Abstract
Assays for the molecular detection of nucleic acids are typically constrained by the level of multiplexing (this is the case for the quantitative polymerase chain reaction (qPCR) and for isothermal amplification), turnaround times (as with microarrays and next-generation sequencing), quantification accuracy (isothermal amplification, microarrays and nanopore sequencing) or specificity for single-nucleotide differences (microarrays and nanopore sequencing). Here we show that a portable and battery-powered PCR assay performed in a toroidal convection chamber housing a microarray of fluorescently quenched oligonucleotide probes allows for the rapid and sensitive quantification of multiple DNA targets with single-nucleotide discrimination. The assay offers a limit of detection of 10 DNA copies within 30 min of turnaround time and a dynamic range spanning 4 orders of magnitude of DNA concentration, and we show its performance by detecting 20 genomic loci and 30 single-nucleotide polymorphisms in human genomic DNA samples, and 15 bacterial species in clinical isolates. Portable devices for the fast and highly multiplexed detection of nucleic acids may offer advantages in point-of-care diagnostics.
Similar content being viewed by others
Main
The detection of specific DNA sequences is central to precision medicine, from pathogen identification to the risk assessment of human genetic diseases to disease prognosis. As the understanding of the genomics of disease improves, translation of this scientific knowledge into actionable clinical practice will be facilitated by DNA diagnostic systems that are simultaneously fast, affordable, sensitive, massively multiplexed, quantitative and easy to operate.
Since the early 2000s, however, DNA-detection technologies have bifurcated into either massively multiplexed but slow systems (next-generation sequencing (NGS)1,2 and microarrays3,4), or rapid but lowly multiplexing assays (quantitative PCR (qPCR)5,6 and isothermal amplification7,8). Two notable exceptions to the slow but powerful or fast but limited trade-off are the Biofire FilmArray multiplex PCR system9 and the Oxford Nanopore high-throughput sequencing system10 (Table 1). However, these two technologies do not offer accurate quantification and are unable to reliably recognize single-nucleotide differences, which are relevant for genetic and metabolic risk assessments11,12, pharmacogenetic drug dosing13, cancer-therapy selection14 and antimicrobial resistance to infectious disease15,16.
Here we present the design and performance of a toroidal chamber for the detection of DNA via PCR. The device allows for scalable and massive multiplexing, rapid turnaround times, single-nucleotide discrimination and precise quantification in a portable, affordable and battery-powered instrument that uses closed consumables to minimize contamination risks (Table 1). The toroidal PCR system is enabled by two technologies: (1) reliable convection PCR using an annular reaction chamber, and (2) a pre-quenched microarray that allows multiplexed readout via spatial separation. Convection PCR achieves thermal cycling of a PCR reaction mixture using passive movement of fluid owing to temperature-induced density differences. Convection PCR was initially proposed and experimentally demonstrated in 2002 using capillary tubes heated to 95 °C at the bottom17. Although a number of publications have explored alternative chambers for convection PCR18,19,20,21, so far demonstrations of real-time amplification in convection PCR have been limited to single-plex assays.
Importantly, the pre-quenched microarray is integrated in the consumable allowing probe hybridization to occur concurrently with convection PCR amplification. Consequently, unlike standard DNA microarrays that visualize spot endpoint fluorescence after 16 h of hybridization, the toroidal PCR system performs real-time detection and quantification of DNA in under 30 min. The quantitative dynamic range is over four logs, with a limit of detection of under ten genomic DNA (gDNA) copies. Use of toehold probes22 or X-probes23 for the microarray further provides single-nucleotide discrimination that enables robust single-nucleotide polymorphism (SNP) genotyping.
Results
Toroidal convection PCR mechanism and chip design
Rayleigh–Bénard thermal convection is the physical principle that as an aqueous solution is heated, it becomes less dense and rises due to gravity; in contrast, a colder solution is denser and falls. In the toroidal PCR system, we designed a chip that includes an annular (donut shaped) reaction chamber in which the DNA sample and PCR reagents are loaded. The chip is then sealed and vertically mounted on one side to a 95 °C heater and on the other side to a 60 °C heater (Fig. 1a,b). The fluid at the 95 °C zone becomes less dense and rises, and subsequently moves to the 60 °C zone where it is cooled and falls to the bottom of the chamber due to increased density.
a, Thermal cycling based on passive fluidic flow, using Rayleigh–Bénard convection. PCR mixture comprising DNA template, primers, DNA polymerase and deoxynucleoside triphosphates (dNTPs) is loaded via the bottom green loading port into the donut-shaped reaction chamber on the toroidal PCR chip. Subsequently, the left side of the chip is heated to 95 °C and the right side to 60 °C. Because water is less dense at higher temperatures, the reaction solution will rise on the left side as it is heated and fall on the right side as it is cooled, achieving autonomous thermal cycles with only two constant-temperature heaters. For each lap around the donut-shaped ‘racetrack’, amplicons have a chance to hybridize to the surface-functionalized probe array printed in the 60 °C zone. b, Picture of a solution of food colouring being loaded into the toroidal PCR chip, with a US dime for size comparison. c, Schematic of pre-quenched microarray probes printed in the 60 °C zone. At each spot, a three-stranded DNA complex is printed in which the fluorophore-functionalized oligo is colocalized to a quencher-labelled oligo via an unmodified bridge oligo. Green regions show gene-specific DNA sequences and blue regions show universal DNA sequences. The double-stranded DNA (dsDNA) amplicon is denatured in the 95 °C zone to become single-stranded DNA (ssDNA). The ssDNA amplicon then binds to and displaces the quencher-labelled oligo from the surface probe, resulting in a local fluorescence increase. d, Overview of the initial chip imaging setup using a Zeiss Axiovert 1M epifluorescence microscope. e, Fluorescence images of the chip microarray zone before (top) and after (bottom) PCR amplification. The field of view is 2 mm wide and each spot has a diameter of 150 µm. Here, probes are printed as adjacent duplicate pairs. The top left and bottom right pair are positive (Pos.) controls, and the bottom middle pairs are negative (Neg.) controls. The remaining 20 pairs of probes target different single-copy regions of the human genome. For this experiment, 10 ng of NA18537 human gDNA (roughly 3,000 haploid genome copies) was used as input, and 20 pairs of corresponding PCR primers were introduced in the reaction to simultaneously amplify all loci of interest within the toroidal PCR chip. f, Time-based spot fluorescence for the experiment shown in e. Plotted is the averaged relative fluorescence from each pair of spots. The rs numbers in each subfigure indicate the human SNP marker included within each amplicon region. Our probes in this experiment only detect the presence of the amplicon and not specific SNP alleles (see also Fig. 5).
On the inner surface of the reaction chamber of the chip, we print a pre-quenched DNA microarray (Fig. 1c) to allow highly multiplexed probe-based readout. Microarrays allow detection of up to hundreds of thousands of different nucleic acid targets using a single fluorescence channel24,25 by spatially separating different probes. However, traditional microarrays are unsuitable for in vitro diagnostic use because they require labour-intensive and open-tube wash steps to suppress fluorescence background. Early commercial attempts at combining real-time PCR with microarrays were based on a PCR amplification reaction followed by a separate hybridization reaction to a microarray26,27. However, these approaches have not been widely adopted and are no longer commercially available.
The team has developed a pre-quenched microarray chemistry, in which unlabelled amplicons induce an increase of the corresponding spot’s fluorescence via displacement of a quencher-functionalized oligonucleotide. Because the PCR amplicons are unlabelled, no wash steps are needed to reduce fluorescent background from excess amplicons, and a highly multiplexed readout for many different DNA targets can be achieved in a closed tube reaction without specialized opto-fluidic equipment.
Once the loaded chip is mounted against the heaters, the PCR reaction begins, and an external camera is used to periodically take pictures of the microarray area (Fig. 1d). At early time points, probe spots on the microarray are dark except for positive control spots (Fig. 1e); at the end of the reaction, the probes that are hybridized to DNA amplicons become bright. Importantly, toroidal PCR amplification occurs concurrently with probe hybridization, so the progress of the amplification reaction can be tracked in real time, unlike standard microarrays (Fig. 1f). This real-time readout allows robust and accurate DNA target quantification based on the time at which the fluorescence substantially increases. In contrast, endpoint fluorescence quantification for standard microarrays is known to be less stable due to sample contents, probe synthesis impurities, illumination non-uniformity, optical aberrance, physical smudges, and other factors.
Specificity, speed, sensitivity and quantification dynamic range
Previous implementations of convection PCR28 have not entered mainstream use, in part because they exhibited poor temperature uniformity that resulted in notable primer-dimer formation and non-specific genomic amplification. Thus, one of the first priorities in building a convection-based massively multiplex qPCR assay is to ensure PCR amplification specificity.
On the basis of our understanding, the fluid circulation in a reaction tube or chamber adopts laminar flow, with the flow velocity dependent on the temperature differential of the different circuit paths. In many reaction chamber designs, there will be regions containing fluid paths with minimal temperature differential that have minimal fluid movement (Supplementary Section 5). These regions may result in disproportionate formation of primer dimers and non-specific amplification. By engineering a donut-shaped reaction chamber in the PCR chip, we remove most of the dead volume, and are able to achieve similar PCR specificity on human gDNA as the commercial Bio-Rad CFX96 instrument (Fig. 2a). In contrast, for this pair of primers, a ‘pizza’-shaped reaction chamber and a capillary tube both result in primer-dimer and non-specific amplicon formation. Our fluid dynamics modelling on ANSYS supports our experimental findings that convection PCR in capillary tubes, ‘pizza’-shaped chambers and square toroidal chambers all result in a considerably higher proportion of slow movement zones (Supplementary Section 5).
a, Toroidal PCR achieves much higher PCR specificity than alternative implementations of convection PCR, enabling diagnostics-grade DNA analysis. Shown here is 2% agarose gel electrophoresis on single-plex PCR amplification products (human TFRC gene) using different systems. Lane 1 shows results of convection PCR performed in a commercial capillary tube and convection PCR instrument (GeneReach49). Hardly any intended amplicon molecules are generated, and the vast majority of amplification products are non-specific amplicons or primer dimers. Lane 2 shows the results on the toroidal PCR system using a non-annular cylindrical reaction chamber (‘pizza’-shaped chamber with the same external sizes as the donut-shaped chamber). There is notable non-specific amplification and primer-dimer formation due to the dead space in the middle with low temperature and circulation velocity. Lane 3 shows the results from the toroidal PCR system. Lane 4 shows the results from using a commercial qPCR instrument (Bio-Rad CFX96) in a 200 µl PCR tube. The experiment was performed once. b, Dependence of circulation velocity on chamber thickness. Faster flow circulation is achieved by increasing the thickness of the toroidal reaction chamber. The dashed line shows expected mean particle velocities based on ANSYS fluid dynamics modelling (Supplementary Section 5). The grey rectangle shows the conditions the current donut-shaped fluidic chamber is being operated at. The image on the right is taken with 1 s exposure time and used for quantification of circulation velocity in a 360-µm-thick chamber. The streaks show movement of fluorescent polystyrene beads (10 µm in diameter). The variation in bead velocities probably corresponds to the position in the z axis, with beads closer to surfaces expected to be slower. c, Pictures of the toroidal PCR chip (360-µm-thick chamber) loaded with red dye to visualize fluid circulation. d, Toroidal PCR sensitivity and dynamic range. As with qPCR, the observed toroidal PCR spot fluorescence follows a sigmoidal curve. Here, we show fluorescence traces for amplification of human gDNA from 35 pg (10 haploid genome copies) to 350 ng (105 copies). The time at which the fluorescence crosses a threshold value (Tt) is dependent on the log of the input DNA concentration, as expected. NTC, no template control. e, Simultaneous amplification and quantification of mixed samples of human and mouse gDNA. We designed primers and probes for the human TFRC gene and the mouse Nono gene, and quantitated each gene for samples wherein the human DNA was fixed at 1,000 copies and mouse DNA varied from 100 to 12,800 copies. The Tt values for human TFRC were unaffected by the changes in the mouse DNA concentration. See Supplementary Section 6 for additional data. f, Sample fluorescence images of experiments shown in e. The mouse probe (left spot) became brighter at earlier times when a larger amount of mouse DNA was used as input.
Next, we aimed to improve the amplification speed within the toroidal PCR chamber because rapid turnaround is highly desirable for point-of-care applications such as pathogen identification. The speed of fluid circulation in the toroidal PCR is impacted by the thickness of the reaction chamber, because in laminar flow, the fluid velocity near a surface approaches zero. Figure 2b shows that the mean circulation velocity increases in thicker chambers, consistent with expectations, and plateauing at roughly 360 µm chamber thickness. At this thickness, for our standard chamber dimensions (10 mm outer diameter, 4 mm inner diameter), the chamber volume is approximately 25 µl, consistent with commercial qPCR reaction volumes. With a 25 s circulation time, a standard 40-cycle PCR protocol can be completed within 20 min.
To evaluate the speed of the amplification and the dynamic range of the toroidal PCR assay, we next ran the toroid chip using different quantities of the NA18537 human cell line gDNA (Fig. 2d), ranging from 10 haploid genomic copies (5 cell equivalents, 30 pg) to 105 haploid genomic copies. Triplicate repeat experiments showed consistent transition time (Tt), defined as the time at which fluorescence first exceeds the threshold of 0.4 RFU. Furthermore, the Tt values for lower input DNA quantities increased as predicted in a log-linear fashion, similar to qPCR and demonstrating a dynamic range of at least four logs. Experimentally, the ∆Tt differed by 86 ± 14 s per tenfold reduction of DNA input, implying a doubling time of 26 ± 4 s. This experimental value is consistent with the calculated value of 26.2 s, based on the observed fluid velocity of 0.84 mm s−1 and mean lap circumference of 22 mm. All three negative control samples (water) did not have spot fluorescence exceeding the threshold, confirming specific PCR amplification and probe detection.
Next, we performed simultaneous quantification of mouse and human DNA on a toroidal PCR chip, to characterize the potential interference of quantification accuracy due to the presence of variable quantities of background DNA (Fig. 2e,f). We observe that as the stoichiometric ratio of mouse DNA to human DNA ranges from 12.8 to 0.1, the value of Tt for human DNA is essentially unaffected. Simultaneously, the log-linearity of the mouse DNA Tt value is also unaffected by the presence of human DNA. Collectively, these results suggest that the toroidal PCR assay could be used for multiplexed gene-expression profiling.
Spot position independence
The toroidal PCR assay leverages the embedded pre-quenched microarray to achieve massive multiplexing. For these multiplexing capabilities to be realized in a diagnostic setting, it is necessary that the observed results are reproducible and consistent across different spots. To characterize interspot consistency, we constructed a 48-spot toroidal PCR chip that includes quintuplet repeat spots for probes against 3 human genes, 3 rat genes and 3 mouse genes, plus 2 positive controls and 1 negative control (Fig. 3a). We observe that although there is a substantial difference in endpoint spot fluorescence intensity, suspected to be primarily due to non-uniform illumination, the values of Tt are well conserved across all five replicate spots for each of the nine genes.
a, Chip layout. Out of 48 spots, 3 were used as positive and negative controls (PC and NC, respectively), 15 spots were used for fivefold replicate detection of 3 human genes (H1, H2 and H3), 15 spots for 3 rat genes (R1, R2 and R3) and 15 spots for 3 mouse genes (M1, M2 and M3). b, Sample fluorescence image at t = 30 min when 4 ng of human gDNA was loaded as input, using PCR primers for all 9 genes (including 3 rat and 3 mouse). Only the human and positive control spots were bright, as expected. There is variation in the endpoint brightness of different probes due to both uneven illumination and to differences in fluorophore coupling and probe spotting efficiency. c, Time-course fluorescence traces for all 9 sets of probe spots. Threshold times are generally consistent across the fivefold replicate spots. d, Fluorescence image (top) and time-course fluorescence traces (bottom) using 4 ng rat DNA input, using the same primers and probes as in b and c. e, Fluorescence image (top) and time-course fluorescence traces (bottom) using 4 ng mouse DNA input.
Portable instrument
Experiments presented thus far have used a commercial fluorescence microscope as the readout instrument. To facilitate the adoption of the toroidal PCR assay for a variety of applications that require portability and rapid turnaround, we next designed and built a portable toroidal PCR instrument (Fig. 4a). Importantly, commercial qPCR instruments require wall power, preventing them from being rapidly deployed to point-of-care or field-use settings where rapid diagnostic testing may be needed. A major component of the power requirement is the need to rapidly cool PCR reaction mixtures from 95 °C (denaturing step) to 60 °C (annealing step). Because toroidal PCR achieves rapid thermal cycling using passive fluidic flow, it requires only constant-temperature heaters with low power consumption. Our breadboard prototype (Fig. 4b) thus is able to run off a mid-size 12 V battery (12 V × 18 Ah) for 9 or 10 experiments and does not require connection to an external power source.
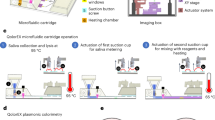
a, Simplified schematic of the loading, heater-clamping and imaging modules. b, Picture of initial breadboard prototype instrument, using an iPhone 6S as a portable camera. This prototype is fully battery powered (12 V) and does not require an external power supply. c, Probe layout for a toroidal PCR chip designed to detect bacterial species frequently observed in nosocomial (hospital acquired) infections. Bacterial species: (1) Enterococcus faecium, (2) Staphylococcus aureus, (3) Klebsiella pneumoniae, (4) Acinetobacter baumannii, (5) Pseudomonas aeruginosa, (6) Enterobacter cloacae, (7) Enterobacter agglomerans, (8) Enterobacter aerogenes, (9) Chlamydia pneumoniae, (10) Mycoplasma pneumoniae, (11) Streptococcus pneumoniae, (12) Bordetella pertussis, (13) Streptococcus agalactiae, (14) Streptococcus pyogenes, (15) Escherichia coli. d, Experimental results on the breadboard prototype using a 0.5 ng clinical isolate sample of C. pneumoniae (left) and a mixture of clinical isolate samples of 0.25 ng K. pneumoniae and 0.25 ng S. pneumoniae (right). e, Picture of functional prototype instrument. f, Experimental results on the functional prototype using a mixture of clinical isolate samples consisting of 1 ng P. aeruginosa, 1 ng E. claocae and 1 ng S. aureus. For historical reasons, input quantity and chip layout differed between c and f. In d and f, fluorescent traces coloured in grey represent the kinetics of signal accumulation from the rest of the microarray probes. See Supplementary Section 9 for probe layout.
There are five main modules in the instrument: (1) a mechanical mechanism to mount the chip against the heaters with sufficient force to ensure good thermal contact; (2) a closed-loop thermal system to ensure the chip is differentially heated to the desired temperatures; (3) an optics setup to illuminate the chip and filter out scattered light to reduce fluorescence background; (4) a camera to acquire fluorescence images; and (5) a microcontroller to coordinate timing of all components. In total, the cost of the components of this breadboard prototype was roughly US$2,500, with the bulk of the cost from the camera (an iPhone 6S, US$600), the optical filters (ThorLabs MF530-43 and Chroma AT575LP, US$500), the light-emitting diode (LED) light source (ThorLabs LED1B, US$325) and the LED driver (ThorLabs M530L4, US$296).
To validate the functionality of this prototype instrument, we constructed a 15-plex bacterial identification panel on the toroidal PCR chip. The DNA sequence encoding the 16S ribosomal RNA and 23S ribosomal RNA in bacteria are mostly conserved but contains nine hypervariable regions with sequences that differ across species but are conserved within strains of the same species. For this 15-plex bacterial panel, we constructed 15 different probes that target distinct 16S sequences (located in the V3 hypervariable region) that serve as signatures for 15 important bacterial species frequently implicated in nosocomial (hospital acquired) infections, including the most common ESKAPE bacteria29. Nosocomial infections cause roughly 20,000 deaths in the United States per year30, and rapid pathogen identification could help inform timely targeted antibiotic treatment that can improve patient outcomes, limit spread and drug resistance15,16.
Our ESKAPE panel performed as we expected on the breadboard prototype instrument, identifying both a single bacteria species and a combination of two bacteria species. Because the current chip and instrument design does not include sample preparation and DNA extraction modules, we performed validation using DNA input from clinically derived isolates and reference strains purchased commercially. We individually tested each DNA sample obtained (see Supplementary Section 9 for additional experimental data on the 15-plex bacterial panel using the breadboard prototype).
Because the breadboard prototype is open to the air, it is vulnerable to ambient dust that can occlude or distort the optics and ambient light that reduces image signal-to-noise ratio. We next built a closed instrument with similar functionality to our breadboard prototype (Fig. 4e). As expected, the signal-to-noise ratio of the fluorescent spots was improved, and the 15-plex bacterial identification panel was able to clearly identify all three bacterial species in a mixed sample of clinical isolates (Fig. 4f). See Supplementary Section 9 for revised chip layout for the production prototype instrument.
SNP genotyping
SNPs are natural variations in the human genome; over 100 million distinct SNP loci have been reported in the human genome31,32. Genome-wide association studies have discovered a wide range of SNPs that correlate with disease risk, type of diabetes33, neurodegenerative disease34,35, hereditary breast36 and colorectal cancers37, and coronary disease38. In addition to human disease-related applications, SNP genotyping can also be used for DNA forensics application39 as well as agricultural seed selection40.
Currently, microarrays are the preferred technology for SNP-genotyping applications, due to its massive multiplexing, high automation and acceptable economics (typically ≤US$60 per Affymetrix array chip). However, analysing a sample using a microarray takes approximately 24 h. Furthermore, because it requires three large specialized instruments for hybridization, washing and imaging, samples need to be transported to central labs for analysis, adding more days to the total turnaround time. While a three-to-seven-day turnaround is acceptable for non-time-sensitive applications, other SNP-genotyping applications require rapid turnaround. One prominent example is pharmacogenetics13; the use of SNP-genotyping information to inform dosage of drugs such as warfarin based on individualized drug metabolism rates41. Information to guide accurate dosage of warfarin to treat patients suffering from stroke or deep vein thrombosis needs to be provided immediately.
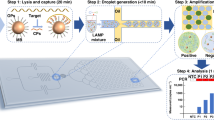
For SNP genotyping, we use the X-probe architecture23 to achieve probe-based single-nucleotide discrimination while limiting the reagent costs of chemically modified DNA (Fig. 5a, see also Supplementary Section 10). For each SNP locus, we design two separate surface-bound X-probes, one to each allele. Human DNA samples that are homozygous at the SNP locus will have only the corresponding allele spot light up, while samples that are heterozygous will have both spots light up (Fig. 5b). The overall workflow from buccal (cheek) swab sample collection takes less than 1 h, including less than 5 min of hands-on time (Fig. 5c). See http://demovideo.torus.bio for a two-minute video of the entire workflow.
a, To simultaneously achieve single-nucleotide discrimination and reduce functionalized oligo costs, we use the X-probe architecture23 for the SNP-genotyping panel here. Due to the thermodynamics and kinetics of the X-probe design, it will bind only perfectly matched DNA sequences, and even DNA sequences differing by a single SNP allele will not favourably bind to the probe. b, Sample experimental results on four SNP loci. Each pair of probes performs genotyping on a different SNP. c, Illustration of workflow for human buccal swab sample testing on the toroidal PCR. Total turnaround time is under 1 h. d, Genotyping 15 SNP loci from buccal swab samples from a family trio. See Supplementary Section 10 for SNP loci covered, probe layout, time-based fluorescence plots. Buccal swab and fingerstick samples were collected with informed consent.
To showcase the validation and accuracy of SNP genotyping by the toroidal PCR assay, we constructed a 30-spot array corresponding to the alternate alleles for 15 different human SNP loci. Initial panel testing showed correct SNP genotype calls for all 15 loci for 3 different human cell line gDNA samples. Next, this panel was applied to buccal swab samples from a family trio of mother, father and child under informed consent, using the production prototype instrument shown in Fig. 4e. Endpoint fluorescence images of the samples are shown in Fig. 5d, but as usual, SNP genotype calls are made using time-based fluorescence traces.
Reproducibility studies
To verify the robustness of the toroidal PCR system and the reproducibility of highly multiplexed panels, we performed replicate experiments a 19-plex toroidal PCR panel to different regions of the human genome (Fig. 6). For each of 3 input DNA concentrations (2 ng, 20 ng and 200 ng), we performed 20 replicate chips, for a total of 60 chips. Although there is some variability in the threshold time Tt, the results are quantitatively consistent with observed standard deviations on Tt having a mean value of less than 0.3 min across 19 loci × 3 concentrations. As with qPCR, different primers have different per-step PCR yields, so standard optimization of primer sequences to achieve more uniform Ct (quantitation cycle threshold) values on qPCR would probably lead to more uniform Tt values on toroidal PCR as well. Furthermore, the relative change in Tt value for tenfold reductions of in input DNA quantity follows a linear trend as expected (Fig. 6b), supporting the ability to do multiplex quantification using toroidal PCR.
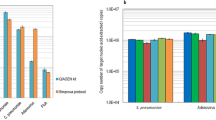
a, Summary of observed threshold time Tt for a 19-plex toroidal PCR panel. Here, we tried 3 different input DNA quantities of the NA18537 human gDNA, ranging from 2 ng to 200 ng. For each quantity, 20 replicate toroidal PCR chips were run, for a total of 60 chips and 1,140 observed Tt values plotted. b, Comparison of Tt values for 20 ng (top) and 2 ng (bottom) input DNA versus 200 ng input DNA. Each point corresponds to the mean Tt values for one of the 19 loci detected. The horizontal and vertical lines denote ±1 s.d. based on the 20 replicate experiments. The solid black line shows the best linear fit, and the dashed black line shows y = x. c, Comparison of toroidal PCR and commercial qPCR instrument (Bio-Rad CFX96). The same primers used for the 19-plex toroidal PCR were split across seven qPCR reactions ranging between 2-plex and 3-plex, using Taqman probes with spectrally distinct fluorophores (FAM, HEX and Cy5). Because the same 19-plex assay was split across seven tubes, sevenfold more input DNA was needed for the Taqman qPCR experiments (140 ng total). d, Comparison of the turnaround time of 19-plex toroidal PCR and 2-/3-plex qPCR. The plotted toroidal PCR Tt values (green) are the mean values from 20 replicate experiments from a. The plotted qPCR Ct values (brown) are the mean values of three replicate experiments. The time to Ct for qPCR reactions is reported based on actual time to Ct, including initial enzyme hotstart activation, temperature ramping, and standard qPCR thermal cycling denaturing and annealing step times. See Supplementary Section 12 for qPCR traces and detailed qPCR protocol. The time to a positive result for toroidal PCR is roughly 1/3 that of the qPCR assays. The total turnaround time for reporting a negative result would be roughly 50% higher for both toroidal PCR and qPCR.
We next compared the performance of the 19-plex toroidal PCR panel to that of qPCR assays on a commercial qPCR instrument. Most commercial qPCR instruments, including the Bio-Rad CFX96 instrument that we used, have five fluorescence colour channels. However, one of these channels (ROX) is typically used as an internal control, and not recommended for use in multiplex qPCR. Another one of the channels is far in the red spectrum, and requires less frequently used fluorophores such as Quasar 705 or Cy5.5. Consequently, multiplex qPCR assays are typically limited to 3-plex (FAM, HEX, Cy5) or 4-plex (FAM, HEX, Cy5, Cy5.5).
We divided our 19-plex toroidal PCR panel into seven tubes of multiplex qPCR assays (five tubes of 3-plex and two tubes of 2-plex, Fig. 6c). To ensure consistency, we used the same primer sequences for the qPCR reactions as for toroidal PCR, but only the relevant primers for each qPCR tube was introduced, consistent with standard qPCR assay design. For each of the two or three amplicons in each qPCR reaction, we used a corresponding number of Taqman probes to independently report detection of each amplicon. Using 20 ng of NA18537 human gDNA in each of the seven multiplex qPCR reactions, we observed Ct values of roughly 25, consistent with expectations for Taqman-based qPCR assays. It takes the toroidal PCR system 15 min to reach Ct = 25, which is threefold faster than traditional qPCR instrument (45 min). On this 19-plex assay, the toroidal PCR required only 1/7 of the input DNA needed for traditional multiplex qPCR. This is an important advantage in diagnostic settings with limited sample availability such as prenatal genetic testing or forensic analysis. On higher-plex panels, toroidal PCR will still hold the advantage of threefold shorter turnaround time, while the low-input-DNA requirement advantage becomes a larger differentiating factor.
Discussion
The toroidal PCR system presented here achieves rapid, sensitive and quantitative detection of many DNA targets from a single sample using a closed, portable and affordable instrument. The lowest input DNA that we showed is 35 pg of human gDNA, corresponding to 10 haploid copies. Below 10 copies, stochastic Poisson sample distribution render detection less reliable; this is well documented for qPCR assays. For example, a sample with nominally 2 molecules of a DNA target has a 14% chance of containing 0 molecules instead.
Although we have limited our studies in this work to arrays of 16 to 48 spots, probe printing is a highly scalable process and we have demonstrated that over 1,000 probes can be printed on a single toroidal PCR chip (Supplementary Section 11). We believe that the toroidal PCR system will be competitive in a range of applications where rapid and highly multiplexed DNA detection is needed in decentralized settings.
With the recent coronavirus pandemic, one apt use case for the toroidal PCR may be in disease screening, for example, at airports to triage patients with similar clinical symptoms, for example, influenza, parainfluenza, human syncytial virus, common cold and severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Furthermore, the high mutability of viruses results in a strong need for molecular surveillance, to identify novel mutations that can result in substrains with different infectivity and disease severity42,43. Single-plex isothermal amplification assays7,8 are vulnerable to low and decreasing clinical sensitivity as virus genomes evolve away from the target sequences single-plex assays are designed to detect. The high multiplexing of the toroidal PCR system can overcome potential clinical false negatives by redundantly detecting many conserved pathogen-specific RNA or complementary DNA sequences.
The single-nucleotide specificity demonstrated by the toroidal PCR system in Fig. 5 allows simultaneous detection of antibiotic resistance with pathogen identification. Antibiotic resistances are typically caused either by gain of a gene (for example, mecA for methicillin resistance) or mutation of a gene (for example, gyrA for fluoroquinone resistance44). While other multiplex PCR systems such as BioFire9 or StatDx45 can detect drug resistance due to gain of gene, they typically lack the molecular specificity needed to identify resistance caused by point mutations. The ability to rapidly identify infections and prescribe effective antimicrobial therapies is especially needed to reduce the mortality and morbidity rate of nosocomial infections in the United States.
The core innovations in the toroidal PCR system are the donut-shaped chamber to allow reliable and low-power convection-based qPCR, and the integrated pre-quenched microarray to allow massively multiplexed readout through spatial separation. As a result, our experimental demonstrations in this paper were ‘DNA in, answer out’ workflows. We did not consider sample preparation and DNA extraction modules. Extensive previous work has reported a variety of different lab-on-a-chip approaches to process blood, nasal/buccal swab, urine and cerebrospinal fluid. Integration with a pre-analytical module will probably be necessary for adoption of the toroidal PCR system in point-of-care settings for infectious disease diagnostic applications.
Another potential application area for toroidal PCR is multigene-expression analysis. Cancer prognosis tests such as Oncotype Dx46,47 use tumour gene-expression profiles to stratify patients for more aggressive treatment such as chemotherapy. Recently, research on host biomarkers48 also suggests that human gene expression can be used to differentiate exposure to bacteria versus viruses and could serve as a complementary set of markers to pathogen DNA. We believe that the ability of toroidal PCR to rapidly and simultaneously detect and quantify many different nucleic acid markers positions it well as a system for performing complex DNA and RNA diagnostics in settings convenient to the patient.
Methods
Surface functionalization
Attachment of the pre-annealed probe to the surface of the fluidic chamber is performed via strain-promoted azide–alkyne cycloaddition between the dibenzocyclooctyne-functionalized probe and the azide (N3)-functionalized surface of the fluidic chamber. N3 surface functionalization is performed according to the following protocol depicted in Supplementary Fig. 1a. Step 1: glass slides are treated with sodium hydroxide solution (10 M) for 30 min in Branson ultrasonic bath (Bransonic CPXH Digital Bath 2800) at the high power of ultrasound generation. Step 2: after washing with copious amount of water and drying at 120 °C, the slides are treated with oxygen plasma for 30 min in Harrick plasma cleaner at high power and maintaining oxygen pressure in range of 0.2–0.3 torr. Step 3: hydroxylated slides are treated with a 5% solution of N-(2-aminoethyl)-2,2,4-trimethyl-1-azasilacyclopentane (Gelest) in dry dichloromethane (Sigma) for 16 h at room temperature, followed by washing with 95% ethanol in water solution; amino-salinized slides are baked in ambient atmosphere at 105 °C for 2 h. Step 4: functionalization of the amino-terminated slides with an isothiocyanate moiety is performed with 10 mM solution of p-phenylene diisothiocyanate (Sigma) in dry pyridine-dimethylformamide (both chemicals are from Sigma) mixture (9:1, v/v) for 2 h at room temperature, followed by washing with 95% ethanol in water solution and drying at 80 °C for 1 h. Step 5: azido-functionalization of the isothiocyanate slides is performed at 37 °C for 2 h with a solution of 11-azido-3,6,9-trioxaundecan-1-amine (50 µl) in 0.1 M phosphate buffer pH 8 (10 ml), followed by washing with copious amount of water and drying at 45 °C. At this stage, the slides are ready for microarray printing or can be stored vacuum-sealed at 4 °C for at least 6 months without loss of conjugation activity. The scheme of the surface functionalization protocol can be found in Supplementary Fig. 1.
Probe annealing
Toehold microarray probes were annealed in 2× PBS buffer according to the following protocol: 90 °C for 3 min, then uniformly decreasing temperature down to 30 °C over a period of 1 h. Concentrations of the probe components for the universal probe architecture (Supplementary Fig. 2a) were as follows: anchor strand −2 µM, arm strand −3 µM, quencher strand −4.5 µM. Concentrations of the probe components for the super universal probe architecture (Supplementary Fig. 2b) were as follows: anchor strand −2 µM, arm strand −3 µM, quencher arm strand −4.5 µM, universal quencher strand −8 µM. Schematics of the DNA probe architecture can be found in Supplementary Fig. 2.
General considerations on thermodynamics of toehold and X-probes
During the probe design, we kept the same toehold binding energy within a range −9.5 to −10.5 kcal mol−1 (for all probes designs) for a fully complementary sequence. By comparison, toehold binding with variant amplicon sequence would have higher energy, and thus not favourable for toehold displacement reaction initiation, which in turn would result in no or weak fluorescence signal increase. The binding energetics for the specific arm sequence (binding between the arm strand and the quencher strand or quencher arm strand) was kept in the range of −22 to −24 kcal mol−1 to prevent spontaneous dissociation of the quencher strand from the probe. Both anchor strand and universal quencher strand had binding energy of around −24 kcal mol−1. Probes designed for SNP discrimination were also equipped with a unique non-homologous double-stranded region with binding energy of around −5 kcal mol−1. Presented probe energetics was calculated for 60 °C and concentration of Mg2+ cations at 6 mM, using the script written in our laboratory.
Microarray spotting solution preparation
Microarray spotting solution was prepared by diluting the annealed probes with Tris-EDTA buffer spiked with MgCl2 to obtain the following final concentration of the components: 1× Tris-EDTA buffer (10 mM Tris, 1 mM EDTA), 10 mM MgCl2, 0.5 µM of the probe, recalculated to the concentration of the anchor strand.
Microarray printing
Microarrays were printed using Scienion S3 microarray printer equipped with PDC070 micronozzles. Printing was preformed at 70% relative humidity maintained in the printing enclosure. After the printing run finished, the arrays were allowed to rest for 15 min at the same humidity and were then dried in a desiccator under reduced pressure for 1 h. The microarrays were stored at room temperature in the vacuum-sealed polyethylene bags until final assembly and use.
DNA oligonucleotides
All DNA oligonucleotides used in the study were purchased from Integrated DNA Technology, except for the strands labelled with the BHQ-2 moiety, which were purchased from Biosearch Technologies. All PCR primers were ordered without purification, but desalted. All fluorophore-labelled oligonucleotides, quencher-labelled oligonucleotides and unlabelled oligonucleotides shorter than 50 nt were ordered with reverse-phase HPLC purification. All unlabelled oligonucleotides longer than 50 nt were ordered with PAGE purification.
Design of primers
Main considerations during primer design were as follows: (1) amplicon length was restricted to 300 bp or shorter; (2) standard free energy of primer binding (ΔG°) was in the range of −11 to −12 kcal mol−1 at 60 °C in a solution with 6 mM Mg2+ concentration; (3) no predicted intermolecular binding with standard free energy more negative than −3 kcal mol−1; (4) no intramolecular secondary structures with standard free energy more negative than −5 kcal mol−1.
Composition of PCR reaction
Toroidal PCR was performed using 15 units of AptaTaq Δ exo DNA Polymerase (Roche) per reaction, 1× AptaTaq Fast PCR buffer and final MgCl2 concentration of 6 mM. The increased concentration of magnesium ions was reasoned by their non-specific absorption on glass surface. The concentration of each of the standard deoxynucleotide triphosphates (dATP, dTTP, dGTP, dCTP) was 0.25 mM. Each PCR reaction mixture was supplied with 5% DMSO and BSA at a concentration of 0.3 µg µl−1. Concentrations of primers varied from 50 to 400 nM depending on the assay run. The toroidal PCR instrument heaters were set to 60 °C (array side) and 95 °C.
Standard (tube) PCR was performed using AptaTaq Δ exo DNA Polymerase (Roche) with 1× AptaTaq Fast PCR buffer at a final MgCl2 concentration of 1.5 mM. The concentration of each of the standard deoxynucleotide triphosphates (dATP, dTTP, dGTP, dCTP) was 0.2 mM. Concentration of primers was 200 nM. PCR amplification regime of 95 °C for 1 min, 30 cycles of 95 °C for 10 s, 60 °C for 10 s, 72 °C for 15 s, and a final elongation of 72 °C for 30 s was used.
Reference DNA samples
Reference human gDNA samples were purchased from Coriell Institute for Medical Research. Reference bacterial gDNA samples were purchased from ATCC collection and Zeptometrix.
DNA extraction
DNA extraction from fresh fingerstick blood samples (200 µl) was performed using Qiagen blood mini kit strictly following manufacturer’s protocol. DNA extraction from buccal swab samples was performed using QuickExtract DNA Extraction Solution from Lucigen (recently acquired by Biosearch Technologies). In brief: after collecting buccal swab samples (SK-2S buccal swab, Isohelix), the swabs were placed and vortexed in 0.5 ml of QuickExtract DNA Extraction Solution. The solution (with the swabs) was heated at 60 °C for 10–15 min, at 95 °C for 3 min, and then cooled to room temperature. Two microlitres of the solution was used as DNA input for SNP-genotyping assays.
Agarose gel electrophoresis
Agarose electrophoresis was performed in 1× TEA buffer (40 mM Tris-HCl, 20 mM acetic acid and 1 mM EDTA, pH 8.6) at the electric field strength of about 10 V cm−1.
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The main data supporting the results in this study are available within the paper and its Supplementary Information. The raw and analysed datasets generated during the study are too large to be publicly shared, yet they are available for research purposes from the corresponding author on reasonable request. Some exemplary raw data are available from figshare with the identifier https://doi.org/10.6084/m9.figshare.14612343.
Code availability
The custom MATLAB code used to analyse fluorescence images is available at https://github.com/KhodakovDA/Toroidal-PCR. MATLAB version R2014b was used.
References
Goodwin, S., McPherson, J. D. & McCombie, W. R. Coming of age: ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351 (2016).
Mardis, E. R. Next-generation sequencing platforms. Annu. Rev. Anal. Chem. 6, 287–303 (2013).
Heller, M. J. DNA microarray technology: devices, systems, and applications. Annu. Rev. Biomed. Eng. 4, 129–153 (2002).
Loven, J. et al. Revisiting global gene expression analysis. Cell 151, 476–482 (2012).
Higuchi, R., Fockler, C., Dollinger, G. & Watson, R. Kinetic PCR analysis: real-time monitoring of DNA amplification reactions. Biotechnology 11, 1026–1030 (1993).
Taylor, S., Wakem, M., Dijkman, G., Alsarraj, M. & Nguyen, M. A practical approach to RT–qPCR publishing data that conform to the MIQE guidelines. Methods 50, S1–S5 (2010).
Gill, P. & Ghaemi, A. Nucleic acid isothermal amplification technologies, a review. Nucleosides Nucleotides Nucleic Acids 27, 224–243 (2008).
Craw, P. & Balachandran, W. Isothermal nucleic acid amplification technologies for point-of-care diagnostics: a critical review. Lab Chip 12, 2469–2486 (2012).
Buss, S. N. et al. Multicenter evaluation of the BioFire FilmArray gastrointestinal panel for etiologic diagnosis of infectious gastroenteritis. J. Clin. Microbiol. 53, 915–925 (2015).
Jain, M., Olsen, H. E., Paten, B. & Akeson, M. The Oxford Nanopore MinION: delivery of nanopore sequencing to the genomics community. Genome Biol. 17, 239 (2016).
McCarthy, M. I. et al. Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat. Rev. Genet. 9, 356–369 (2008).
MacArthur, J. et al. The new NHGRI-EBI catalog of published genome-wide association studies (GWAS catalog). Nucleic Acids Res. 45, D896–D901 (2017).
Hauser, A. S. et al. Pharmacogenomics of GPCR drug targets. Cell 172, 41–54 (2018).
Weitzel, J. N., Blazer, K. R., MacDonald, D. J., Culver, J. O. & Offit, K. Genetics, genomics, and cancer risk assessment: state of the art and future directions in the era of personalized medicine. CA Cancer J. Clin. 61, 327–359 (2011).
Koser, C. U., Ellington, M. J. & Peacock, S. J. Whole-genome sequencing to control antimicrobial resistance. Trends Genet. 30, 401–407 (2014).
Lowy, F. D. Antimicrobial resistance: the example of Staphylococcus aureus. J. Clin. Investig. 111, 1265–1273 (2003).
Krishnan, M., Ugaz, V. M. & Burns, M. A. PCR in a Rayleigh–Benard convection cell. Science 298, 793–793 (2002).
Agrawal, N. & Ugaz, V. M. A buoyancy-driven compact thermocycler for rapid PCR. Clin. Lab. Med. 27, 215–223 (2007).
Agrawal, N., Hassan, Y. A. & Ugaz, V. M. A pocket-sized convective PCR thermocycler. Angew. Chem. Int. Ed. 46, 4316–4319 (2007).
Braun, D., Goddard, N. L. & Libchaber, A. Exponential DNA replication by laminar convection. Phys. Rev. Lett. 91, 158103 (2003).
Hsieh, Y. F. et al. A real-time convective PCR machine in a capillary tube instrumented with a CCD-based fluorometer. Sens. Actuators B 183, 434–440 (2013).
Zhang, D. Y., Chen, S. X. & Yin, P. Optimizing the specificity of nucleic acid hybridization. Nat. Chem. 4, 208–214 (2012).
Wang, J. S. & Zhang, D. Y. Simulation-guided DNA probe design for consistently ultraspecific hybridization. Nat. Chem. 7, 545–553 (2015).
Gresham, D., Dunham, M. J. & Botstein, D. Comparing whole genomes using DNA microarrays. Nat. Rev. Genet. 9, 291–302 (2008).
Do, J. H. & Choi, D. K. Normalization of microarray data: single-labeled and dual-labeled arrays. Mol. Cells 22, 254–261 (2006).
Bogaerts, P. et al. Analytical validation of a novel high multiplexing real-time PCR array for the identification of key pathogens causative of bacterial ventilator-associated pneumonia and their associated resistance genes. J. Antimicrob. Chemother. 68, 340–347 (2013).
Dijksman, J. F. & Pierik, A. Mathematical analysis of the real time array PCR (RTA PCR) process. Chem. Eng. Sci. 71, 496–506 (2012).
Miaoa, G. et al. Free convective PCR: from principle study to commercial applications—a critical review. Anal. Chim. Acta 1108, 177e197 (2000).
Pendleton, J. N., Gorman, S. P. & Gilmore, B. F. Clinical relevance of the ESKAPE pathogens. Expert Rev. Anti Infect. Ther. 11, 297–308 (2013).
Haque, M., Sartelli, M., McKimm, J. & Bakar, M. A. Health care-associated infection—an overview. Infect. Drug Resist. 11, 2321–2333 (2018).
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
Glusman, G., Caballero, J., Mauldin, D. E., Hood, L. & Roach, J. C. Kaviar: an accessible system for testing SNV novelty. Bioinformatics 27, 3216–3217 (2011).
Fuchsberger, C. et al. The genetic architecture of type 2 diabetes. Nature 536, 41–47 (2016).
Jansen, I. E. et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 51, 404–413 (2019).
Chang, D. et al. A meta-analysis of genome-wide association studies identifies 17 new Parkinson’s disease risk loci. Nat. Genet. 49, 1511–1516 (2017).
Robson, M. et al. Olaparib for metastatic breast cancer in patients with a germline BRCA mutation. N. Engl. J. Med. 377, 523–533 (2017).
Lynch, H. T., Snyder, C. L., Shaw, T. G., Heinen, C. D. & Hitchins, M. P. Milestones of Lynch syndrome: 1898–2015. Nat. Rev. Cancer 15, 181–194 (2015).
Nikpay, M. et al. A comprehensive 1000 Genomes-based genome-wide association meta-analysis of coronary artery disease. Nat. Genet. 47, 1121–1130 (2015).
Eduardoff, M. et al. Inter-laboratory evaluation of SNP-based forensic identification by massively parallel sequencing using the Ion PGM. Forensic Sci. Int. Genet. 17, 110–121 (2015).
Elshire, R. J. et al. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6, e19379 (2011).
Mega, J. L. et al. Genetics and the clinical response to warfarin and edoxaban: findings from the randomised, double-blind ENGAGE AF-TIMI 48 trial. Lancet 385, 2280–2287 (2015).
Hadfield, J. et al. Nextstrain: real-time tracking of pathogen evolution. Bioinformatics 34, 4121–4123 (2018).
Mousavizadeh, L. & Ghasemi, S. Genotype and phenotype of COVID-19: their roles in pathogenesis. J. Microbiol. Immunol. Infect. 54, 159–163 (2020).
Devasia, R. et al. High proportion of fluoroquinolone-resistant Mycobacterium tuberculosis isolates with novel gyrase polymorphisms and a gyrA region associated with fluoroquinolone susceptibility. J. Clin. Microbiol. 50, 1390–1396 (2012).
Parina, M. et al. Multicenter evaluation of the QIAstat Respiratory Panel—a new rapid highly multiplexed PCR based assay for diagnosis of acute respiratory tract infections. PLoS ONE 15, e0230183 (2020).
Dowsett, M. et al. Comparison of PAM50 risk of recurrence score with oncotype DX and IHC4 for predicting risk of distant recurrence after endocrine therapy. J. Clin. Oncol. 31, 2783–2790 (2013).
Kapasi, A. J., Dittrich, S., Gonzalez, I. J. & Rodwell, T. C. Host biomarkers for distinguishing bacterial from non-bacterial causes of acute febrile illness: a comprehensive review. PLoS ONE 11, e0160278 (2016).
Correia, C. N. et al. Circulating microRNAs as potential biomarkers of infectious disease. Front. Immunol. 8, 118 (2017).
Chou, W. P. et al. Rapid DNA amplification in a capillary tube by natural convection with a single isothermal heater. BioTechniques 50, 1 (2011).
Author information
Authors and Affiliations
Contributions
D.K. and D.Y.Z. conceived the project. D.K. and D.Y.Z. designed the primers and probes. D.K. designed and constructed the chip. D.K., J.L. and J.X.Z. designed and constructed the instrument. D.K. and D.Y.Z. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
There are patents issued on toehold probes (WO2012058488A1) and X-probes (WO2015094429A1) used in this work. There are patents pending on the toroidal PCR chip (WO2017172760A1) and toroidal PCR instrument (WO2020056292A1) presented in this work. D.K. declares a competing interest in the form of employment at and equity ownership in Torus Biosystems. J.L. declares a competing interest in the form of consulting for Torus Biosystems. J.X.Z. declares a competing interest in the form of employment at Nuprobe. D.Y.Z. declares a competing interest in the form of consulting for and equity ownership in Nuprobe, Torus Biosystems and Pana Bio.
Additional information
Peer review information Nature Biomedical Engineering thanks Xianbo Qiu, Tim Stakenborg and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary methods, figures, tables, references and video captions.
Supplementary Video 1
Real-time label-free fluorescent-signal accumulation registered from a microarray in a research-grade toroidal PCR.
Supplementary Video 2
Real-time label-free fluorescent-signal accumulation registered from a microarray in the functional toroidal PCR.
Supplementary Video 3
Real-time label-free fluorescent-signal accumulation registered from a microarray in the battery-powered breadboard toroidal PCR.
Supplementary Table 1
Sequences of primers and probes.
Rights and permissions
About this article
Cite this article
Khodakov, D., Li, J., Zhang, J.X. et al. Highly multiplexed rapid DNA detection with single-nucleotide specificity via convective PCR in a portable device. Nat Biomed Eng 5, 702–712 (2021). https://doi.org/10.1038/s41551-021-00755-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-021-00755-4
This article is cited by
-
Precise in-field molecular diagnostics of crop diseases by smartphone-based mutation-resolved pathogenic RNA analysis
Nature Communications (2023)
-
Analytical device miniaturization for the detection of circulating biomarkers
Nature Reviews Bioengineering (2023)
-
Rapid, tunable, and multiplexed detection of RNA using convective array PCR
Communications Biology (2023)
-
Multiplexed RNA profiling by regenerative catalysis enables blood-based subtyping of brain tumors
Nature Communications (2023)
-
Application of 4D two-colour LIF to explore the temperature field of laterally confined turbulent Rayleigh–Bénard convection
Experiments in Fluids (2023)