Experiments were performed a minimum of three independent times with similar results. (h-j) Cell viability (h), caspase activation (i), and anchorage-independent growth in soft agar (j) was measured in Att and Det H460 pLKO and shMLKL cells at 24h. Viability was measured using AlamarBlue, caspase activation using CaspaseGlo 3/7 assay, and after 7 days, images of soft agar growth taken following INT-violet staining. Colony count was determined using ImageJ. n=6 biological independent biological replicates for soft agar and n=3 independent biological samples for viability and caspase measurements. All results are presented as means +/- SEM and statistical significance was calculated using a Student’s two tailed t-test. N.S., not significant. Unprocessed original scans of blots are available in Supplementary Fig. 9
Supplementary Figure 4 PGAM5 is downstream of RIPK1-mediated loss of viability.
(a) Immunoblotting against PGAM5 and β actin was used to confirm the PGAM5 knockdown (shPGAM5) in MCF-10A cells grown in Att for 24h. Experiments were performed a minimum of three independent times with similar results. (b) MCF-10A pLKO and shPGAM5 cells transfected with empty vector (EV) or RIPK1 wildtype (RIPK1 WT) were grown in Att for 24h. Lysates were then immunoblotted for total RIPK1 and β actin. Experiments were performed a minimum of three independent times with similar results. (c, d) Cell viability was measured using AlamarBlue following transfection of MCF-10A pLKO (c) and shPGAM5 (d) cells after being grown in Att for 24h. n=3 independent biological samples. (e) Immunoblotting against PGAM5 and β actin was used to confirm the PGAM5 knockdown (shPGAM5) in H460 cells grown in Att for 24h. Experiments were performed a minimum of three independent times with similar results. (f) H460 pLKO and shPGAM5 cells transfected with empty vector (EV) or RIPK1 wildtype (RIPK1 WT) were grown in Att for 24h. Lysates were immunoblotted for Myc tag, and β actin to confirm the overexpression of Myc-tagged-RIPK1. Experiments were performed a minimum of three independent times with similar results. (g, h) Cell viability was measured using AlamarBlue following transfection H460 pLKO (g) and shPGAM5 (h) cells after being grown in Att for 24h. n=3 independent biological samples. All results are presented as means +/- SEM and statistical significance was calculated using a Student’s two tailed t-test. p<0.05 is statistically significant. N.S., not significant. Unprocessed original scans of blots are available in Supplementary Fig. 9.
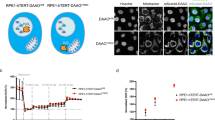
Supplementary Figure 5 Inhibition of RIPK1 or PGAM5 blocks ECM-detachment-induced mitophagy.
(a) H460 cells were grown in Att or Det for 24h treated with DMSO, Nec1 (10μM), BAFA (10nM) or CCCP (10μM). Cells were then cytospun onto slides, fixed, and stained with Mito-Tracker Red (200nM) (red) and DAPI (blue). Representative images of each condition are shown (40x). (b) Relative fluorescence measurements for each condition in (a) were determined using Fiji. For fluorescence calculations, n = 5 images. (c) H460 pLKO, shRIPK1, shPGAM5, and shPINK1 cells grown in Att or Det for 24h. Cells were then cytospun onto slides, fixed, and stained with Mito-Tracker Red (200nM) (red) and DAPI (blue). Representative images of each condition are shown (40x). (d) Relative fluorescence measurements for each condition in (c) were determined using Fiji. For fluorescence calculations, n = 5 images. (e) 10A-Bcl2 cells were grown in Att or Det for 24h treated with DMSO, Nec1 (10μM), or Nec1 (10μM) and CCCP (10μM). Cell viability was measured using AlamarBlue. Viability measurements are normalized to Att 10A-Bcl2 DMSO. n=3 independent biological samples. All results are presented as means +/- SEM and statistical significance was calculated using a Student’s two tailed t-test. p<0.05 is statistically significant. Scale bars, 100 μm.
Supplementary Figure 6 Deficiency of ATG5 halts RIPK1-dependent mitophagy during ECM-detachment and mitochondrial ROS generation during ECM-detachment is a consequence of RIPK1-mediated mitophagy.
(a) Immunoblotting against ATG5-ATG12 conjugate and GAPDH was used to confirm the ATG5 knockout clone in MCF-10A cells using CRISPR/Cas9. Experiments were performed a minimum of three independent times with similar results. (b) MCF-10A EV, ATG5KO, and HA-ATG5-R/E cells were grown in Att or Det for 24h treated with DMSO or Nec1 (10μM). Cell viability was measured using AlamarBlue. n=3 independent biological samples. (c) MCF-10A EV, ATG5KO, and HA-ATG5-R/E cells were grown in Att or Det for 24h treated with DMSO or Nec1 (10μM). Lysates were immunoblotted for Tom20 and β actin. Experiments were performed a minimum of three independent times with similar results. (d) 10A-Bcl2 cells were grown in Att or Det for 24h treated with DMSO, Nec1 (10μM), BAFA (10nM), MitoTEMPO (10μM) or CCCP (10μM). Cells were then cytospun onto slides, fixed, and stained with MitoSOX Red (5μM) (red) and DAPI (blue). Representative images of each condition are shown (40x). (e) Relative fluorescence measurements for each condition in (d) were determined using Fiji. For fluorescence calculations, n = 5 images. (f) H460 cells were grown in Att or Det for 24h treated with DMSO, Nec1 (10μM), BAFA (10nM), MitoTEMPO (10μM) or CCCP (10μM). Cells were then cytospun onto slides, fixed, and stained with MitoSOX Red (5μM) (red) and DAPI (blue). Representative images of each condition are shown (40x). (g) Relative fluorescence measurements for each condition in (f) were determined using Fiji. For fluorescence calculations, n = 5 images. All results are presented as means +/- SEM and statistical significance was calculated using a Student’s two tailed t-test. p<0.05 is statistically significant. N.S., not significant. Scale bars, 100 μm. Unprocessed original scans of blots are available in Supplementary Fig. 9.
Supplementary Figure 7 RIPK1 inhibition, but not ROS neutralization, rescues the loss of mitochondria during ECM-detachment as a consequence of IDH2.
(a) H460 EV cells were grown in Att or Det for 24h treated with DMSO, Nec1 (10μM), MitoTEMPO (10μM), or M. Malate (2.5mM). Cells were then cytospun onto slides, fixed, and stained with Mito-Tracker Red (200nM) (red) and DAPI (blue). Representative images of each condition are shown (40x). Experiments were performed a minimum of three independent times with similar results. (b) Relative fluorescence measurements for each condition in (a) were determined using Fiji. For fluorescence calculations, n = 5 images. (c) H460 IDH1KO cells were grown in Att or Det for 24h treated with DMSO, Nec1 (10μM), MitoTEMPO (10μM), or M. Malate (2.5mM). Cells were then cytospun onto slides, fixed, and stained with Mito-Tracker Red (200nM) (red) and DAPI (blue). Representative images of each condition are shown (40x). Experiments were performed a minimum of three independent times with similar results. (d) Relative fluorescence measurements for each condition in (c) were determined using Fiji. For fluorescence calculations, n = 5 images. (e) H460 IDH2KO cells were grown in Att or Det for 24h treated with DMSO, Nec1 (10μM), MitoTEMPO (10μM), or M. Malate (2.5mM). Cells were then cytospun onto slides, fixed, and stained with Mito-Tracker Red (200nM) (red) and DAPI (blue). Representative images of each condition are shown (40x). Experiments were performed a minimum of three independent times with similar results. (f) Relative fluorescence measurements for each condition in (e) were determined using Fiji. For fluorescence calculations, n = 5 images. All results are presented as means +/- SEM and statistical significance was calculated using a Student’s two tailed t-test. p<0.05 is statistically significant. N.S., not significant. Scale bars, 100 μm.
Supplementary Figure 8 IDH2 activity is required for cell survival when RIPK1 is inhibited.
(a-c) H460 EV (a), H460 IDH1KO (b), or H460 IDH2KO (c) cells were grown in Att or Det for 24h treated with DMSO, Nec1 (10μM), MitoTEMPO (10μM), or M. Malate (2.5mM). Cell viability was measured using AlamarBlue. n=3 independent biological samples. (d-f) H460 EV (d), H460 IDH1KO (e), or H460 IDH2KO (f) cells were grown in Att or Det for 24h treated with DMSO, Nec1 (10μM), or M. Malate (2.5mM). The ratio of NADPH/NADP+ was measured using NADP/NADPH-Glo assay. n=3 independent biological samples. (g) H460 EV (g), H460 IDH1KO (h), or H460 IDH2KO (i) cells were grown in Att or Det for 24h treated with H2O or H2O2. Cells were then cytospun onto slides, fixed, and stained with CellROX Green (5 μM) (green) and DAPI (blue). Representative images of each condition are shown (40x). n=3 independent biological samples. All results are presented as means +/- SEM and statistical significance was calculated using a Student’s two tailed t-test. p<0.05 is statistically significant. N.S., not significant. Scale bars, 20 μm.
Supplementary Figure 9
Unprocessed images of all gels and blots
Supplementary information
Supplementary Information
Supplementary Figures 1–9 and Supplementary Table legends.
Supplementary Table 1
Statistics source data.
Supplementary Table 2
Primer sequences.
Rights and permissions
About this article
Cite this article
Hawk, M.A., Gorsuch, C.L., Fagan, P. et al. RIPK1-mediated induction of mitophagy compromises the viability of extracellular-matrix-detached cells. Nat Cell Biol 20, 272–284 (2018). https://doi.org/10.1038/s41556-018-0034-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0034-2
This article is cited by
-
Extracellular matrix remodeling in tumor progression and immune escape: from mechanisms to treatments
Molecular Cancer (2023)
-
ATF4/CEMIP/PKCα promotes anoikis resistance by enhancing protective autophagy in prostate cancer cells
Cell Death & Disease (2022)
-
Mitophagy and reactive oxygen species interplay in Parkinson’s disease
npj Parkinson's Disease (2022)
-
The role of ROS in tumour development and progression
Nature Reviews Cancer (2022)
-
Mitochondrial fission links ECM mechanotransduction to metabolic redox homeostasis and metastatic chemotherapy resistance
Nature Cell Biology (2022)