Abstract
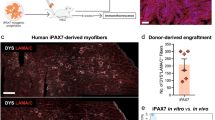
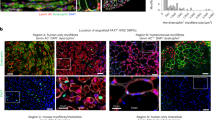
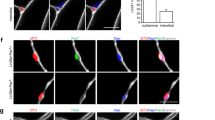
Human pluripotent stem cells (hPSCs) can be directed to differentiate into skeletal muscle progenitor cells (SMPCs). However, the myogenicity of hPSC-SMPCs relative to human fetal or adult satellite cells remains unclear. We observed that hPSC-SMPCs derived by directed differentiation are less functional in vitro and in vivo compared to human satellite cells. Using RNA sequencing, we found that the cell surface receptors ERBB3 and NGFR demarcate myogenic populations, including PAX7 progenitors in human fetal development and hPSC-SMPCs. We demonstrated that hPSC skeletal muscle is immature, but inhibition of transforming growth factor-β signalling during differentiation improved fusion efficiency, ultrastructural organization and the expression of adult myosins. This enrichment and maturation strategy restored dystrophin in hundreds of dystrophin-deficient myofibres after engraftment of CRISPR–Cas9-corrected Duchenne muscular dystrophy human induced pluripotent stem cell-SMPCs. The work provides an in-depth characterization of human myogenesis, and identifies candidates that improve the in vivo myogenic potential of hPSC-SMPCs to levels that are equal to directly isolated human fetal muscle cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Chong, J. J. H. et al. Human embryonic-stem-cell-derived cardiomyocytes regenerate non-human primate hearts. Nature 510, 273–277 (2014).
Schwartz, S. D. et al. Human embryonic stem cell-derived retinal pigment epithelium in patients with age-related macular degeneration and Stargardt’s macular dystrophy: follow-up of two open-label phase 1/2 studies. Lancet 385, 509–516 (2014).
Steinbeck, J. A. et al. Optogenetics enables functional analysis of human embryonic stem cell-derived grafts in a Parkinson’s disease model. Nat. Biotechnol. 33, 204–209 (2015).
Hrvatin, S. et al. Differentiated human stem cells resemble fetal, not adult, β cells. Proc. Natl Acad. Sci. USA 111, 3038–3043 (2014).
Witty, A. D. et al. Generation of the epicardial lineage from human pluripotent stem cells. Nat. Biotechnol. 32, 1026–1035 (2014).
Yang, X., Pabon, L. & Murry, C. E. Engineering adolescence: maturation of human pluripotent stem cell-derived cardiomyocytes. Circ. Res. 114, 511–523 (2014).
Jonsson, M. K. et al. Application of human stem cell-derived cardiomyocytes in safety pharmacology requires caution beyond hERG. J. Mol. Cell. Cardiol. 52, 998–1008 (2012).
Borchin, B., Chen, J. & Barberi, T. Derivation and FACS-mediated purification of PAX3+/PAX7+ skeletal muscle precursors from human pluripotent stem cells. Stem Cell Rep. 21, 620–631 (2013).
Xu, C. et al. A zebrafish embryo culture system defines factors that promote vertebrate myogenesis across species. Cell 155, 909–921 (2013).
Chal, J. et al. Differentiation of pluripotent stem cells to muscle fiber to model Duchenne muscular dystrophy. Nat. Biotechnol. 33, 962–969 (2015).
Shelton, M. et al. Derivation and expansion of PAX7-positive muscle progenitors from human and mouse embryonic stem cells. Stem Cell Rep. 3, 516–529 (2014).
Chal, J. et al. Generation of human muscle fibers and satellite-like cells from human pluripotent stem cells in vitro. Nat. Protoc. 11, 1833–1850 (2016).
Swartz, E. W. et al. A novel protocol for directed differentiation of C9orf72-associated human induced pluripotent stem cells into contractile skeletal myotubes. Stem Cells Transl. Med. 5, 1461–1472 (2016).
Abujarour, R. et al. Myogenic differentiation of muscular dystrophy-specific induced pluripotent stem cells for use in drug discovery. Stem Cells Transl. Med. 3, 149–160 (2014).
Kimura, E. et al. Cell-lineage regulated myogenesis for dystrophin replacement: a novel therapeutic approach for treatment of muscular dystrophy. Hum. Mol. Genet. 17, 2507–2517 (2008).
Darabi, R. et al. Human ES- and iPS-derived myogenic progenitors restore dystrophin and improve contractility upon transplantation in dystrophic mice. Cell Stem Cell 10, 610–619 (2012).
Darabi, R. et al. Functional skeletal muscle regeneration from differentiating embryonic stem cells. Nat. Med. 14, 134–143 (2008).
Sambasivan, R. & Tajbakhsh, S. Skeletal muscle stem cell birth and properties. Semin. Cell Dev. Biol. 18, 870–882 (2007).
Xi, H. et al. In vivo human somitogenesis guides somite development from hPSCs. Cell Rep. 18, 1573–1585 (2017).
Loh, K. M. et al. Mapping the pairwise choices leading from pluripotency to human bone, heart, and other mesoderm cell types. Cell 166, 451–467 (2016).
Yin, H., Price, F. & Rudnicki, M. A. Satellite cells and the muscle stem cell niche. Physiol. Rev. 93, 23–67 (2013).
Biressi, S. et al. Myf5 expression during fetal myogenesis defines the developmental progenitors of adult satellite cells. Dev. Biol. 379, 195–207 (2013).
Tierney, M. T. et al. Autonomous extracellular matrix remodeling controls a progressive adaptation in muscle stem cell regenerative capacity during development. Cell Rep. 14, 1940–1952 (2016).
Sacco, A., Doyonnas, R., Kraft, P., Vitorovic, S. & Blau, H. M. Self-renewal and expansion of single transplanted muscle stem cells. Nature 456, 502–506 (2008).
Young, C. S. et al. A single CRISPR–Cas9 deletion strategy that targets the majority of DMD patients restores dystrophin function in hiPSC-derived muscle cells. Cell Stem Cell 18, 533–540 (2016).
Tierney, M. T. & Sacco, A. Satellite cell heterogeneity in skeletal muscle homeostasis. Trends Cell Biol. 26, 434–444 (2016).
Castiglioni, A. et al. Isolation of progenitors that exhibit myogenic/osteogenic bipotency in vitro by fluorescence-activated cell sorting from human fetal muscle. Stem Cell Rep. 2, 92–106 (2014).
Montarras, D. et al. Direct isolation of satellite cells for skeletal muscle regeneration. Science 309, 2064–2067 (2005).
Cerletti, M. et al. Highly efficient, functional engraftment of skeletal muscle stem cells in dystrophic muscles. Cell 134, 37–47 (2008).
Godfrey, C. et al. How much dystrophin is enough: the physiological consequences of different levels of dystrophin in the mdx mouse. Hum. Mol. Genet. 24, 4225–4237 (2015).
Sharp, P. S., Bye-a-Jee, H. & Wells, D. J. Physiological characterization of muscle strength with variable levels of dystrophin restoration in mdx mice following local antisense therapy. Mol. Ther. 19, 165–171 (2011).
Chan, Y. & Walmsley, R. P. Learning and understanding the Kruskal–Wallis one-way analysis-of-variance-by-ranks test for differences among three or more independent groups. Phys. Ther. 77, 1755–1762 (1997).
Choi, I. Y. et al. Concordant but varied phenotypes among Duchenne muscular dystrophy patient-specific myoblasts derived using a human iPSC-based model. Cell Rep. 15, 2301–2312 (2016).
Xu, X. et al. Human satellite cell transplantation and regeneration from diverse skeletal muscles. Stem Cell Rep. 5, 419–434 (2015).
Cahan, P. & Daley, G. Q. Origins and implications of pluripotent stem cell variability and heterogeneity. Nat. Rev. Mol. Cell Biol. 14, 357–368 (2013).
Osafune, K. et al. Marked differences in differentiation propensity among human embryonic stem cell lines. Nat. Biotechnol. 26, 313–315 (2008).
Cusella-De Angelis, M. G. et al. Differential response of embryonic and fetal myoblasts to TGF beta: a possible regulatory mechanism of skeletal muscle histogenesis. Development 120, 925–933 (1994).
Schiaffino, S., Rossi, A. C., Smerdu, V., Leinwand, L. A. & Reggiani, C. Developmental myosins: expression patterns and functional significance. Skelet. Muscle 5, 22 (2015).
Pagliuca, F. W. et al. Generation of functional human pancreatic β cells in vitro. Cell 159, 428–439 (2104).
Maroof, A. M. et al. Directed differentiation and functional maturation of cortical interneurons from human embryonic stem cells. Cell Stem Cell 12, 559–572 (2013).
Evseenko, D. et al. Mapping the first stages of mesoderm commitment during differentiation of human embryonic stem cells. Proc. Natl Acad. Sci. USA 107, 13742–13747 (2010).
Alexander, M. S. et al. CD82 is a marker for prospective isolation of human muscle satellite cells and is linked to muscular dystrophies. Cell Stem Cell 19, 800–807 (2016).
Figeac, N., Serralbo, O., Marcelle, C. & Zammit, P. S. ErbB3 binding protein-1 (Ebp1) controls proliferation and myogenic differentiation of muscle stem cells. Dev. Biol. 386, 135–151 (2014).
Golding, J. P., Calderbank, E., Partridge, T. A. & Beauchamp, J. R. Skeletal muscle stem cells express anti-apoptotic ErbB receptors during activation from quiescence. Exp. Cell Res. 313, 341–356 (2007).
Van Ho, A. T. et al. Neural crest cell lineage restricts skeletal muscle progenitor cell differentiation through neuregulin1–ErbB3 signaling. Dev. Cell 21, 273–287 (2011).
Deponti, D. et al. The low-affinity receptor for neurotrophins p75NTR plays a key role for satellite cell function in muscle repair acting via RhoA. Mol. Biol. Cell 20, 3620–3627 (2009).
Esteves de Lima, J., Bonnin, M. A., Birchmeier, C. & Duprez, D. Muscle contraction is required to maintain the pool of muscle progenitors via YAP and NOTCH during fetal myogenesis. eLife 5, e15593 (2016).
Manceau, M. et al. Myostatin promotes the terminal differentiation of embryonic muscle progenitors. Genes Dev. 22, 668–681 (2008).
Arnett, A. L. et al. Adeno-associated viral (AAV) vectors do not efficiently target muscle satellite cells. Mol. Ther. Methods Clin. Dev. 1, 14038 (2014).
Quarta, M. et al. An artificial niche preserves the quiescence of muscle stem cells and enhances their therapeutic efficacy. Nat. Biotechnol. 34, 752–759 (2016).
Rommel, C. et al. Mediation of IGF-1-induced skeletal myotube hypertrophy by PI(3)K/Akt/mTOR and PI(3)K/Akt/GSK3 pathways. Nat. Cell Biol. 3, 1009–1013 (2001).
Young, C. S. et al. Creation of a novel humanized dystrophic mouse model of Duchenne muscular dystrophy and application of a CRISPR/Cas9 gene editing therapy. J. Neuromuscul. Dis. 4, 139–145 (2017).
Nguyen, H. T. & Morris, G. E. Use of epitope libraries to identify exon-specific monoclonal antibodies for characterization of altered dystrophins in muscular dystrophy. Am. J. Hum. Genet. 52, 1057–1066 (1993).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
da Huang, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
da Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Lapan, A. D., Rozkalne, A. & Gussoni, E. Human fetal skeletal muscle contains a myogenic side population that expresses the melanoma cell-adhesion molecule. Hum. Mol. Genet. 21, 3668–3680 (2012).
Inman, G. J. et al. SB-431542 is a potent and specific inhibitor of transforming growth factor-β superfamily type I activin receptor-like kinase (ALK) receptors ALK4, ALK5, and ALK7. Mol. Pharmacol. 62, 65–74 (2002).
Tojo, M. et al. The ALK-5 inhibitor A-83-01 inhibits Smad signaling and epithelial-to-mesenchymal transition by transforming growth factor-β. Cancer Sci. 96, 791–800 (2005).
Acknowledgements
We thank S. Younesi, J. Marshall, M. Emami, E. Korsakova, K. Saleh, V. Rezek, J. Wen and C. Kumagai-Cresse for helping with stem cell culture and mouse experiments. Furthermore, we thank E. Mokhonova for performing the western blots. We also thank J. Morgan’s laboratory for training on cell engraftments. The following cores were used: CDMD Muscle Phenotyping and Imaging Core, High Throughput and Cell Repository Core, and Bioinformatics and Genomics Core; the Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research at UCLA (BSCRC) Flow Cytometry Core; the UCLA Technology Center for Genomics and Bioinformatics, JCCC Electron Microscopy Core, the CFAR Flow Cytometry Core (NIH P30CA016042, 5P30AI028697); and the UCLA Humanized Mouse Core (CFAR, NIAID AI028697). M.R.H. is the recipient of a BSCRC and Schaffer Fellowship, a CDMD-Cure Duchenne Fellowship and a CDMD-NIH Paul Wellstone Center Training Fellowship (U54 AR052646). D.E. was funded by an NIH grant K01AR061415, Department of Defense (DoD) grant W81XWH-13-1-0465 and California Institute for Regenerative Medicine (CIRM) grant RB5-07230. Funding was provided by the National Institute of Arthritis and Musculoskeletal and Skin Diseases (NIAMS) R01AR064327 to A.D.P., and NIAMS 5P30AR05723 to M.J.S., the CDMD at UCLA, the NIH/NCATS, the UCLA CTSI (UL1TR000124), the BSCRC Research Award (A.D.P.), the Rose Hills Foundation Research Award to A.D.P., and a CIRM Inception and CIRM Quest (DISC1-08823 and DISC2-08824 to A.D.P.).
Author information
Authors and Affiliations
Contributions
M.R.H., B.V.H. and A.D.P. conceived the study. M.R.H., B.V.H. and M.J.S. designed the experiments. A.E., B.V.H., S.F.N. and M.R.H. analysed the data. M.R.H., B.V.H., J.H., K.P., W.F., M.J., H.X. and C.S.Y. conducted the experiments. M.R.H. and A.D.P. wrote the manuscript. M.R.H., A.D.P., B.V.H., J.H., K.P., C.S.Y., M.J.S. and S.F.N. edited the manuscript. Funding acquisition, M.R.H., A.D.P. and D.E.; Resources, D.E.; A.D.P. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figures 1–7 and Supplementary References.
Supplementary Table 1
Engraftment quantification of all human fetal and hPSC skeletal muscle in all mdx-NSG mice.
Supplementary Table 2
Gene Lists of key biological processes upregulated by CD31-CD45-NCAM+ cultured fetal muscle cells and HNK1-NCAM+ hPSC-SMPCs.
Supplementary Table 3
Gene Lists of key biological processes upregulated by directly-isolated fetal muscle cells and hPSC-SMPCs.
Supplementary Table 4
Gene Lists of key biological processes upregulated by directly-isolated fetal muscle cells and CD31-CD45-NCAM+ cultured fetal muscle cells.
Supplementary Table 5
Gene primer lists.
Supplementary Table 6
Antibody lists for IF and FACS.
Supplementary Table 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Hicks, M.R., Hiserodt, J., Paras, K. et al. ERBB3 and NGFR mark a distinct skeletal muscle progenitor cell in human development and hPSCs. Nat Cell Biol 20, 46–57 (2018). https://doi.org/10.1038/s41556-017-0010-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-017-0010-2
This article is cited by
-
Motor neurons and endothelial cells additively promote development and fusion of human iPSC-derived skeletal myocytes
Skeletal Muscle (2024)
-
The adult environment promotes the transcriptional maturation of human iPSC-derived muscle grafts
npj Regenerative Medicine (2024)
-
Nanoparticles systemically biodistribute to regenerating skeletal muscle in DMD
Journal of Nanobiotechnology (2023)
-
3D human induced pluripotent stem cell–derived bioengineered skeletal muscles for tissue, disease and therapy modeling
Nature Protocols (2023)
-
Regenerating human skeletal muscle forms an emerging niche in vivo to support PAX7 cells
Nature Cell Biology (2023)