Abstract
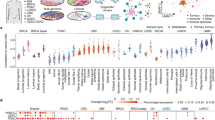
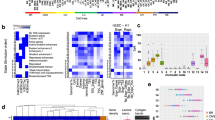
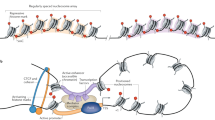
In cancer, linking epigenetic alterations to drivers of transformation has been difficult, in part because DNA methylation analyses must capture epigenetic variability, which is central to tumour heterogeneity and tumour plasticity. Here, by conducting a comprehensive analysis, based on information theory, of differences in methylation stochasticity in samples from patients with paediatric acute lymphoblastic leukaemia (ALL), we show that ALL epigenomes are stochastic and marked by increased methylation entropy at specific regulatory regions and genes. By integrating DNA methylation and single-cell gene-expression data, we arrived at a relationship between methylation entropy and gene-expression variability, and found that epigenetic changes in ALL converge on a shared set of genes that overlap with genetic drivers involved in chromosomal translocations across the disease spectrum. Our findings suggest that an epigenetically driven gene-regulation network, with UHRF1 (ubiquitin-like with PHD and RING finger domains 1) as a central node, links genetic drivers and epigenetic mediators in ALL.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
DNA-methylation and RNA-seq data are available at the Gene Expression Omnibus repository under the accession number GSE116229.
References
Mullighan, C. G. The molecular genetic makeup of acute lymphoblastic leukemia. Hematology Am. Soc. Hematol. Educ. Program 2012, 389–396 (2012).
Hunger, S. P. & Mullighan, C. G. Acute lymphoblastic leukemia in children. N. Engl. J. Med. 373, 1541–1552 (2015).
Grobner, S. N. et al. The landscape of genomic alterations across childhood cancers. Nature 555, 321–327 (2018).
Ma, X. et al. Pan-cancer genome and transcriptome analyses of 1,699 paediatric leukaemias and solid tumours. Nature 555, 371–376 (2018).
Kulis, M. et al. Whole-genome fingerprint of the DNA methylome during human B cell differentiation. Nat. Genet. 47, 746–756 (2015).
Figueroa, M.E. et al. Integrated genetic and epigenetic analysis of childhood acute lymphoblastic leukemia. J. Clin. Invest. 123, 3099–3111 (2013).
Nordlund, J. et al. Genome-wide signatures of differential DNA methylation in pediatric acute lymphoblastic leukemia. Genome Biol. 14, r105 (2013).
Mullighan, C. G. et al. CREBBP mutations in relapsed acute lymphoblastic leukaemia. Nature 471, 235–239 (2011).
Mar, B. G. et al. Mutations in epigenetic regulators including SETD2 are gained during relapse in paediatric acute lymphoblastic leukaemia. Nat. Commun. 5, 3469 (2014).
Milani, L. et al. DNA methylation for subtype classification and prediction of treatment outcome in patients with childhood acute lymphoblastic leukemia. Blood 115, 1214–1225 (2010).
Hogan, L.E. et al. Integrated genomic analysis of relapsed childhood acute lymphoblastic leukemia reveals therapeutic strategies. Blood 118, 5218–5226 (2011).
Lee, S. T. et al. Epigenetic remodeling in B-cell acute lymphoblastic leukemia occurs in two tracks and employs embryonic stem cell-like signatures. Nucleic Acids Res. 43, 2590–2602 (2015).
Wahlberg, P. et al. DNA methylome analysis of acute lymphoblastic leukemia cells reveals stochastic de novo DNA methylation in CpG islands. Epigenomics 8, 1367–1387 (2016).
Jenkinson, G., Pujadas, E., Goutsias, J. & Feinberg, A. P. Potential energy landscapes identify the information-theoretic nature of the epigenome. Nat. Genet. 49, 719–729 (2017).
Jenkinson, G., Abante, J., Feinberg, A. P. & Goutsias, J. An information-theoretic approach to the modeling and analysis of whole-genome bisulfite sequencing data. BMC Bioinformatics 19, 87 (2018).
Jenkinson, G., Abante, J., Koldobskiy, M. A., Feinberg, A. P. & Goutsias, J. Ranking genomic features using an information-theoretic measure of epigenetic discordance. BMC Bioinformatics 20, 175 (2019).
Landan, G. et al. Epigenetic polymorphism and the stochastic formation of differentially methylated regions in normal and cancerous tissues. Nat. Genet. 44, 1207–1214 (2012).
Wang, F. et al. CellMethy: identification of a focal concordantly methylated pattern of CpGs revealed wide differences between normal and cancer tissues. Sci. Rep. 5, 18037 (2015).
Lilljebjorn, H. et al. Identification of ETV6-RUNX1-like and DUX4-rearranged subtypes in paediatric B-cell precursor acute lymphoblastic leukaemia. Nat. Commun. 7, 11790 (2016).
Zhang, J. et al. Deregulation of DUX4 and ERG in acute lymphoblastic leukemia. Nat. Genet. 48, 1481–1489 (2016).
Tsuzuki, S., Taguchi, O. & Seto, M. Promotion and maintenance of leukemia by ERG. Blood 117, 3858–3868 (2011).
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Sharov, A. A. et al. Responsiveness of genes to manipulation of transcription factors in ES cells is associated with histone modifications and tissue specificity. BMC Genomics 12, 102 (2011).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Heerema, N. A. et al. Dicentric (9;20)(p11;q11) identified by fluorescence in situ hybridization in four pediatric acute lymphoblastic leukemia patients. Cancer Genet. Cytogenet. 92, 111–115 (1996).
Felice, M. S. et al. Prognostic impact of t(1;19)/TCF3–PBX1 in childhood acute lymphoblastic leukemia in the context of Berlin–Frankfurt–Munster-based protocols. Leuk. Lymphoma 52, 1215–1221 (2011).
Pui, C. H., Carroll, W. L., Meshinchi, S. & Arceci, R. J. Biology, risk stratification, and therapy of pediatric acute leukemias: an update. J. Clin. Oncol. 29, 551–565 (2011).
Bhojwani, D. et al. ETV6-RUNX1-positive childhood acute lymphoblastic leukemia: improved outcome with contemporary therapy. Leukemia 26, 265–270 (2012).
Paulsson, K. et al. The genomic landscape of high hyperdiploid childhood acute lymphoblastic leukemia. Nat. Genet. 47, 672–676 (2015).
Greaves, M. A causal mechanism for childhood acute lymphoblastic leukaemia. Nat. Rev. Cancer 18, 471–484 (2018).
De Braekeleer, E. et al. Acute lymphoblastic leukemia associated with RCSD1–ABL1 novel fusion gene has a distinct gene expression profile from BCR–ABL1 fusion. Leukemia 27, 1422–1424 (2013).
Goyama, S. et al. UBASH3B/Sts-1–CBL axis regulates myeloid proliferation in human preleukemia induced by AML1–ETO. Leukemia 30, 728–739 (2016).
Wernicke, C. M. et al. MondoA is highly overexpressed in acute lymphoblastic leukemia cells and modulates their metabolism, differentiation and survival. Leuk. Res. 36, 1185–1192 (2012).
Zhang, R. et al. A possible 5′-NRIP1/UHRF1-3′ fusion gene detected by array CGH analysis in a Ph+ ALL patient. Cancer Genet. 204, 687–691 (2011).
Sidhu, H. & Capalash, N. UHRF1: the key regulator of epigenetics and molecular target for cancer therapeutics. Tumour Biol. https://doi.org/10.1177/1010428317692205 (2017).
Ashraf, W. et al. The epigenetic integrator UHRF1: on the road to become a universal biomarker for cancer. Oncotarget 8, 51946–51962 (2017).
Chow, M. et al. Maintenance and pharmacologic targeting of ROR1 protein levels via UHRF1 in t(1;19) pre-B-ALL. Oncogene 37, 5221–5232 (2018).
Gibcus, J. H. & Dekker, J. The hierarchy of the 3D genome. Mol. Cell 49, 773–782 (2013).
Chow, M. L., Kim, D., Kamath, S., Peng, D. & Luu, M. Use of antiviral medications in drug reaction with eosinophilia and systemic symptoms (DRESS): a case of infantile DRESS. Pediatr. Dermatol. 35, e114–e116 (2018).
Yan, F. et al. Inhibition effect of siRNA-downregulated UHRF1 on breast cancer growth. Cancer Biother. Radiopharm. 26, 183–189 (2011).
Yan, F., Wang, X., Shao, L., Ge, M. & Hu, X. Analysis of UHRF1 expression in human ovarian cancer tissues and its regulation in cancer cell growth. Tumour Biol. 36, 8887–8893 (2015).
Ge, T. T., Yang, M., Chen, Z., Lou, G. & Gu, T. UHRF1 gene silencing inhibits cell proliferation and promotes cell apoptosis in human cervical squamous cell carcinoma CaSki cells. J. Ovarian Res. 9, 42 (2016).
Iacobucci, I. & Mullighan, C. G. Genetic basis of acute lymphoblastic leukemia. J. Clin. Oncol. 35, 975–983 (2017).
Lilljebjorn, H. & Fioretos, T. New oncogenic subtypes in pediatric B-cell precursor acute lymphoblastic leukemia. Blood 130, 1395–1401 (2017).
Reddy, K. L. & Feinberg, A. P. Higher order chromatin organization in cancer. Semin. Cancer Biol. 23, 109–115 (2013).
Shen, H. & Laird, P. W. Interplay between the cancer genome and epigenome. Cell 153, 38–55 (2013).
Timp, W. et al. Large hypomethylated blocks as a universal defining epigenetic alteration in human solid tumors. Genome Med. 6, 61 (2014).
Duran-Ferrer, M. et al. The proliferative history shapes the DNA methylome of B-cell tumors and predicts clinical outcome. Nat. Cancer 1, 1066–1081 (2020).
Pujadas, E. & Feinberg, A. P. Regulated noise in the epigenetic landscape of development and disease. Cell 148, 1123–1131 (2012).
Feinberg, A. P., Koldobskiy, M. A. & Gondor, A. Epigenetic modulators, modifiers and mediators in cancer aetiology and progression. Nat. Rev. Genet. 17, 284–299 (2016).
Zheng, S. C., Widschwendter, M. & Teschendorff, A. E. Epigenetic drift, epigenetic clocks and cancer risk. Epigenomics 8, 705–719 (2016).
Landau, D. A. et al. Locally disordered methylation forms the basis of intratumor methylome variation in chronic lymphocytic leukemia. Cancer Cell 26, 813–825 (2014).
Li, S. et al. Distinct evolution and dynamics of epigenetic and genetic heterogeneity in acute myeloid leukemia. Nat. Med. 22, 792–799 (2016).
Pan, H. et al. Epigenomic evolution in diffuse large B-cell lymphomas. Nat. Commun. 6, 6921 (2015).
Chan, T. E., Stumpf, M. P. H. & Babtie, A. C. Gene regulatory network inference from single-cell data using multivariate information measures. Cell Syst. 5, 251–267.e3 (2017).
Giovinazzo, H. et al. A high-throughput screen of pharmacologically active compounds for inhibitors of UHRF1 reveals epigenetic activity of anthracycline derivative chemotherapeutic drugs. Oncotarget 10, 3040 (2019).
Lefebvre, J. L., Kostadinov, D., Chen, W. V., Maniatis, T. & Sanes, J. R. Protocadherins mediate dendritic self-avoidance in the mammalian nervous system. Nature 488, 517–521 (2012).
El Hajj, N., Dittrich, M. & Haaf, T. Epigenetic dysregulation of protocadherins in human disease. Semin. Cell Dev. Biol. 69, 172–182 (2017).
Dias, S., Mansson, R., Gurbuxani, S., Sigvardsson, M. & Kee, B. L. E2A proteins promote development of lymphoid-primed multipotent progenitors. Immunity 29, 217–227 (2008).
Semerad, C. L., Mercer, E. M., Inlay, M. A., Weissman, I. L. & Murre, C. E2A proteins maintain the hematopoietic stem cell pool and promote the maturation of myelolymphoid and myeloerythroid progenitors. Proc. Natl Acad. Sci. USA 106, 1930–1935 (2009).
Hunger, S. P. et al. The t(1;19)(q23;p13) results in consistent fusion of E2A and PBX1 coding sequences in acute lymphoblastic leukemias. Blood 77, 687–693 (1991).
Inaba, T. et al. Fusion of the leucine zipper gene HLF to the E2A gene in human acute B-lineage leukemia. Science 257, 531–534 (1992).
Wu, H., Caffo, B., Jaffee, H. A., Irizarry, R. A. & Feinberg, A. P. Redefining CpG islands using hidden Markov models. Biostatistics 11, 499–514 (2010).
Ernst, J. & Kellis, M. Chromatin-state discovery and genome annotation with ChromHMM. Nat. Protoc. 12, 2478–2492 (2017).
Ernst, J. & Kellis, M. Large-scale imputation of epigenomic datasets for systematic annotation of diverse human tissues. Nat. Biotechnol. 33, 364–376 (2015).
Encode Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Press, W. H., Teukolsky, S. A., Vetterling, W. T. & Flannery, B. P. Numerical Recipes. The Art of Scientific Computing (Cambridge Univ. Press, 2007).
Benjamini, Y. & Yekutieli, D. The control of the false discovery rate in multiple testing under dependency. Ann. Stat. 29, 1165–1188 (2001).
Stasinopoulos, D. M. & Rigby, R. A. Generalized additive models for location scale and shape (GAMLSS) in R. J. Stat. Softw. 23, 1–46 (2007).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Kharchenko, P. V., Silberstein, L. & Scadden, D. T. Bayesian approach to single-cell differential expression analysis. Nat. Methods 11, 740–742 (2014).
Guo, G. et al. Serum-based culture conditions provoke gene expression variability in mouse embryonic stem cells as revealed by single-cell analysis. Cell Rep. 14, 956–965 (2016).
Jiao, Y., Widschwendter, M. & Teschendorff, A. E. A systems-level integrative framework for genome-wide DNA methylation and gene expression data identifies differential gene expression modules under epigenetic control. Bioinformatics 30, 2360–2366 (2014).
Fisher, R. A Statistical Methods, Experimental Design, and Statistical Inference (Oxford Univ. Press, 1990).
Acknowledgements
This work was supported by US National Institutes of Health grants R01 CA65438 and DP1 DK119129 to A.P.F., R01 HG006282 to H.J., US National Science Foundation grant CCF-1656201 to J.G., St. Baldrick’s Foundation fellowship and funding from Unravel Pediatric Cancer to M.A.K. M.A.K. is a Damon Runyon–Sohn Pediatric Cancer Fellow supported by the Damon Runyon Cancer Research Foundation (DRSG-15P-16). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
A.P.F. and M.A.K. designed the study and led the biological experiments. A.P.F. and J.G. supervised all aspects of the research. C.L.B., K.R.R. and P.A.B. provided the primary patient material and disease-specific expertise. A.I., C.C. and R.T. performed library preparation and sequencing. V.A.R.D. performed the single-cell RNA-seq experiments. E.P. performed WGBS quality control, preprocessing and bisulfite alignment. H.J. and W.Z. performed statistical analysis of bulk and single-cell RNA-seq data. G.J., J.A. and J.G. developed the data analysis methods. G.J. and J.A. implemented the data analysis methods. A.P.F., G.J., J.G. and M.A.K. analysed the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Biomedical Engineering thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures and table captions.
Supplementary Tables
WGBS samples, statistics and clinical features, ranked lists of genes, bulk RNA-seq results, single-cell RNA-seq data, gene regulatory-network modules, median JSD of comparisons, and more.
Rights and permissions
About this article
Cite this article
Koldobskiy, M.A., Jenkinson, G., Abante, J. et al. Converging genetic and epigenetic drivers of paediatric acute lymphoblastic leukaemia identified by an information-theoretic analysis. Nat Biomed Eng 5, 360–376 (2021). https://doi.org/10.1038/s41551-021-00703-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-021-00703-2