Abstract
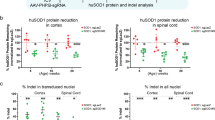
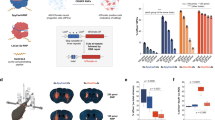
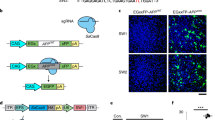
Technologies that can safely edit genes in the brains of adult animals may revolutionize the treatment of neurological diseases and the understanding of brain function. Here, we demonstrate that intracranial injection of CRISPR–Gold, a nonviral delivery vehicle for the CRISPR–Cas9 ribonucleoprotein, can edit genes in the brains of adult mice in multiple mouse models. CRISPR–Gold can deliver both Cas9 and Cpf1 ribonucleoproteins, and can edit all of the major cell types in the brain, including neurons, astrocytes and microglia, with undetectable levels of toxicity at the doses used. We also show that CRISPR–Gold designed to target the metabotropic glutamate receptor 5 (mGluR5) gene can efficiently reduce local mGluR5 levels in the striatum after an intracranial injection. The effect can also rescue mice from the exaggerated repetitive behaviours caused by fragile X syndrome, a common single-gene form of autism spectrum disorders. CRISPR–Gold may significantly accelerate the development of brain-targeted therapeutics and enable the rapid development of focal brain-knockout animal models.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Cho, S. W., Kim, S., Kim, J. M. & Kim, J. S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat. Biotechnol. 31, 230–232 (2013).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR–Cas system. Cell 163, 759–771 (2015).
Swiech, L. et al. In vivo interrogation of gene function in the mammalian brain using CRISPR–Cas9. Nat. Biotechnol. 33, 102–106 (2015).
Mingozzi, F. & High, K. A. Immune responses to AAV vectors: overcoming barriers to successful gene therapy. Blood 122, 23–36 (2013).
Ishida, K., Gee, P. & Hotta, A. Minimizing off-target mutagenesis risks caused by programmable nucleases. Int. J. Mol. Sci. 16, 24751–24771 (2015).
Watakabe, A. et al. Comparative analyses of adeno-associated viral vector serotypes 1, 2, 5, 8 and 9 in marmoset, mouse and macaque cerebral cortex. Neurosci. Res. 93, 144–157 (2015).
Staahl, B. T. et al. Efficient genome editing in the mouse brain by local delivery of engineered Cas9 ribonucleoprotein complexes. Nat. Biotechnol. 35, 431–434 (2017).
Kazdoba, T. M., Leach, P. T., Silverman, J. L. & Crawley, J. N. Modeling fragile X syndrome in the Fmr1 knockout mouse. Intractable Rare Dis. Res. 3, 118–133 (2014).
Persico, A. M. & Napolioni, V. Autism genetics. Behav. Brain Res. 251, 95–112 (2013).
Ji, N. Y. & Findling, R. L. Pharmacotherapy for mental health problems in people with intellectual disability. Curr. Opin. Psychiatry 29, 103–125 (2016).
Politte, L. C., Henry, C. A. & McDougle, C. J. Psychopharmacological interventions in autism spectrum disorder. Harv. Rev. Psychiatry 22, 76–92 (2014).
Bear, M. F., Huber, K. M. & Warren, S. T. The mGluR theory of fragile X mental retardation. Trends Neurosci. 27, 370–377 (2004).
Bear, M. F. Therapeutic implications of the mGluR theory of fragile X mental retardation. Genes Brain Behav. 4, 393–398 (2005).
Dölen, G. & Bear, M. F. Role for metabotropic glutamate receptor 5 (mGluR5) in the pathogenesis of fragile X syndrome. J. Physiol. 586, 1503–1508 (2008).
Osterweil, E. K., Krueger, D. D., Reinhold, K. & Bear, M. F. Hypersensitivity to mGluR5 and ERK1/2 leads to excessive protein synthesis in the hippocampus of a mouse model of fragile X syndrome. J. Neurosci. 30, 15616–15627 (2010).
Tao, J. et al. Negative allosteric modulation of mGluR5 partially corrects pathophysiology in a mouse model of Rett syndrome. J. Neurosci. 36, 11946–11958 (2016).
Silverman, J. L. et al. Negative allosteric modulation of the mGluR5 receptor reduces repetitive behaviors and rescues social deficits in mouse models of autism. Sci. Transl. Med. 4, 131ra151 (2012).
Jacquemont, S. et al. Epigenetic modification of the FMR1 gene in fragile X syndrome is associated with differential response to the mGluR5 antagonist AFQ056. Sci. Transl. Med. 3, 64ra61 (2011).
Raspa, M., Wheeler, A. C. & Riley, C. Public health literature review of fragile X syndrome. Pediatrics 139, S153–S171 (2017).
Lee, K. et al. Nanoparticle delivery of Cas9 ribonucleoprotein and donor DNA in vivo induces homology-directed DNA repair. Nat. Biomed. Eng. 1, 889–901 (2017).
Feng, G. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000).
Choi, G. B. et al. Driving opposing behaviors with ensembles of piriform neurons. Cell 146, 1004–1015 (2011).
Fried, I., Mukamel, R. & Kreiman, G. Internally generated preactivation of single neurons in human medial frontal cortex predicts volition. Neuron 69, 548–562 (2011).
McMahon, M. A. & Cleveland, D. W. Gene therapy: gene-editing therapy for neurological disease. Nat. Rev. Neurol. 13, 7–9 (2017).
Xie, N. et al. Reactivation of FMR1 by CRISPR/Cas9-mediated deletion of the expanded CGG-repeat of the fragile X chromosome. PLoS ONE 11, e0165499 (2016).
Park, C. Y. et al. Reversion of FMR1 methylation and silencing by editing the triplet Repeats in fragile X iPSC-derived neurons. Cell Rep. 13, 234–241 (2015).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Chen, Y. S. et al. Size-dependent impairment of cognition in mice caused by the injection of gold nanoparticles. Nanotechnology 21, 485102 (2010).
Ferreira, G. K. et al. Effect of acute and long-term administration of gold nanoparticles on biochemical parameters in rat brain. Mater. Sci. Eng. C Mater. Biol. Appl. 79, 748–755 (2017).
Almad, A. A. & Maragakis, N. J. Glia: an emerging target for neurological disease therapy. Stem Cell Res. Ther. 3, 37 (2012).
Sukoff Rizzo, S. J. & Crawley, J. N. Behavioral phenotyping assays for genetic mouse models of neurodevelopmental, neurodegenerative, and psychiatric disorders. Annu. Rev. Anim. Biosci. 5, 371–389 (2017).
Kim, H., Lim, C. S. & Kaang, B. K. Neuronal mechanisms and circuits underlying repetitive behaviors in mouse models of autism spectrum disorder. Behav. Brain Funct. 12, 3 (2016).
Spencer, C. M. et al. Modifying behavioral phenotypes in Fmr1 KO mice: genetic background differences reveal autistic-like responses. Autism Res. 4, 40–56 (2011).
Sungur, A., Vörckel, K. J., Schwarting, R. K. & Wöhr, M. Repetitive behaviors in the Shank1 knockout mouse model for autism spectrum disorder: developmental aspects and effects of social context. J. Neurosci. Methods 234, 92–100 (2014).
Ding, Q., Sethna, F. & Wang, H. Behavioral analysis of male and female Fmr1 knockout mice on C57BL/6 background. Behav. Brain Res. 271, 72–78 (2014).
Graham, D. R. & Sidhu, A. Mice expressing the A53T mutant form of human alpha-synuclein exhibit hyperactivity and reduced anxiety-like behavior. J. Neurosci. Res. 88, 1777–1783 (2010).
Masuda, T., Tsuda, M., Tozaki-Saitoh, H. & Inoue, K. Lentiviral transduction of cultured microglia. Methods Mol. Biol. 1041, 63–67 (2013).
Balcaitis, S., Weinstein, J. R., Li, S., Chamberlain, J. S. & Möller, T. Lentiviral transduction of microglial cells. Glia. 50, 48–55 (2005).
Burke, B., Sumner, S., Maitland, N. & Lewis, C. E. Macrophages in gene therapy: cellular delivery vehicles and in vivo targets. J. Leukoc. Biol. 72, 417–428 (2002).
Kim, H. J. et al. Introduction of stearoyl moieties into a biocompatible cationic polyaspartamide derivative, PAsp(DET), with endosomal escaping function for enhanced siRNA-mediated gene knockdown. J. Control. Release 145, 141–148 (2010).
Miyata, K. et al. Polyplexes from poly(aspartamide) bearing 1,2-diaminoethane side chains induce pH-selective, endosomal membrane destabilization with amplified transfection and negligible cytotoxicity. J. Am. Chem. Soc. 130, 16287–16294 (2008).
Tabebordbar, M. et al. In vivo gene editing in dystrophic mouse muscle and muscle stem cells. Science 351, 407–411 (2016).
Zhang, X., Servos, M. R. & Liu, J. Instantaneous and quantitative functionalization of gold nanoparticles with thiolated DNA using a pH-assisted and surfactant-free route. J. Am. Chem. Soc. 134, 7266–7269 (2012).
Lee, H. Y. et al. Bidirectional regulation of dendritic voltage-gated potassium channels by the fragile X mental retardation protein. Neuron 72, 630–642 (2011).
Khan, M. & Gasser, S. Generating primary fibroblast cultures from mouse ear and tail tissues. J. Vis. Exp. 107, 53565 (2016).
Lin, S., Staahl, B. T., Alla, R. K. & Doudna, J. A. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. elife 3, e04766 (2014).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Güuell, M., Yang, L. & Church, G. M. Genome editing assessment using CRISPR Genome Analyzer (CRISPR-GA). Bioinformatics 30, 2968–2970 (2014).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
McFarlane, H. G. et al. Autism-like behavioral phenotypes in BTBR T+tf/J mice. Genes Brain Behav. 7, 152–163 (2008).
Thomas, A. et al. Marble burying reflects a repetitive and perseverative behavior more than novelty-induced anxiety. Psychopharmacology 204, 361–373 (2009).
Acknowledgements
We thank J. Doudna for advice, B. Staahl for discussions and technical support, and H. Kim, A. Rao and K. Kataoka for technical support. We thank M. A. Bhat and members of the Bhat Lab for technical support. We thank M. West in the CIRM/QB3 Shared Stem Cell facility for technical support. This work was supported by the National Institutes of Health grant R01EB023776 to N.M, and by the National Science Foundation grant 1456862 to R.B.
Author information
Authors and Affiliations
Contributions
B.L., K.L., R.B., N.M. and H.Y.L. designed the research, and B.L., K.L., S.P., R.G.-R., A.C., H.M.P., V.B. and H.Y.L. performed the experiments and analyses. R.G.-R. and S.P. generated the videos. B.L., K.L., N.M. and H.Y.L. wrote the manuscript. H.Y.L. supervised the research. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
K.L., H.M.P. and N.M. are co-founders of GenEdit Inc. The remaining authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures, tables and video captions.
Supplementary Video 1
Marble-bury assay.
Supplementary Video 2
Empty-cage observations.
Rights and permissions
About this article
Cite this article
Lee, B., Lee, K., Panda, S. et al. Nanoparticle delivery of CRISPR into the brain rescues a mouse model of fragile X syndrome from exaggerated repetitive behaviours. Nat Biomed Eng 2, 497–507 (2018). https://doi.org/10.1038/s41551-018-0252-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-018-0252-8
This article is cited by
-
Engineering self-deliverable ribonucleoproteins for genome editing in the brain
Nature Communications (2024)
-
Nanomaterials for intelligent CRISPR-Cas tools: improving environment sustainability
Environmental Science and Pollution Research (2024)
-
A potential paradigm in CRISPR/Cas systems delivery: at the crossroad of microalgal gene editing and algal-mediated nanoparticles
Journal of Nanobiotechnology (2023)
-
Techniques for investigating lncRNA transcript functions in neurodevelopment
Molecular Psychiatry (2023)
-
A temporally resolved DNA framework state machine in living cells
Nature Machine Intelligence (2023)