Abstract
The coronavirus disease 2019 (COVID-19) pandemic has highlighted the need for rapid and sensitive protein detection and quantification in simple and robust formats for widespread point-of-care applications. Here, we report on nanobody-functionalized organic electrochemical transistors with a modular architecture for the rapid quantification of single-molecule-to-nanomolar levels of specific antigens in complex bodily fluids. The sensors combine a solution-processable conjugated polymer in the transistor channel and high-density and orientation-controlled bioconjugation of nanobody–SpyCatcher fusion proteins on disposable gate electrodes. The devices provide results after 10 min of exposure to 5 μl of unprocessed samples, maintain high specificity and single-molecule sensitivity in human saliva and serum, and can be reprogrammed to detect any protein antigen if a corresponding specific nanobody is available. We used the sensors to detect green fluorescent protein, and severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) and Middle East respiratory syndrome coronavirus (MERS-CoV) spike proteins, and for the COVID-19 screening of unprocessed clinical nasopharyngeal swab and saliva samples with a wide range of viral loads.
Similar content being viewed by others
Main
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) had caused 2.5 million fatalities as of February 2021 (https://covid19.who.int) and may remain a worldwide health and economic burden for several years1. Owing to airborne transmission2 and a large share of symptom-free but infective carriers3,4, this virus defies traditional suppression measures that are based on the isolation of symptomatic individuals. Molecular testing has, therefore, become an essential tool in global attempts to curb the coronavirus disease 2019 (COVID-19) pandemic5. Testing currently relies on PCR with reverse transcription (RT-PCR)—a well-established diagnostic method with close to single-molecule sensitivity6. However, RT-PCR requires complex sample processing in specialized, well-instrumented laboratories. Owing to the limited capacity of these laboratories and the required logistics, results are typically communicated one to several days after sample collection. This delay in isolating infected carriers is severely hampering the suppression of infection chains7.
In a point-of-care (POC) setting, lateral flow immunoassays could offer an alternative as they detect viral protein (rather than RNA) directly in unprocessed patient samples and give qualitative results within 15–30 min. Yet, fast immunoassays detect viral loads above 50 million copies per ml, which is useful for the isolation of highly infectious individuals but not sensitive enough for diagnostic purposes8. In contrast, enzymatic nucleic acid detection-based POC assays generally have to trade off sensitivity and robustness against sample processing and reaction times9. Thus, the COVID-19 crisis highlights a critical limitation of our current POC diagnostics toolbox. What has remained elusive so far is rapid protein detection at the single-molecule level in a format that is simple and robust enough for widespread POC or bedside application. Here, we present a bioelectronic sensor technology that fulfils these requirements.
Recent years have seen major progress in the development of transistors that transduce and, importantly, amplify biological reactions or binding events into an electrical readout10,11,12,13,14. Organic electrochemical transistors (OECTs) have emerged as alternative bioelectronic transducers that outperform all other electrolyte-gated transistors and perform favorably compared with most solid-state technologies, which themselves have record high transconductance values15. OECTs operate in aqueous media and integrate ion-permeable conjugated polymers in their channel, which are volumetrically (de)doped by electrolyte ions injected by a gate electrode. The volumetric coupling between ionic and electronic charges in the channel makes OECTs powerful voltage amplifiers. As such, even a few binding events at the gate electrode can cause large modulations in the channel current16,17,18. This efficient on-site amplification of input signal allows for miniaturization and low-noise recordings, which should make OECTs particularly suitable for POC applications17. Despite these advantages, there are only a few reports of label-free OECT immunosensors19,20. More generally, current electronic protein sensors appear to translate poorly into real-world applications. Common drawbacks include complex sensor designs, the need for tightly controlled environments, reliance on regular off-the-shelf antibodies and their chemical immobilization in random orientations21.
We overcame these limitations in protein detection with a bioelectronic approach that advances OECT technology on several levels, ranging from chemical materials to biological recognition (Fig. 1). Key features in this process were: (1) the use of a novel organic semiconductor that allowed for a practical implementation of an accumulation-mode OECT; and (2) the controlled biofunctionalization of the sensor surface with (3) custom-engineered nanobodies22. The accumulation-mode OECT combined high output signal strength and stability with very low input power requirement. The bioassembly strategy coupled recombinant proteins in a precisely defined orientation at very high physical density.
We demonstrate that the resulting sensor is fast (<15 min from sample to result) and works with unprocessed saliva or serum samples. The sensor is largely reusable, easy to manufacture and highly modular. It reliably and specifically detects single protein molecules on millimeter-sized disposable electrodes in uncontrolled, ambient environments. The sensor is also quantitative and distinguishes concentrations over eight to ten orders of magnitude (attomolar to nanomolar). Experience with three different target proteins shows that our sensor technology is broadly applicable and only limited by the availability of antigen-specific nanobodies. We validated the performance with clinical unprocessed nasopharyngeal swab and saliva samples from patients with COVID-19 and demonstrate a sensitivity comparable to RT-PCR methods.
Results and discussion
Biofunctionalization
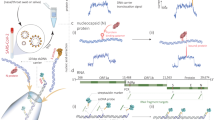
Nanobodies22 are compact recognition modules made from the antigen-binding domain (VHH) of an unusual class of heavy-chain-only antibodies found in Camelids23. Although several therapeutic24 and diagnostic applications25 have been developed, nanobodies have not yet been combined with OECT technology. One report described the functionalization of a conventional field effect transistor (FET) with nanobodies. This sensor relied on random chemical immobilization of nanobodies on carbon nanotubes and reached a detection limit of ~1 pM for green fluorescent protein (GFP)26. To improve on these previous studies, we capitalized on the fact that nanobodies (and modified versions thereof) can be efficiently expressed in Escherichia coli. We designed a recombinant protein in which the well-characterized anti-GFP nanobody27 is fused, through a flexible peptide linker, to a SpyCatcher domain (Fig. 2a). This SpyCatcher domain is specifically recognized by a short SpyTag peptide, triggering the autocatalytic formation of a covalent isopeptide bond linking both of them with very high stability. Originally engineered from a bacterial adhesion protein28, the SpyTag/SpyCatcher protein conjugation system has been used for several applications29 but has not yet been applied to FET or OECT biosensors. The only related application of a nanobody–SpyTag fusion was a passive, impedance-based sensor for the detection of microalgae30, which relied on the random chemical immobilization of separated SpyCatcher proteins as a capture reagent.
a, Operation. The gate electrode is exposed to a mix of sample (such as saliva) and binding buffer (1), rinsed with PBS (2) and mounted on top of the OECT channel for signal acquisition (3). b, Gate functionalization layers. Chem-SAMs and bio-SAMs are self-assembled on the gate electrode surface. c, Molecular architecture. A synthetic SpyTag peptide is chemically coupled to the HDT monolayer to form a chem-SAM. The nanobody–SpyCatcher fusion protein then attaches itself to this chemical layer through the autocatalytic formation of a covalent SpyCatcher–SpyTag bond, forming the bio-SAM. The nanobody domain defines sensor specificity and is interchangeable.
a, Annotated sequence of the recombinant fusion protein combining a nanobody recognition module (blue) with the SpyCatcher domain (brown), with an intervening flexible linker (dashed line). b, Structural model of the complete biorecognition layer assembled from HDT and SpyTag (chem-SAM) and nanobody–SpyCatcher fusion protein (bio-SAM) (based on PDB structures 4MLI and 3OBO). Unstructured, flexible linker regions are indicated by dashed lines drawn to scale. c, High-resolution S 2p XPS spectra of the gold electrode recorded after immobilization of the HDT layer, SpyTag peptide and nanobody–SpyCatcher protein (BSA added to the biofunctionalization solution). Spectra for the three functionalization steps are shifted vertically for clarity. d, QCM-D profile for the coupling of the SpyTag peptide and nanobody–SpyCatcher fusion protein (without BSA) in real time, starting from an HDT-coated gold electrode.
For our sensor, we wanted to avoid steps where the proteins (nanobody or SpyCatcher) are chemically modified or immobilized in a way that could partially impair their function. Instead, we opted for the immobilization of a chemically modified SpyTag, obtained through regular commercial peptide synthesis, on a 1,6-hexanedithiol (HDT) self-assembled monolayer (SAM), thus forming a combined chemical SAM (chem-SAM) on top of the gold gate electrode. The anti-GFP nanobody27–SpyCatcher fusion protein was genetically encoded, gene synthesized and produced in E. coli. The purified protein construct was then incubated under physiological conditions with the chem-SAM, thus completing the sensing surface with a self-assembled biological SAM (bio-SAM). This strategy leads to a defined biofunctionalization layer in a precisely controlled molecular configuration and orientation, which can be modeled at atomic resolution (Fig. 2b). A flexible eight-amino-acid glycine–serine linker separates nanobody and SpyCatcher domains dynamically and sterically. Taking into account unstructured residues contributed by the nanobody and SpyCatcher domains, the 4-nm-long nanobody domain is probably separated from the SpyCatcher adapter by about 30 amino acids, forming a ~12-nm-long flexible molecular leash (Fig. 2a,b). Additional flexibility is introduced by short glycine–serine and alkane linkers that separate the chemically synthesized SpyTag sequence from the underlying HDT, thus also giving the SpyCatcher domain some freedom to move and rearrange itself above the chem-SAM.
The three immobilization steps (HDT, SpyTag peptide and nanobody–SpyCatcher) were monitored through X-ray photoelectron spectroscopy (XPS), quartz crystal microbalance with dissipation (QCM-D), cyclic voltammetry and electrochemical impedance spectroscopy (EIS). Upon functionalization with the HDT layer, the gold electrode displayed characteristic thiol–gold (S–Au) and free thiol (–SH) peaks in its high-resolution S 2p XPS spectrum31,32, indicating the upright orientation of HDT linked to gold through only one of the two –SH moieties (with the other terminal pointing away from the surface) (Fig. 2c and Supplementary Table 1). As we successively introduced SpyTag peptide and then the nanobody fusion protein on top of the chem-SAM, a new peak appeared at about 163.2 eV, which we attributed to the S–C bond from protein methionine residues (Fig. 2c)33. High-resolution C 1s and N 1s XPS spectra further demonstrated the bio-SAM formation by revealing C–O, C–N, and C–OOR bonds and nitrogen groups on the surface (Supplementary Fig. 1 and Supplementary Discussion 1). The formation of all layers was further corroborated by EIS and cyclic voltammetry measurements, which showed a decrease in the electrochemical capacitance of the gold electrode and an increase in its charge transfer resistance with the addition of (insulating) chem-SAM and bio-SAM (Supplementary Fig. 2 and Supplementary Discussion 2).
To assess the packing density of biomolecules on the surface, we monitored by QCM-D the immobilization of (1) the maleimide-modified SpyTag peptide and (2) the nanobody–SpyCatcher fusion protein on the chem-SAM. We quantified the masses gained from the two conjugates to be 114 and 406 ng cm−2, respectively (Fig. 2d, Supplementary Fig. 3 and Supplementary Discussion 3). Based on molecular weights of 1.76 and 28.4 kDa for the SpyTag peptide and nanobody–SpyCatcher protein, respectively, 39 × 1012 SpyTag peptides and 8.6 × 1012 nanobody–SpyCatcher molecules were coupled per cm2. This density translates to a mean inter-particle distance of only 3.4 nm between the SpyCatcher domains at the base of the bio-SAM. Given the size of this domain (~4 nm × 2.5 nm), we are thus approaching the maximum coupling density that is physically feasible. The formation of this exceptionally high-density biorecognition layer is probably further facilitated by the compact nanobody, which has similar dimensions to the SpyCatcher domain but is given additional freedom to pack and reposition itself through the flexible inter-domain linker (Fig. 2b).
Optimization of protein detection
We used the detection of GFP with the GFP nanobody-conjugated gate electrode as a model to optimize multiple parameters of the sensor design. Initial optimization rounds addressed unspecific binding, which was eliminated by adding bovine serum albumin (BSA) and a mild detergent (Tween-20) to both the nanobody–SpyCatcher protein immobilization and the target-binding solutions (Supplementary Discussion 3). Comparing QCM-D binding traces with (Supplementary Fig. 3) and without BSA (Fig. 2d), we suggest that BSA may be primarily capturing contaminating proteins in solution rather than binding to and blocking the surface in a classical sense. This view is supported by QCM-D traces recorded with another nanobody construct (Supplementary Fig. 4), and by the absence of fluorescently labeled BSA on the gate electrode surface (Supplementary Fig. 5). Nevertheless, the inclusion of BSA in the functionalization protocol was vital for the sensor performance as the addition of BSA to SpyCatcher protein and to the target-binding buffer outperformed other blocking strategies (Supplementary Fig. 6). Next, we optimized the OECT geometry through systematic variation of gate and channel sizes. The highest sensitivity towards GFP was attained with an OECT with a gate electrode area of 0.64 mm2, a channel width of 100 µm and a length of 10 µm (Supplementary Fig. 7).
The optimized poly(3,4-ethylenedioxythiophene) doped with poly(styrene sulfonate) (PEDOT:PSS) OECTs were operated in phosphate-buffered saline (PBS) under ambient conditions by varying the gate voltage (VG) between −0.6 and 0.6 V while the drain voltage (VD) was swept from 0 to −0.6 V. We assessed the sensing signal by monitoring the drain current (ID) as a function of VG at a fixed VD = −0.6 V. The reference (blank) response of the sensor was acquired by exposing the nanobody-functionalized gate electrode to PBS (Fig. 3b). The same electrode was then incubated for 10 min with a 5-µl drop of binding buffer containing GFP, rinsed twice in PBS and then stacked vertically on top of the channel separated by PBS (pH 7.4; ionic strength = 0.162 M) to complete the OECT biosensor (Fig. 1a). See Supplementary Fig. 8 for a comparison with a longer incubation time. After gate exposure to GFP, ID underwent a notable decrease at all of the gate voltages applied. As the GFP concentration increased, this sensor response became more evident (Fig. 3b). Similarly, the transconductance of the devices decreased as GFP interacted with the gate electrode (Supplementary Fig. 9a). In contrast, the device characteristics did not change if the gate electrode was incubated in solutions containing equivalent molar concentrations of non-target molecules such as an FRB–mCherry fusion (Fig. 3c and Supplementary Fig. 9b) and lysozyme (Supplementary Fig. 10). Lysozyme is abundant in saliva, which we envisioned as a potential biological medium for our SARS-CoV-2 application. mCherry is a red fluorescent protein with low sequence homology but very high structural similarity to GFP and thus challenged the sensor specificity.
a, Chemical structure of the PEDOT:PSS channel material. b,c, Steady-state characteristics of the depletion-mode PEDOT:PSS OECT gated with a GFP nanobody-functionalized electrode are shown when exposed to its target GFP (b) or its non-target mCherry (c). The arrow indicates increasing protein concentration. d, Resulting normalized response of the sensor to GFP, mCherry and lysozyme. e, Chemical structure of the novel p(g0T2-g6T2) channel material. f,g, Steady-state characteristics of the accumulation-mode OECT after exposure to GFP target (f) or non-target mCherry (g). The arrow indicates increasing protein concentration. h, Normalized response to GFP, mCherry and lysozyme. The normalized response was calculated as \({\rm{normalized}}\,{\rm{response}} = (|I_{\mathrm{D}} - I_0|)/I_0\) (the current change normalized to the baseline signal). The average value of the normalized response is plotted. The error bars in d and h represent the standard deviation calculated from at least three gate electrodes. Note that the PEDOT:PSS channel of the depletion-mode OECT is initially on (that is, high ID) and switches off (that is, low ID) upon the injection of cations (VG sweeping from −0.6 to +0.6 V). Conversely, the p(g0T2-g6T2) channel of the accumulation-mode OECT is initially off and switches on upon the injection of anions (VG sweeping from +0.2 to −0.6 V). In both cases, analyte binding reduces the capacitive coupling between the gate and the channel and lowers the gating efficiency and hence the magnitude of ID.
The trend in ID as well as gm can be clearly attributed to the specific binding of GFP to the nanobody-functionalized gate electrode. Fluorescence images confirmed the specific capture of GFP but not mCherry (Supplementary Fig. 11). The interaction between the nanobody and the GFP increases the impedance at the gate electrode/electrolyte interface (Supplementary Fig. 12), resulting in an increase in the potential decrease therein, which leads to a less efficient capacitive coupling between the gate and the channel (Supplementary Discussion 4). Hence, we measure a reduction in the current and its modulation as GFP binding weakens the electrical driving force acting on the ions. The OECT here provides voltage-to-current transduction and acts as a transconductance amplifier. Torsi and co-workers20 observed similar changes in PEDOT:PSS OECT characteristics, where the ID decreased upon a protein binding event at the functionalized gate electrode. We note that the decrease in ID is not due to a degradation of the channel over consecutive I–V cycles (Supplementary Fig. 13). Neither did we observe a particular change in the gate current (IG) upon GFP binding, and the value of IG was six orders of magnitude lower than the on current of the OECT (Supplementary Fig. 14). Within the gate voltage range that we operate the devices in, no Faradaic reactions that generate current occur at the gate electrode upon analyte binding.
To quantify the sensor response and minimize device-to-device variations, we calculated the normalized response of the OECT by normalizing the target protein-induced change in ID at a single VD and VG to its value previously measured after exposure to the blank solution. The normalized response versus GFP calibration curve revealed that the nanobody-functionalized OECT detects GFP with a lowest limit of detection (LOD) of 23 fM (Fig. 3d). The normalized response was <5% even for the highest concentrations of negative controls (mCherry and lysozyme), while it exceeded 30% for higher (nanomolar) GFP concentrations. Overall, PEDOT:PSS-based OECT biosensors were sensitive to the presence of 6 × 104 GFP molecules in 5 µl of buffer and responded within a dynamic range of six orders of magnitude. This sensitivity is on par with commercial colorimetric enzyme-linked immunosorbent assays, which take hours to complete, require larger sample volumes and have a far narrower dynamic range34.
Sensing with accumulation-mode OECTs
OECTs can be constructed for either depletion- or accumulation-mode operation. Currently, the most often used channel material is PEDOT:PSS; this is intrinsically doped and therefore conductive by default. PEDOT:PSS OECTs thus operate in depletion-mode. Application of a positive gate voltage VG injects cations into the channel, which compensate for the depleted holes and switch the transistor off. For continuous and long-term use, the depletion-mode operation may negatively affect material stability and power consumption as it implies high operating currents and relatively high VG to run the device. Recent studies expanded the available OECT channel materials through the chemical design of undoped, conjugated backbones, which were functionalized with hydrophilic side chains that facilitate ion injection and transport in the film16,35. OECTs comprising these materials operate in accumulation-mode (that is, the channel is initially off and generates a current only upon application of a small gate voltage)17. This operation mode allows for low-power electronics applications, improves device stability and is more compatible with integrated circuit designs that conventionally involve accumulation-mode transistors. Nevertheless, accumulation-mode OECTs for biochemical sensing have not yet been reported. We therefore replaced PEDOT:PSS with a recently designed conjugated polymer, p(g0T2-g6T2)36 (Fig. 3e). p(g0T2-g6T2) is a mixed (ionic and electronic) conductor. A negative VG pushes anions into the film that compensate for the holes injected from the metal contacts, thus turning the device on (Supplementary Fig. 15). The maximum gm (15 mS) occurs at a VG of around −0.3 V (Supplementary Fig. 15b). The transconductance efficiency (gm obtained per unit current) is very high at low VG (with the maximum slope of 60 mV dec−1 at VG ≈ 0.15 V; Supplementary Fig. 15c) due to the exponential ID versus VG characteristics. This behavior is similar to the subthreshold region of operation in traditional inorganic metal oxide semiconductor FETs37. The material also shows exceptional operational and environmental stability (Supplementary Fig. 16), thus overcoming the limitations of the currently available organic mixed (semi)conductors36.
Figure 3f shows the typical transfer characteristics of p(g0T2-g6T2) OECTs gated with the nanobody-functionalized gold electrodes incubated in solutions with varying GFP concentrations. As GFP bound to the gate, ID decreased for all gate voltages. This decrease was accompanied by a significant shift in the threshold voltage (Vth) towards more negative values (Supplementary Fig. 17). Similar to PEDOT:PSS sensors, p(g0T2-g6T2) OECTs showed no significant response to mCherry (Fig. 3g) and lysozyme (Supplementary Fig. 18) (at closer inspection, our data suggest a minor cross-reaction of the anti-GFP nanobody with the structurally similar mCherry). The sensor responded to a GFP concentration as low as 4.7 aM (normalized response = 7%) (Fig. 3h) with a dynamic range spanning ten orders of magnitude. We calculated the lowest LOD for these devices to be 14 aM, corresponding to 42 molecules in the 5 μl sample. Compared with the otherwise identical PEDOT:PSS-based sensor, the accumulation-mode OECT thus showed a 1000-fold improved sensitivity (aM compared with fM) and yet a larger dynamic range (ten orders of magnitude, from aM to nM, compared with six orders of magnitude). Moreover, we noted a much lower operating voltage. The target response was maximized (and the off-target response was minimized) at VG = VD = −0.1 V, compared with VG = VD = −0.6 V for the PEDOT:PSS OECT (Supplementary Fig. 19). Such characteristics would, in principle, allow for nanobody–OECT reader devices with a very compact build and long battery life.
The ability to detect the binding of analytes to only a few out of 55 billion receptors on our gate electrode surface seems intriguing. Monitoring of single-molecule binding to large gate electrodes was previously demonstrated in an electrolyte-gated organic FET recognizing immunoglobulin G as the target13. Macchia and colleagues14 attributed single-molecule detection to a combination of: (1) the 1000-fold amplification of input to output signal provided by their transistor; and (2) the perturbation of a tight hydrogen-bonding network in their chem-SAM, which they suggested to propagate as a defect over larger areas of the electrode13,38. In contrast, our sensor does not have a hydrogen-bonding network in or near the chem-SAM. However, the OECTs developed in this work have higher gains (the gm is on the order of mS rather than µS of typical electrolyte-gated organic FETs), permitting the amplification of input signals by a factor of one million. Indeed, without the amplification endowed by the OECT circuitry, we cannot resolve gate capacitance changes at such low concentrations (Supplementary Fig. 12a,b). A further improvement over previous designs is the very dense coupling of compact and intact receptor units in a, presumably, near-optimal binding orientation. Assuming the domain is diffusing within a 20-nm layer above the sensor surface (Fig. 2b), we obtain a local nanobody concentration of ~700 µM, which is far above the dissociation constant. Each target molecule diffusing into this capture layer will therefore necessarily bind (Supplementary Discussion 5). Once bound, the analyte is extremely unlikely to escape the sensor surface owing to a low koff rate39 (~1.5 × 10−4 s−1, corresponding to an 80-min half-life, which is unmeasurable according to our own surface plasmon resonance (SPR) measurements; Supplementary Fig. 20). The likelihood of analyte escape is further reduced by a high possibility of re-binding within the high-density receptor phase40. Our gate electrode thus operates as a kinetic trap for target molecules, while unspecific contaminants are washed away due to their faster off rates. Analyte diffusion from the binding volume to the sensor surface40 is probably a major bottleneck41, which we address through a large gate surface42 and intermittent mixing during incubation. Moreover, the high receptor density makes unproductive collisions between analyte and surface very unlikely. The large gate area and high receptor density also combine to a high binding capacity (corresponding to an analyte concentration of 18 nM in the 5 µl solution volume), which increases the sensor’s dynamic range. In summary, sensor performance is thus driven by the high transconductance, the large binding electrode surface and a very high receptor coupling density and quality.
Detection of SARS-CoV-2 and Middle East respiratory syndrome coronavirus (MERS-CoV) antigens
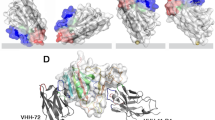
The VHH72 nanobody was originally raised against the SARS-CoV receptor-binding domain (RBD). Subsequently, VHH72 was shown to also bind SARS-CoV-2 at a slightly reduced affinity, and it became the first publicly available nanobody sequence against this new target43. Our VHH72–SpyCatcher fusion construct expressed well in E. coli and could be purified to a high yield (54 mg l−1 culture) as a monomeric protein. Although SARS-CoV-2 was our primary diagnostic target, we also designed and expressed constructs for the detection of MERS-CoV based on the previously reported nanobodies VHH83 (ref. 44), VHH04 (ref. 44) and VHH55 (ref. 43). The VHH04–SpyCatcher fusion yielded high-quality monomeric protein and was advanced to OECT experiments. Both VHH72 and VHH04 bind to the RBD of the homotrimeric (SARS or MERS, respectively) coronavirus spike protein45, which protrudes from the virus surface in multiple (about 100) copies. The RBD is a small (27-kDa) protein domain within the larger S1 subunit (76 kDa) and is directly responsible for recognition and binding of the specific host cell receptor. Detection of RBD would therefore be a good indicator for the presence of infectious viral particles. SPR experiments confirmed the binding of both nanobody fusion proteins with their respective, recombinantly expressed SARS-CoV, SARS-CoV-2 or MERS-CoV target proteins (Supplementary Fig. 21). As expected, VHH04 showed the highest affinity (KD = 0.1 nM) and slowest koff (2 × 10−4 s−1) for its target MERS-CoV S1. VHH72 bound its primary target SARS-CoV(1) RBD with lower affinity (KD = 7 nM), owing to a faster dissociation (3 × 10−3 s−1). The interaction with the closely related RBD of SARS-CoV-2 showed a mixed signal, where a smaller share of fast-on, fast-off binding events was distorting the main slow-on, slow-off binding regimen (KD = 23 nM; koff = 3 × 10−4 s−1). Such a signal was also reported for the interaction with a more recently developed nanobody46 and may indicate an intrinsic heterogeneity of this RBD.
Nevertheless, SARS-CoV nanobody-functionalized OECTs showed an excellent response to the SARS-CoV-2 RBD and S1 subunit, regardless of the channel material used (Fig. 4a,b). Incubation of the same gate electrodes with various concentrations of GFP did not evoke any current response, demonstrating the high specificity of the VHH72-functionalized gate for the viral proteins. The device also responded to the original target of VHH72—the SARS-CoV-1 RBD (Supplementary Fig. 22). In a direct comparison, the higher affinity of VHH72 for the SARS-CoV-1 RBD compared with the SARS-CoV-2 RBD only translated into a small signal increase. In contrast, the larger SARS-CoV-2 S1 spike protein (comprising the same RBD) generated much larger current changes than the isolated RBD. This increase in the normalized response may stem from higher affinity or size or both. We assume that the S1 subunit will, at least partially, form trimers in solution (all concentrations given in this study refer to the monomer) and thus benefit from avidity effects with substantially increased affinity (and prolonged residence) on the gate surface. Compared with the RBD alone, either the S1 monomer or the trimer would also cover a larger area on the functionalized surface than the isolated RBD. Akin to our GFP sensors, p(g0T2-g6T2) OECTs showed higher sensitivities than PEDOT:PSS devices. p(g0T2-g6T2) OECTs detected SARS-CoV-2 S1 at 4.7 aM with a 30% change in the normalized response (s.d. = 7% at most), translating to a nominal LOD of 18 zM. In fact, a single-molecule in our 5 µl measurement volume corresponded to a concentration of 0.33 aM, which appeared to be easily detected.
a,b, Normalized response of SARS-CoV nanobody-functionalized OECTs to SARS-CoV-2 RBD or S1 and GFP for channels comprising PEDOT:PSS (a) and p(g0T2-g6T2) (b). c, Response of SARS-CoV nanobody-functionalized OECTs comprising p(g0T2-g6T2) channels to SARS-CoV-2 proteins, GFP and MERS-CoV S1 spiked into human saliva. d, Normalized response of SARS-CoV-2 sensors at single-molecule concentrations. The theoretical number of spike protein molecules within the 5 µl sample is indicated above each data point. e, Normalized response of SARS-CoV-2 sensors to randomly selected saliva samples containing various amounts of SARS-CoV-2 S1. One of the samples (1.25 nM target protein) was measured with the GFP nanobody gate as a negative control. In a–e, the blank measurements were obtained after incubation in PBS. The average value of the normalized response is plotted. The error bars show the s.e.m. from three gate electrodes measured on one channel. f, Response of MERS nanobody-functionalized OECTs comprising p(g0T2-g6T2) channels to MERS-CoV S1, SARS-CoV-2 S1 and GFP spiked into human saliva. The average normalized response is shown, along with error bars representing the s.d. calculated from measurements with at least three gate electrodes. The normalized response was calculated as \({\rm{normalized}}\, {\rm{response}} = (|I_{\mathrm{D}} - I_0|)/I_0\) (the current change normalized to the baseline signal).
Having confirmed the single-molecule sensitivity and high specificity of SARS-CoV nanobody–OECTs when target proteins were captured in the binding buffer, we turned our attention to sensor performance in saliva. For RT-PCR detection of SARS-CoV-2, saliva samples match nasal swabs in sensitivity and specificity47,48,49 and are clearly the better medium for POC or field applications. We therefore challenged SARS-CoV nanobody-functionalized OECTs comprising p(g0T2-g6T2) channels with human saliva samples into which we spiked predetermined amounts of target and non-target proteins. The normalized response increased with the concentration of SARS-CoV-2 proteins in the saliva and showed only negligible change when the same gate electrode was exposed to GFP (Fig. 4c). As before, the sensor response already picked up at the single-molecule concentration threshold (Fig. 4d). The sensitivity for SARS-CoV-2 S1 in saliva (LOD = 1.2 × 10−21 M) was comparable to that in buffer (LOD = 1.8 × 10−20 M), indicating that the complex composition of saliva did not hamper the association between the nanobody and its target. The OECT could also detect SARS-CoV-2 S1 mixed into human serum (Supplementary Fig. 23), indicating its potential use for a wide range of other diagnostic applications.
So far, we have characterized sensor responses through the repeated exposure of one gate electrode to increasing target concentrations (all of the results are averages over several electrodes gating several channels each). To better emulate real-world screening conditions, we directly exposed individual gate electrodes to a single random concentration of SARS-CoV-2 S1 in saliva (Fig. 4e). The normalized response values varied accordingly and corresponded to the values obtained in the regular dose curves. In contrast, even high concentrations of SARS-CoV-2 S1 did not evoke any response on a gate functionalized with GFP nanobody. Nevertheless, the determination of exact concentration values is currently hindered by variation between individual gate electrodes and channels, which is exacerbated by stochastic variations in molecule numbers at very low concentrations. The manual analyte incubation and mixing may cause additional error. When moving towards practical applications, accurate concentration measurements will probably require a more streamlined and automated fabrication and operation of the device.
MERS-CoV is the only close relative of SARS-CoV-2 in active circulation50. The MERS-CoV nanobody-functionalized gate electrode detected single MERS-CoV S1 protein molecules in saliva, with a nominal detection range from 100 zM to 1 nM (Fig. 4f). A weak cross-reaction of VHH04 with SARS-CoV-2 target protein could already be anticipated from SPR measurements (Supplementary Fig. 21). In contrast, despite the high structural similarity between the two proteins, the SARS-CoV sensor did not respond to MERS-CoV S1 (Fig. 4c). Remarkably, the MERS-CoV sensor again achieved single-molecule sensitivity and ten orders of magnitude dynamic range in complex biological media without previous optimization to this particular target (see Supplementary Table 2 for the LODs of all of the sensors used in this study and Supplementary Table 3 for the sensitivity values). Our findings thus demonstrate the sensitivity and selectivity of nanobody–OECTs in complex biological media and at physiologically and clinically relevant protein concentrations. We expect that the nanobody–OECT technology can be rapidly adapted to detect any protein antigen for which nanobodies are available. A comparison of our OECT sensor performance with other similar thin-film transistor-based devices developed for immunosensing indicates the superiority of this sensor technology (for example, in terms of dynamic range, power consumption, miniaturization and compatibility with unprocessed human samples; Supplementary Table 4). OECTs also compare favorably to optical methods for single-molecule detection, such as flow-based immuonassays51 and plasmonic nanosensors52: Targets are not limited by size, and the lack of labeling allows for very short sample preparation and device operation times.
Detection of SARS-CoV-2 in clinical samples
We first validated our sensors on human nasopharyngeal swab samples collected from outpatients at a clinical testing facility. Swabs were received in a commercial universal transport medium (UTM; proprietary composition) and analyzed after 2 days. A calibration curve with SARS-CoV-2 S1 protein confirmed sensor operation in this UTM, albeit at lowered sensitivity (the LOD for SARS-CoV-2 S1 in UTM was 1.9 × 10−14 M) (Supplementary Fig. 24). All four nasal swab samples that had previously been tested as positive were confirmed to be positive by the OECT. Notably, all samples had low viral loads (RT-PCR cycle threshold = 30 to 37; close to the detection limit (cycle threshold = 40)). For a direct comparison of detection limits, we prepared a tenfold dilution series in UTM of one patient sample, mixed each dilution 1:1 with the binding buffer and subjected it side by side to sensor measurements, as well as RNA extraction and RT-PCR (Fig. 5b). The sensor’s above-noise response appeared to outperform RT-PCR by one order of magnitude.
a, Normalized sensor response for nasopharyngeal swab samples (n = 11) from healthy volunteers (1–7) and COVID-19-positive samples from walk-in patients (8–11). Cycle threshold values from RT-PCR are indicated above the positive samples. b, Sensor response for a tenfold dilution series of nasal swab sample number 8, along with RT-PCR results after RNA extraction from the same diluted samples (values above the bars). ND, not detected. The error bars in a and b indicate the s.d. measured from two gate electrodes combined with three channels. All positive samples were also measured using two GFP nanobody gate electrodes each. c, Average sensor response for saliva samples (n = 13) taken from healthy volunteers (12–17) and outpatients who had recently tested positive for COVID-19 (18–24). RT-qPCR was performed on each sample and the viral copy numbers in the 5 µl sample volume are indicated above the bars. The error bars show the s.e.m. from three gate electrodes measured on one channel. Measurements with one GFP nanobody gate per COVID-19-positive sample served as an additional negative control. As before, the normalized response was calculated as \({\rm{normalized}} \, {\rm{response}} = (|I_{\mathrm{D}} - I_0|)/I_0\) (the current change normalized to the baseline signal). K, thousand; M, million.
In a second validation, saliva was collected from six healthy volunteers and seven volunteers who had recently been confirmed as COVID-19 positive. Each sample was diluted fourfold in a modified, virus-inactivating lysis buffer and subjected to three independent measurements. Positive samples were additionally measured with one GFP nanobody gate electrode each. Quantitative PCR with reverse transcription (RT-qPCR) on each sample (calibrated against N-gene complementary DNA) confirmed the six negatives and revealed a wide range of viral loads from 2,700 to 1.8 × 1012 copies per ml for the others (Supplementary Table 5 and Supplementary Fig. 25). Despite substantial variation in individual sensor readings, average OECT signals reliably agreed with RT-qPCR on the presence or absence of SARS-CoV-2 in all samples (Fig. 5c and Supplementary Fig. 26). However, in contrast with the previous protein measurements, there was no correlation between sensor signal and the viral loads established by RT-PCR. However, the dilution series of virus material from a nasal swab in commercial UTM shows that a concentration-dependent signal can be achieved (Fig. 5b). We suspect that our sensor response is obscured by the co-recruitment of spike proteins attached to viral debris of largely varying size and composition. Our preliminary experience with patient samples thus suggests a qualitative sensor performance on par with RT-PCR and RT-qPCR, but also points to areas that need to be optimized for its application as a diagnostic device.
Outlook
We have introduced a label-free electrochemical immunosensing technology with single-molecule sensitivity and a large dynamic range. The nanobody–OECT biosensor detects specific protein molecules from unprocessed samples under ambient conditions after only 10 min of incubation with the disposable biofunctionalized gate electrode. The sensor can be rapidly adapted to any target for which nanobodies exist or can be raised. After optimizing the sensor for GFP detection, the simple exchange of nanobodies gave attomolar sensitivities for SARS-CoV-2 and MERS spike proteins in saliva or buffers without any further modification to the sensor or protocol. Preliminary tests with samples from patients with COVID-19 and healthy volunteers demonstrate a sensor accuracy and sensitivity on par with RT-PCR. For COVID-19, the detection of intact spike protein may, in fact, be a better indicator of infectious virus than RT-PCR, as the latter is also detecting membrane-associated viral RNA fragments long after the infection has been cleared53.
We attribute the performance of our sensor to several synergistic design choices. The OECT architecture outperforms other transistor technologies in terms of signal amplification and responds to changes far from the sensor surface, despite its simplicity and miniaturized geometry. These characteristics allowed us to tether biorecognition domains through long flexible linkers. A bioconjugation strategy, based on the autocatalytic assembly of a protein adapter with a chemically synthesized peptide, allowed for the oriented coupling of chemically unmodified recognition proteins at an exceptionally high density, which was limited only by the size of the proteins used. The choice of nanobodies over classic antibodies (or antibody fragments) further improved the density and robustness of the biorecognition layer. The resulting three-dimensional and flexible capture layer displayed a kinetically controlled target-binding regimen that is not limited by equilibrium dissociation constants. Moreover, a new solution-processable organic semiconductor enabled the sensor to operate in accumulation-mode, which further improved the sensitivity and stability by providing large changes in the output current for operation at very low-biasing conditions.
Several additional features recommend this sensor architecture for POC applications. The transistor base capable of recording from multiple micrometer-scale channels is reusable, allowing a stable electronic base to be combined with disposable biofunctionalized gate electrodes. Power consumption of the sensor is minimal and compatible with a hand-held battery-driven reader. Test results are currently obtained in less than 15 min, and we expect that changes in our 10-min incubation protocol can further accelerate the procedure. Real-time measurements are technically feasible when the sensor is integrated with microfluidics. Our current laboratory-scale prototype relies on careful handling by skilled and experienced operators. However, the functionalization is simple and modular, and both the gate electrode and the transistor base are easy to fabricate. The device operation consists of four easy steps: blank measurement, incubation, rinsing and sample measurement. Device assembly and packaging will now have to be engineered into, for instance, a single-housing flow-cell format that can be used by minimally trained personnel. The speed, performance and versatility of our nanobody-functionalized OECT, and its compatibility with many sample types, suggest that this biosensor technology can complement or replace a wide range of clinical and non-clinical diagnostic assays.
Methods
Materials
Sodium chloride, Tween-20, glycerol, HEPES, BSA, (3-glycidyloxypropyl) trimethoxysilane (GOPS), sodium dodecylbenzenesulfonate, ethylene glycol, HDT, 1,3-propanedithiol, 3-mercaptopropionic acid, albumin–fluorescein isothiocyanate conjugated bovine protein, human serum and PBS (pH 7.4) were purchased from Sigma–Aldrich and used as received. PEDOT:PSS (PH1000) was received from Heraeus. All aqueous solutions were prepared with ultrapure water (Millipore Milli-Q). p(g0T2-g6T2) was synthesized according to a procedure reported previously36. The protein purification materials used were: agar, LB broth, 2x YT broth, kanamycin, glucose, isopropyl β-d-1-thiogalactopyranoside, BugBuster (Novagen), cOmplete Protease Inhibitor mix (Sigma–Aldrich), Benzonase (Novagen), egg lysozyme (Fluka), Tris(2 carboxyethyl)phosphine, Tris(hydroxymethyl)aminomethane hydrochloride (Tris-HCl), imidazole, glycerol, dithiothreitol (DTT), ethylenediaminetetraacetic acid (EDTA), d-desthiobiotin and 10K Amicon Ultra spin concentrators (Milipore). The following purification columns and SPR materials were purchased from GE Healthcare: HisTrap HP (5 ml), StrepTrap HP (5 ml), Superdex75 16/600 and Biacore NTA SPR sensor chips (BR100034). The MCA–SpyTag peptide was from GenScript (Peptide Synthesis Service). The following viral target proteins were purchased from Sino Biological: SARS-CoV RBD (40150-V08B2), SARS-CoV-2 RBD (40592-V08H), SARS-CoV-2 S1 (40591-V08B1) and MERS-CoV S1 (40069-V08H). The Universal Transport Medium kit (UTM proprietary composition) was purchased from Noble Biosciences.
Fabrication of OECTs and the gate electrode
OECTs were microfabricated on glass substrates based on established protocols using standard photolithography and parylene C peel-off techniques54,55. The process starts with the first layer of photoresist (AZ5214) being spin coated and exposed to ultraviolet light using contact aligner to create Au electrodes and interconnection pads. The photoresist patterns were generated with AZ 726 MIF developer, followed by metal sputtering of 10 nm Cr and 100 nm Au and a standard lift-off process using hot dimethyl sulfoxide. Next, we coated the second layer of photoresist AZ9260 on the substrates and developed them using AZ developer. A parylene C layer was deposited to insulate the gold interconnects. The OECT channel was patterned by reactive ion etching, using a second layer of parylene C, which was peeled off to yield a length of 10 µm and a width of 100 µm. The aqueous dispersion of PEDOT:PSS containing ethylene glycol (5 vol%), sodium dodecylbenzenesulfonate (0.25 vol%) and GOPS (1 wt%) was sonicated for 30 min and then spin coated (3,000 r.p.m.; 45 s) on the substrates, leading to a film thickness of ~160 nm. The PEDOT:PSS OECTs were annealed at 140 °C for 1 h to activate GOPS and avoid dissolution of the polymer film in aqueous medium. p(g0T2-g6T2) films were spin coated (800 r.p.m.; 45 s) from a chloroform solution (5 g l−1) on the substrates to yield a film thickness of ~70 nm in the channel. All devices were rinsed with deionized water before use.
To fabricate the gate electrode, we used 175-µm-thick Kapton (polyimide) substrates. The Kapton was first cut with a CO2 laser (Universal Laser Systems; PLS6.75) into a square geometry that defined the sensor active area (0.8 × 0.8 mm). We then sonicated the substrates, first in isopropyl alcohol and then in deionized water for 30 min each. We sputtered 10 nm chromium or titanium and 100 nm Au onto these cleaned substrates. Before the functionalization, the electrodes were electrochemically cleaned in 10 mM sulfuric acid (H2SO4) using cyclic voltammetry. Ten cyclic voltammetry cycles with a potential between −0.2 and 1.2 V were applied at a scan rate of 100 mV s−1.
Biofunctionalization of gate electrodes
The chem-SAM solution was prepared in absolute ethanol containing 1 mM HDT, as described by others56. The Au electrodes were immersed in this solution for 1 h32. The electrodes were rinsed with absolute ethanol and dried under an N2 stream. The electrodes were then incubated for 1 h with the synthetic maleimide-modified SpyTag peptide (0.1 mg ml−1) in PBS, then washed in PBS. The electrodes were then incubated for 1 h with nanobody–SpyCatcher fusion protein (anti-GFP, anti-SARS-CoV-1 or anti-MERS-CoV) diluted to 50 µM in sensor binding buffer (100 mM HEPES (pH 7.4), 150 mM NaCl, 0.05% vol/vol Tween-20, 0.02% wt/vol NaN3 and 0.1% wt/vol BSA), then rinsed with PBS. Nanobody-functionalized gate electrodes were stored for up to 1 week at 4 °C in the sensor binding buffer until use.
XPS
XPS analysis was performed using a Kratos AXIS Supra instrument equipped with a monochromatic Al Kα X-ray source (1468.6 eV). We operated the source at 75 W under ultra-high vacuum conditions (~10−9 mbar). The spectra were recorded in a hybrid mode using electrostatic and magnetic lenses and an aperture slot of 300 × 700 µm. The survey and high-resolution spectra were acquired at fixed analyzer pass energies of 80 and 20 eV, respectively. We mounted the samples in a floating mode to avoid differential charging. The spectra were then acquired under charge neutralization conditions. We calibrated the spectra to a reference of C 1s at 284.8 eV. The Tougaard method was used for background subtraction, which was deconvoluted using Gaussian and Lorentzian methods with XPSPEAK41.
QCM-D
QCM-D measurements were conducted using a QSense analyzer (QE401; Biolin Scientific). The piezoelectrically active gold sensor (0.7854 cm2) was first modified with HDT SAM under the same reaction conditions and then mounted into the QCM-D setup. First, the QCM-D signals, including the change in frequency (Δf) and dissipation (ΔD) were stabilized in PBS. Second, the peptide solution (0.1 mg ml−1 SpyTag peptide in PBS) was injected into the chamber with a flow rate of 100 µl min−1 controlled by a peristaltic pump. After ensuring that the sensor was fully covered with the solution, we stopped the pump for 1 h and rinsed the sensor surface with PBS injected into the system for 15 min. The same procedure was employed to treat the surface with SpyCatcher-linked nanobody solution (50 µM in the binding buffer). All QCM-D data presented in this work were recorded at the seventh overtone. To quantify the mass accumulating on the sensor (Δm) and the thickness of the layers during the functionalization steps, we used the Sauerbrey equation (1)57:
where n is the overtone number selected for the calculations and −17.7 is a constant calculated based on the resonant frequency, active area, density and shear modulus of the quartz crystal sensor. The mass of the binding molecules on the sensor surface was then estimated using their molecular weight. The (Sauerbrey) thickness (nm), ds, was calculated as:
where Δm is the change in mass (ng), A is the area (cm2) and r is the estimated density of the layer (g cm−3; assumed to be the same as water, which is taken as 1 g cm−3).
Electrochemical characterization
The electrochemical characteristics of the gold electrode were investigated before and after the formation of chem-SAM and bio-SAM using EIS and cyclic voltammetry performed in a three-electrode setup using a potentiostat (Autolab PGSTAT128N with Nova software; Metrohm). We used a platinum wire and an Ag/AgCl electrode as the counter electrode and reference electrodes, respectively, while the gold electrode was connected as the working electrode. All measurements were carried out in 5 ml of 10 mM PBS (pH 7.4) containing 10 mM [Fe(CN)6]3−/4−. For cyclic the voltammetry measurements, the potential window was between −0.2 and 0.5 V and the scan rate was 100 mV s−1. Electrochemical impedance spectra were recorded at a d.c. voltage of 0 V versus the open circuit potential (Voc) and an a.c. modulation of 10 mV over a frequency range of 0.1–100,000 Hz. For the analysis of GFP nanobody-functionalized electrodes, the electrodes were incubated with GFP solutions for 10 min and washed in 10 mM PBS before the measurements. The data were analyzed using Nova software with the appropriate equivalent circuit modeling.
Proteins and peptides
Nanobody–SpyCatcher fusion proteins were designed based on available structures (nanobody: Protein Data Bank (PDB) ID 4PFE58; SpyCatcher PDB ID 4MLI59) with the nanobody placed at the amino-terminal end of the fusion protein in order to orient the common VHH target-binding interface towards the bulk solution, away from the sensing surface. Protein sequences were reverse-translated and codon-optimized for expression in E. coli with an in-house Python script based on DNA Chisel60. Plasmids for protein expression were gene synthesized by Twist Bioscience (United States) or Bio Basic (Canada) in our customized expression vector pJE411c with kanamycin resistance and modified with an RBS insulator (BCD2) cassette61 for improved translation initiation. Plasmids were transformed into E. coli BL21 (DE3) and starter cultures were inoculated overnight from a single colony. 1 l production cultures in 2x YT medium with 50 mg l−1 kanamycin and 1% glucose were inoculated 1:100, grown at 37 °C and 250 r.p.m. to and optical density measured at 600 nm of 0.8, induced with 0.5 mM β-d-1-thiogalactopyranoside and incubated with shaking for 18 h at 25 °C. Cells were harvested by centrifugation for 10 min at 6000 × g and 4 °C, washed once with cold PBS, resuspended in lysis buffer (25 mM Tris-HCl (pH 7.4), 500 mM NaCl, 10 mM imidazole, 10% glycerol, SIGMAFAST protease inhibitor, 25 U ml−1 Benzonase HC (Milipore) and 2 mM DTT) and homogenized with a cell disruptor (Constant Systems). Earlier purifications of mCherry, GFP nanobody and msfGFP used different lysis methods (chemical lysis with BugBuster (Novagen) for mCherry; sonication for the other two). Lysates were cleared by centrifugation at 87,000 × g for 45 min and the supernatant was filtered through Miracloth tissue (Milipore) and subjected to affinity chromatography on an ÄKTA FPLC (GE Healthcare) using either StrepTrap HP or HisTrap HP columns (GE Healthcare), depending on the purification tag. The Strep-tag binding buffer was 100 mM Tris-HCl (pH 8), 150 mM NaCl, 1 mM EDTA, 5% glycerol and 0.5 mM Tris(2 carboxyethyl)phosphine, and elution was performed with 2.5 mM desthiobiotin in binding buffer. The His-tag binding buffer was 25 mM Tris-HCl (pH 7.4), 500 mM NaCl, 10 mM imidazole, 10% glycerol and 2 mM DTT, and elution was performed with a four-step imidazole gradient up to 0.5 M. Fractions were pooled and concentrated using 10K Amicon Ultra (Milipore), followed by gel filtration on a Superdex75 16/600 column (GE Healthcare) into 20 mM HEPES (pH 7.5), 300 mM NaCl, 10% glycerol and 50 µM EDTA. After spin concentration, aliquots were snap-frozen in liquid nitrogen and stored at −80 °C. Protein purity, quality and accurate molar mass were monitored by sodium dodecyl sulfate polyacrylamide gel electrophoresis, as well as size-exclusion chromatography multi-angle light scattering on a DAWN HELEOS II and OptiLab T-rEx (Wyatt). Protein concentrations were determined on a NanoDrop spectrophotometer by absorbance at 280 nm using sequence-specific extinction coefficients (https://web.expasy.org/protparam/).
SARS-CoV-1/2 and MERS target proteins, expressed and purified from HEK293 cells or insect cell culture, were received lyophilized from Sino Biological (China) and dissolved to a standard concentration of 0.25 mg ml−1, per the manufacturer’s instructions, then aliquoted, snap-frozen in liquid nitrogen and stored at −80 °C. Amino-terminally maleimide-labeled SpyTag peptides were synthesized by GenScript Biotech (Singapore), received lyophilized, dissolved in PBS and stored at −20 °C.
Protein dilutions
Target and non-target proteins were thawed on ice and centrifuged at 15,000 r.p.m. at 4 °C for 30–45 min to remove any potential aggregates (although no aggregation was observed). Sino Biological proteins were then used as provided for the preparation of dilution series starting at 320 or 640 nM. Equivalent dilutions of the Sino Biological storage buffer by itself did not give any sensor response. Weak background sensor signals were recorded from dilutions of DTT. In-house proteins were therefore stored or exchanged into DTT-free buffer before use. The higher-concentrated proteins from in-house production were first diluted to intermediate concentrations that could still be validated and corrected spectrophotometrically. Protein dilutions were prepared in standard sensor binding buffer (100 mM HEPES (pH 7.4), 150 mM NaCl, 0.05% vol/vol Tween-20, 0.02% wt/vol NaN3 and 0.1% wt/vol BSA), which was modified for saliva spike-in measurements to include complete protease inhibitor cocktail with EDTA (Sigma–Aldrich) at four times the manufacturer-recommended concentration (giving a twofold concentration in the final 1:1 mixture with saliva). BSA was not included in this saliva buffer. For measurements in the regular binding buffer, fourfold dilution series were prepared in 96-well microplates over 23 steps starting from 320 nM. For measurements in serum, saliva and UTM, target protein dilution series were prepared in the appropriate buffer (standard or saliva binding buffer) starting from 640 nM so that final concentrations were identical after 1:1 mixture with serum, saliva or UTM.
SPR measurements
SPR measurements were performed on a Biacore T200 instrument using Ni-NTA sensor chips (GE Healthcare) and a modified running buffer (20 mM HEPES (pH 7.4), 150 mM NaCl, 0.05% Tween-20, 0.02% wt/vol NaN3 and 50 µM EDTA, prepared at room temperature and filtered) mirroring the sensor binding buffer. All analyte proteins were desalted into the running buffer (HiPrep 26/10 column; GE Healthcare) before the preparation of dilution series in the same running buffer. The GFP nanobody–GFP interaction was measured with the His10-tagged nanobody immobilized (ligand) and msfGFP in solution. For the remaining measurements, since all viral target proteins arrived with a non-cleavable His-tag, target proteins were immobilized instead of the nanobody. All ligand proteins gave stable immobilization responses with minimal signal loss over time. His tags were removed from nanobody–SpyCatcher proteins by overnight cleavage with excess 3C protease (produced in house), followed by gel filtration on a Superdex 200 Increase 10/300GL column. All ligand proteins were immobilized to equal loading levels of around 100 RU at 10 µl min−1 flow rates. Binding and unbinding experiments were run at a flow rate of 80 µl min−1. Two replicates of each dilution series were prepared and measured in the course of the same experiment. Between each measurement cycle, the sensor surface was regenerated with 0.35 M EDTA in running buffer and re-charged with 0.5 mM NiSO4 in water. Biacore results were analyzed with the manufacturer analysis software version 2.1 following standard procedures (double subtraction of the reference channel and buffer injection signal) applying a 1:1 binding model and simultaneous curve fitting. The SARS-CoV-2 RBD results were instead fit to a heterologous ligand binding model.
Fluorescence imaging
Imaging was performed on a DMI8 inverted fluorescence microscope (Leica Microsystems) coupled with a pE-4000 fluorescence illumination system (CoolLED), and the images were processed using ImageJ software. Electrodes were placed between two cover slips in the presence of PBS, to keep the flexible material in plane and in focus. The presence or absence of BSA was studied by adding fluorescently labeled BSA (albumin–fluorescein isothiocyanate conjugated bovine protein) to the nanobody–SpyCatcher solution during functionalization of the sensing surface. The capture of GFP was examined by imaging already functionalized gold electrodes before and after incubation with GFP or control solutions.
OECT characterization and sensor operation
We used a Keithley Source Meter operating by LabVIEW code to apply drain (VD) and gate voltages (VG) and to obtain gate and channel currents (IG and ID) under ambient conditions. A polydimethylsiloxane well was glued on top of the channels and filled with 75 µl of 10 mM PBS. The steady-state transistor characteristics were obtained by measuring ID versus VD at various VG, for PEDOT:PSS applied between −0.6 and 0.6 V with 0.1-V steps (2.5 V s−1), and for p(g0T2-g6T2) from 0.2 to −0.6 V with 0.05-V steps (1.25 V s−1). VD was swept from 0 to −0.6 V. We first chose a channel and obtained its transfer curve (ID versus VG) in PBS using the nanobody-functionalized gate electrode incubated for 10 min in the buffer solution (buffer, saliva, UTM or serum, in the absence of target proteins). The currents obtained were used as the baseline signal (I0). The same gate electrode was then incubated with 5 µl of the solution (buffer, saliva, UTM or serum) containing the protein target for 10 min (pipetting for 30 s every 3 min). The electrode was washed thoroughly with PBS to remove any unbound proteins. To obtain calibration curves, we prepared various concentrations of the target protein and collected device data starting from the most diluted one. The normalized response of the sensor was calculated according to the following equation:
where ID is the current response of the sensor to an analyte solution that the gate was exposed to.
According to the International Union of Pure and Applied Chemistry, the LOD was calculated as the concentration leading to a response that equalled the average of the noise level plus three times the noise standard deviation13,62. The level of noise \(\left( {\frac{{{\Delta}I}}{{I_0}} \pm 3\delta } \right)\) was taken from the relative current variation in negative control sensors.
where \(\frac{{{\Delta}I}}{{I_0}}\) is the average response of the blank sample, δ is the relative standard deviation and a and b are the intercept and slope of the calibration curve, respectively.
Clinical sample preparation and testing
The clinical samples (saliva samples and nasopharyngeal swabs) used in this study (Fig. 5) were collected from human subjects as part of registered protocols approved by the Institutional Review Board of the King Faisal Specialist Hospital and Research Center and King Abdullah University of Science and Technology (KAUST) Institutional Biosafety and Bioethics Committee (IBEC). All volunteers provided signed consent to participate in the study. Nasopharyngeal swabs collected from outpatients with COVID-19 and healthy subjects were stored in UTM at 4 °C and tested within 1–2 d of collection. Saliva was collected in 50-ml tubes and always measured the same day. The saliva used for original sensor characterization with recombinant proteins (Fig. 4) was self-collected in the morning before food or tooth brushing, filtered and stored in aliquots at −20 °C. All volunteers provided signed consent to participate in the study and provide saliva. The buffer composition for measurements in clinical samples was modified in the course of the study: nasal swabs were measured after 1:1 dilution in the original buffer used for the saliva spike-in experiments (100 mM HEPES (pH 7.4), 150 mM NaCl, 0.05% vol/vol Tween-20, 0.02% wt/vol NaN3 and 4x cOmplete). Raw saliva samples were measured after 1:3 dilution in a virus-inactivating lysis buffer (50 mM Tris (pH 7.4), 250 mM NaCl, 1% Nonidet P-40, 0.02% NaN3, 0.5% BSA and 4x cOmplete). Human male AB plasma (serum) was purchased from Sigma–Aldrich and used as received. All protocols and procedures involving human saliva, nasopharyngeal swabs and serum were approved by the KAUST IBEC (under approval numbers 18IBEC11 and 20IBEC25) and National Committee of BioEthics, Saudi Arabia (registration number HAP-02-J-042). The study conducted at the hospital was approved by the Office of Research Affairs of the King Faisal Specialist Hospital and Research Centre (RAC #220 0014).
Reporting Summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data supporting the results of this study are available within the paper and its Supplementary Information. All of the data generated during this study and used to produce the figures are available from Figshare at https://doi.org/10.6084/m9.figshare.14420588.
References
Kissler, S. M., Tedijanto, C., Goldstein, E., Grad, Y. H. & Lipsitch, M. Projecting the transmission dynamics of SARS-CoV-2 through the postpandemic period. Science 368, 860–868 (2020).
Lewis, D. Mounting evidence suggests coronavirus is airborne—but health advice has not caught up. Nature 583, 510–513 (2020).
Oran, D. P. & Topol, E. J. Prevalence of asymptomatic SARS-CoV-2 infection. Ann. Intern. Med. https://doi.org/10.7326/M20-3012 (2020).
Long, Q.-X. et al. Clinical and immunological assessment of asymptomatic SARS-CoV-2 infections. Nat. Med. 26, 1200–1204 (2020).
WHO. WHO Director-General’s Opening Remarks at the Media Briefing on COVID-19—16 March 2020 (World Health Organization, 2020); https://www.who.int/dg/speeches/detail/who-director-general-s-opening-remarks-at-the-media-briefing-on-covid-19---16-march-2020
Corman, V. M. et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eurosurveillance 25, 2000045 (2020).
Ferretti, L. et al. Quantifying SARS-CoV-2 transmission suggests epidemic control with digital contact tracing. Science 368, eabb6936 (2020).
Krueger, L. J. et al. Evaluation of the accuracy, ease of use and limit of detection of novel, rapid, antigen-detecting point-of-care diagnostics for SARS-CoV-2. Preprint at medRxiv https://doi.org/10.1101/2020.10.01.20203836 (2020).
Niemz, A., Ferguson, T. M. & Boyle, D. S. Point-of-care nucleic acid testing for infectious diseases. Trends Biotechnol. 29, 240–250 (2011).
Wang, W. U., Chen, C., Lin, K., Fang, Y. & Lieber, C. M. Label-free detection of small-molecule–protein interactions by using nanowire nanosensors. Proc. Natl Acad. Sci. USA 102, 3208–3212 (2005).
Ren, R. et al. Nanopore extended field-effect transistor for selective single-molecule biosensing. Nat. Commun. 8, 586 (2017).
Nakatsuka, N. et al. Aptamer–field-effect transistors overcome Debye length limitations for small-molecule sensing. Science 362, 319–324 (2018).
Macchia, E. et al. Single-molecule detection with a millimetre-sized transistor. Nat. Commun. 9, 3223 (2018).
Macchia, E. et al. About the amplification factors in organic bioelectronic sensors. Mater. Horiz. 7, 999–1013 (2020).
Khodagholy, D. et al. High transconductance organic electrochemical transistors. Nat. Commun. 4, 2133 (2013).
Inal, S., Rivnay, J., Suiu, A.-O., Malliaras, G. G. & McCulloch, I. Conjugated polymers in bioelectronics. Acc. Chem. Res. 51, 1368–1376 (2018).
Rivnay, J. et al. Organic electrochemical transistors. Nat. Rev. Mater. 3, 17086 (2018).
Ohayon, D. & Inal, S. Organic bioelectronics: from functional materials to next-generation devices and power sources. Adv. Mater. 32, 2001439 (2020).
Gentili, D. et al. Integration of organic electrochemical transistors and immuno-affinity membranes for label-free detection of interleukin-6 in the physiological concentration range through antibody–antigen recognition. J. Mater. Chem. B 6, 5400–5406 (2018).
Macchia, E. et al. Ultra-sensitive protein detection with organic electrochemical transistors printed on plastic substrates. Flex. Print. Electron. 3, 034002 (2018).
Trilling, A. K., Beekwilder, J. & Zuilhof, H. Antibody orientation on biosensor surfaces: a minireview. Analyst 138, 1619–1627 (2013).
Muyldermans, S. Nanobodies: natural single-domain antibodies. Annu. Rev. Biochem. 82, 775–797 (2013).
Hamers-Casterman, C. et al. Naturally occurring antibodies devoid of light chains. Nature 363, 446–448 (1993).
Steeland, S., Vandenbroucke, R. E. & Libert, C. Nanobodies as therapeutics: big opportunities for small antibodies. Drug Discov. Today 21, 1076–1113 (2016).
De Meyer, T., Muyldermans, S. & Depicker, A. Nanobody-based products as research and diagnostic tools. Trends Biotechnol. 32, 263–270 (2014).
Filipiak, M. S. et al. Highly sensitive, selective and label-free protein detection in physiological solutions using carbon nanotube transistors with nanobody receptors. Sens. Actuators B Chem. 255, 1507–1516 (2018).
Rothbauer, U. et al. A versatile nanotrap for biochemical and functional studies with fluorescent fusion proteins. Mol. Cell. Proteom. 7, 282–289 (2008).
Zakeri, B. et al. Peptide tag forming a rapid covalent bond to a protein, through engineering a bacterial adhesin. Proc. Natl Acad. Sci. USA 109, E690–E697 (2012).
Keeble, A. H. & Howarth, M. Insider information on successful covalent protein coupling with help from SpyBank. Methods Enzymol. 617, 443–461 (2019).
Oloketuyi, S. et al. Electrochemical immunosensor functionalized with nanobodies for the detection of the toxic microalgae Alexandrium minutum using glassy carbon electrode modified with gold nanoparticles. Biosens. Bioelectron. 154, 112052 (2020).
Esplandiu, M. J. & Noeske, P.-L. M. XPS investigations on the interactions of 1,6-hexanedithiol/Au(111) layers with metallic and ionic silver species. Appl. Surf. Sci. 199, 166–182 (2002).
Love, J. C., Estroff, L. A., Kriebel, J. K., Nuzzo, R. G. & Whitesides, G. M. Self-assembled monolayers of thiolates on metals as a form of nanotechnology. Chem. Rev. 105, 1103–1170 (2005).
Weser, U. in Cation Ordering and Electron Transfer (eds Gleitzer, C. et al.) 145–160 (Springer, 1985).
Zhang, S., Garcia-D’Angeli, A., Brennan, J. P. & Huo, Q. Predicting detection limits of enzyme-linked immunosorbent assay (ELISA) and bioanalytical techniques in general. Analyst 139, 439–445 (2013).
Moser, M., Ponder, J. F., Wadsworth, A., Giovannitti, A. & McCulloch, I. Materials in organic electrochemical transistors for bioelectronic applications: past, present, and future. Adv. Funct. Mater. 29, 1807033 (2019).
Moser, M. et al. Side chain redistribution as a strategy to boost organic electrochemical transistor performance and stability. Adv. Mater. 32, 2002748 (2020).
Venkatraman, V. et al. Subthreshold operation of organic electrochemical transistors for biosignal amplification. Adv. Sci. 5, 1800453 (2018).
Macchia, E. et al. Label-free and selective single-molecule bioelectronic sensing with a millimeter-wide self-assembled monolayer of anti-immunoglobulins. Chem. Mater. 31, 6476–6483 (2019).
Kubala, M. H., Kovtun, O., Alexandrov, K. & Collins, B. M. Structural and thermodynamic analysis of the GFP:GFP-nanobody complex. Protein Sci. 19, 2389–2401 (2010).
Schuck, P. & Zhao, H. The role of mass transport limitation and surface heterogeneity in the biophysical characterization of macromolecular binding processes by SPR biosensing. Methods Mol. Biol. 627, 15–54 (2010).
Sheehan, P. E. & Whitman, L. J. Detection limits for nanoscale biosensors. Nano Lett. 5, 803–807 (2005).
Macchia, E., Manoli, K., Di Franco, C., Scamarcio, G. & Torsi, L. New trends in single-molecule bioanalytical detection. Anal. Bioanal. Chem. 412, 5005–5014 (2020).
Wrapp, D. et al. Structural basis for potent neutralization of betacoronaviruses by single-domain Camelid antibodies. Cell 181, 1004–1015.e15 (2020).
Raj, S. V. et al. Chimeric camel/human heavy-chain antibodies protect against MERS-CoV infection. Sci. Adv. 4, eaas9667 (2018).
Walls, A. C. et al. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 181, 281–292.e6 (2020).
Hanke, L. et al. An alpaca nanobody neutralizes SARS-CoV-2 by blocking receptor interaction. Nat. Commun. 11, 4420 (2020).
Pasomsub, E. et al. Saliva sample as a non-invasive specimen for the diagnosis of coronavirus disease 2019: a cross-sectional study. Clin. Microbiol. Infect. 27, 285.E1–285.E4 (2021).
Wyllie, A. L. et al. Saliva or nasopharyngeal swab specimens for detection of SARS-CoV-2. N. Engl. J. Med. 383, 1283–1286 (2020).
Butler-Laporte, G. et al. Comparison of saliva and nasopharyngeal swab nucleic acid amplification testing for detection of SARS-CoV-2: a systematic review and meta-analysis. JAMA Intern. Med. 181, 353–360 (2021).
Azhar, E. I., Hui, D. S. C., Memish, Z. A., Drosten, C. & Zumla, A. The Middle East respiratory syndrome (MERS). Infect. Dis. Clin. 33, 891–905 (2019).
Todd, J. et al. Ultrasensitive flow-based immunoassays using single-molecule counting. Clin. Chem. 53, 1990–1995 (2007).
Spitzberg, J. D., Zrehen, A., van Kooten, X. F & Meller, A. Plasmonic-nanopore biosensors for superior single-molecule detection. Adv. Mater. 31, 1900422 (2019).
Alexandersen, S., Chamings, A. & Bhatta, T. R. SARS-CoV-2 genomic and subgenomic RNAs in diagnostic samples are not an indicator of active replication. Nat. Commun. 11, 6059 (2020).
Ohayon, D. et al. Biofuel powered glucose detection in bodily fluids with an n-type conjugated polymer. Nat. Mater. 19, 456–463 (2020).
Wustoni, S., Savva, A., Sun, R., Bihar, E. & Inal, S. Enzyme-free detection of glucose with a hybrid conductive gel electrode. Adv. Mater. Interfaces 6, 1800928 (2019).
Qu, D. et al. 1,6-Hexanedithiol self-assembled monolayers on Au(111) investigated by electrochemical, spectroscopic, and molecular mechanics methods. J. Phys. Chem. C. 114, 497–505 (2010).
Höök, F., Rodahl, M., Brzezinski, P. & Kasemo, B. Energy dissipation kinetics for protein and antibody–antigen adsorption under shear oscillation on a quartz crystal microbalance. Langmuir 14, 729–734 (1998).
Eshaghi, M. et al. Rational structure-based design of bright GFP-based complexes with tunable dimerization. Angew. Chem. Int. Ed. Engl. 54, 13952–13956 (2015).
Li, L., Fierer, J. O., Rapoport, T. A. & Howarth, M. Structural analysis and optimization of the covalent association between SpyCatcher and a peptide Tag. J. Mol. Biol. 426, 309–317 (2014).
Zulkower, V. & Rosser, S. DNA Chisel, a versatile sequence optimizer. Bioinformatics 36, 4508–4509 (2020).
Mutalik, V. K. et al. Precise and reliable gene expression via standard transcription and translation initiation elements. Nat. Methods 10, 354–360 (2013).
Long, G. L. & Winefordner, J. D. Limit of detection. A closer look at the IUPAC definition. Anal. Chem. 55, 712A–724A (1983).
Acknowledgements
Figure 1 was produced by X. Pita, a scientific illustrator at KAUST. We thank all of the members of the KAUST Rapid Research Response Team (R3T) for COVID-19, especially S. Hamdan, for contributions in this study. We thank S. Mfarrej and A. K. Subudhi for providing access to and assisting with the experiments in the Biosafety Level 2+ experimental room at KAUST. We thank the KAUST Health team (operated by Dr. Soliman Fakeeh Hospital, Jeddah), including D. Buttigieg and M. Habib, for providing clinical samples. We thank staff at the King Faisal Specialist Hospital and Research Center (Riyadh), particularly A. Alzahrani, M. Alsanea and F. Alhadeq, for help with organizing and hosting some of the clinical studies. We thank the KAUST nanofabrication core laboratory team, D. Rosas Villalva and U. Buttner for help with device fabrication and integration. This work was initiated thanks to the KAUST Impact Acceleration Fund (IAF) program. The research reported in this publication was supported by funding from the Office of Sponsored Research (OSR) at KAUST under award numbers REI/1/4204-01, REI/1/4229-01, OSR-2018-CRG7-3709, OSR-2018-CARF/CCF-3079, OSR-2015-CRG4-2572 and OSR-4106 CPF2019. We acknowledge EC FP7 Project SC2 (610115), EC H2020 (643791) and EPSRC Projects EP/G037515/1, EP/M005143/1 and EP/L016702/1.
Author information
Authors and Affiliations
Contributions
S.I., S.T.A. and R.G. conceived of the research, designed the experiments, supervised the work and wrote the manuscript. K.G., S.W. and A.K. fabricated the devices and performed the OECT experiments. K.G. and S.W. functionalized the gate electrodes. A.K. performed the EIS measurements and conducted the modelling. S.W. performed the cyclic voltammetry and XPS measurements. K.G. conducted the QCM-D experiments. R.G. and E.D.-G. designed and produced the recombinant proteins and performed the Biacore experiments. A.H. took the fluorescence microscope images and developed the LabVIEW codes to operate the OECTs. A.N.A. collected the clinical samples. M.S. and A.P. provided the RT-qPCR results. F.S.A. supervised the nasal swab collection and conducted the RT-PCR with A.A.A. and R.G. M.M. and I.M. provided the p-type material. All authors were involved in the discussion and participated in manuscript input.
Corresponding authors
Ethics declarations
Competing interests
S.I., S.T.A., R.G., K.G., S.W. and A.K. have filed a patent application (US Patent application no. 63/047,547; “Nanobody-functionalized electrochemical transistors and methods of making and using thereof”) related to this work.
Additional information
Peer review information Nature Biomedical Engineering thanks Howard Katz and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Discussion, Figs. 1–26, Tables 1–7 and references.
Rights and permissions
About this article
Cite this article
Guo, K., Wustoni, S., Koklu, A. et al. Rapid single-molecule detection of COVID-19 and MERS antigens via nanobody-functionalized organic electrochemical transistors. Nat Biomed Eng 5, 666–677 (2021). https://doi.org/10.1038/s41551-021-00734-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-021-00734-9
This article is cited by
-
Advances in nanobiosensors during the COVID-19 pandemic and future perspectives for the post-COVID era
Nano Convergence (2024)
-
Understanding asymmetric switching times in accumulation mode organic electrochemical transistors
Nature Materials (2024)
-
Designing organic mixed conductors for electrochemical transistor applications
Nature Reviews Materials (2024)
-
A hybrid transistor with transcriptionally controlled computation and plasticity
Nature Communications (2024)
-
Rapid deep learning-assisted predictive diagnostics for point-of-care testing
Nature Communications (2024)