Abstract
The ongoing severe acute respiratory syndrome-coronavirus (SARS-CoV-2) has triggered the coronavirus pandemic (COVID-19) that has claimed hundreds of thousands of lives worldwide. This virus spreads predominantly by human-to-human transmission via respiratory droplets. However, the presence of this virus in the fecal and anal swabs of infected patients has triggered the need for research into its waterborne transmission. The various environmental factors that impact the persistence of coronavirus in different water matrices include temperature, UV exposure, organic matter, disinfectants as well as adversarial microorganisms. This review summarizes the most recent research data on the effect of various factors on coronavirus in aqueous environments. The available data suggest that: (i) increasing temperature decreases the overall persistence of the virus; (ii) the presence of organic matter can increase the survivability of coronavirus; (iii) chlorine is the most effective and economic disinfectant; (iv) membrane bioreactors in wastewater treatment plants are hosts of competitive microorganisms that can inactivate coronaviruses; (v) ultraviolet irradiation is another effective option for virus inactivation. However, the inactivation disinfection kinetics of coronaviruses are yet to be fully understood. Thus, further research is needed to understand its fate and transport with respect to the water cycle so that effective strategies can be adopted to curb its effects. These strategies may vary based on geographic, climatic, technical, and social conditions around the globe. This paper explores possible approaches and especially the conditions that local communities and authorities should consider to find optimal solutions that can limit the spread of this virus.
Similar content being viewed by others
Introduction
As of June 2020, the novel coronavirus (CoV) pandemic has claimed more than five-hundred thousand lives worldwide with a total of more than 13 million cases since its first emergence in Wuhan, China in December 20191. On 30th January 2020, this outbreak was declared a Public Health Emergency of International Concern and on 11th February 2020, the World Health Organization (WHO) named it COVID-192. Subsequently, on 11th March 2020, the WHO declared this outbreak to be a pandemic3. Since last 18 years, there have been three outbreaks by different strains of this viral family: Severe Acute Respiratory Syndrome (SARS-CoV, China 2003), Middle East Respiratory Syndrome (MERS-CoV, Saudi Arabia 2012), and the ongoing COVID-19 (SARS-CoV-2, China 2019).
The coronaviruses (order: Nidovirales, family: Coronaviridae, subfamily: Coronavirinae) are single-stranded RNA viruses with size ranging from 60 to 220 nm. These viruses replicate using a nested set of RNAs, as is evident from the name of the order Nidovirales (“nido” meaning nest). The Coronavirinae subfamily is categorized into four genera: alpha, beta, gamma, and delta coronaviruses. Of these, the first two genera infect human beings. The alpha human coronaviruses (HCoVs) include HCoV-229E and HVoV-NL63, while the beta-human coronaviruses include HCoV-HKU1, HCoV-OC43, MERS-CoV, SARS-CoV, and SARS-CoV-24. It has been observed that the genome sequence of SARS-CoV-2 has 82% similarity to that of SARS-CoV5. The coronavirus is an enveloped virus with crownlike spikes on its surface. This viral envelope protects the genetic material in its life cycle when it is traveling between hosts’ cells. The function of the envelope protein includes the development of the envelope and pathogenesis. The viral envelope contains the helically structured nucleoprotein bound RNA. The RNA genomes of these viruses are known to be very large with length ranging from 27 to 32 kb6,7. The virion structure of coronavirus is illustrated in Fig. 1. The primary mode of transmission of this infection is via respiratory droplets of an infected individual that may be caused by breathing, coughing, sneezing, or other forms of bio-aerosols. The wide range of symptoms of this disease include fever, cough, shortness of breath, respiratory diseases, diarrhea, and other such health issues which often prove to be fatal as can be observed from the number of deaths due to COVID-19 since December 2019.
Although the major route of transmission of coronavirus is by droplets, historical evidence suggests that the possible spread of such pathogenic viruses via the fecal-oral pathway cannot be ruled out. There have been many instances when the fecal-oral pathway has led to the waterborne transmission of such pathogenic viruses that pose a great risk to public health5,6,7,8,9,10,11,12,13,14. Chan et al. reported the detection of SARS-CoV RNA in the fecal discharge of infected patients since the 5th day of the illness during the 2003 SARS-CoV epidemic15. Additionally, it was observed that the number of fecal samples testing positive for the presence of SARS-CoV RNA increased gradually with time and spiked at day 11 of the illness. Traces of the RNA were still at a detectable level even after 30 days of the illness. The viable presence of SARS-CoV RNA was also detected in the wastewater of two hospitals in Beijing treating SARS infected patients during the 2003 epidemic16. In a sporadic instance, the leaking of a sewage pipe in a residential complex of Amoy Gardens in Hong Kong led to the aerosolization of infected wastewater containing SARS-CoV. This later resulted in a cluster of coronavirus cases in Hong Kong17,18,19,20. Recently, Holshue et al. reported the detection of SARS-CoV-2 RNA in the fecal discharge of a patient on days 2 as well as 7 of the illness in Washington, USA21. Thus, existing data support the viable presence of the RNA of SARS-CoV-2 in the fecal discharge of infected patients and sewage of hospitals that can enter the urban water cycle via faulty plumbing, spillover, or contamination of surface water. Hence, it becomes imperative to study the viability of the virus in different water matrices, especially wastewater.
The present review summarizes the knowledge available on coronaviruses with respect to the urban water cycle and the environment. The influence of various environmental factors that can impact the survivability of the coronavirus in aqueous systems has been emphasized and summarized with available literature. The outcome of this study is aimed at enhancing our understanding of the persistence of the virus in different water matrices and being cognizant of the knowledge gaps so that it can be strengthened. This can aid us in risk assessment and management during future viral outbreaks.
Fate and transport of coronavirus through the urban water cycle
The urban water cycle consists of the following main components: (i) wastewater treatment plants (WWTPs), (ii) drinking water distribution system, and (iii) stormwater runoff management. During a viral outbreak, human viruses are excreted in the form of fecal matter, urine, vomit, and other discharges and can enter municipal wastewater by the sewage system. Faulty plumbing or leakage like detachment of vent pipe, drain to sewer, sewage disposal/lavatory, and flushing problems (Fig. 2). can lead to the aerosolization of the virus-infected droplets, as described excellently by a CNN report22. This can lead to further secondary transmission as is evident from the incident at Amoy Gardens, Hong Kong in the 2003 SARS epidemic that resulted in 319 clustered cases19,23. A cycle focusing on the occurrence and fate of infective viruses in wastewater as well as its possible locations that can lead to human exposure has been highlighted by Fig. 3 (adapted after Wigginton et al.24). The virus infected wastewater is transported to the WWTPs, where the untreated wastewater can possibly infect the workers of the plant if bioaerosols are generated. Moreover, a spillover or severe weather event can also lead to the contamination of surface water. Recreational activities like swimming, fishing, kayaking, etc. can lead to further transmission of the virus. The wastewater undergoes a wide variety of physicochemical as well as biological treatment in the WWTPs, leaving behind residual biosolids that are often land-applied23. During the land-application process, there is an additional possibility of human exposure if the viruses manage to survive the treatment. Infected surface water bodies may also serve as sources of drinking water supply. The incoming contaminated surface water undergoes physical and chemical treatment in the drinking water treatment plant before distribution. The manifestation of the viable virus in drinking water is more prominent in cases where the concentration of residual disinfectant is less. The incoming drinking water pipes and outgoing sewage pipes often form an array in the underground. Thus, the leakage of sewage pipes can contaminate the drinking water pipes in the vicinity. This represents the urban water cycle entailing the fate and transport of infective viruses. Additionally, the possibility of fecal-oral transmission of viruses has substantial consequences in those infected areas suffering from poor sanitation and open defecation problem. Currently, 2 billion people of the world do not have access to basic sanitation facilities, of which 673 million still defecate in open areas which include drains and water bodies25.
Reproduced with permission from Wigginton et al.24 (Royal Society of Chemistry, 2015).
The presence of SARS-CoV-2 in the fecal matter of infected patients has been validated by many studies5,15,26. Table 1 summarizes the detection of SARS-CoV-2 in the urine and feces of infected patients in various countries around the globe performed by qPCR. These studies also support the possibility of a fecal-oral mode of transmission thereby strengthening the cycle of transport and occurrence of the virus in water and wastewater as illustrated by Fig. 3. However, the detection of the viral genome or fragments of genetic material in the fecal discharge does not necessarily guarantee the presence of an entire live virus particle that can cause contagion. A disadvantage of the widely used qPCR technique for viral quantification is it can detect viral presence in human samples, but it does not convey any information regarding the level of infectivity of the virus. Further, viruses excreted through fecal matter tend to remain aggregated in the water matrix due to the Brownian movement or nucleation of aquatic particles. This ultimately leads to low recovery rates of virus extraction for biological assays27,28. Nonetheless, recent studies involve attempts in isolation of viable SARS-CoV-2 from the fecal matter of infected patients14, observation of live virion structure from human excrement using electron microscopy29, and identification of live virions from feces30. Thus, currently, it is considered that SARS-CoV-2 may be viable in water, wastewater as well as sewage, with possible fecal-oral transmissibility, though its viral load and level of infection are yet to be researched.
Fate of SARS-CoV-2 in wastewater
Recently many researchers around the world have studied the fate of SARS-CoV-2 in different wastewater matrices as well as river water samples. In one of the first studies in this context, Medema et al. had reported the presence of SARS-CoV-2 in the sewage samples of the Netherlands during the emergence of COVID-19 by detecting two genes of the virus31. Around 58% of the collected samples tested positive for the presence of the viral RNA (nucleic acid detection). Further, the temporal pattern of the detection agreed with the timeline of the emergence of the pandemic in that region. This in turn validated the detection of the virus in the sewage. Rimoldi et al. had examined the presence of viral RNA in raw and treated wastewater samples as well as river water samples of Italy32. A total of 100% of the raw and treated wastewater samples tested positive and negative, respectively for the detection of viral RNA. However, 50% of the river water samples tested positive for the virus, though with no cytopathic effect after 48 and 72 h of inoculation during the vitality test of the virus. The presence of the virus in surface waters could perhaps be attributed to the discharge of untreated wastewater or possible overflow of sewage due to combined sewer overflows (CSOs) of urban runoff and domestic effluents. Combined sewer systems are quite common in central Europe (70%) as well as in 772 cities of the United States of America33,34. Thus, it becomes imperative to strictly monitor urban runoff and domestic effluents during the current pandemic.
To quantify viral load in the environment and initiate wastewater surveillance, researchers have studied the genome concentration of SARS-CoV-2 RNA in various wastewater matrices, like raw wastewater, primary, secondary and tertiary effluents, sludge samples, and river water. In most of the cases, the viral loading was quite high in raw/untreated wastewater with a moderate concentration in treated/secondary/tertiary wastewater. The viral loadings have been reported in terms of copies of viral genomes per liter of the water matrix and/or cycle threshold (Ct) which is the number of cycles needed for the fluorescent signal to exceed the threshold (background levels) during PCR35. Taking Ct into account, most of the recent studies on the presence of the SARS-CoV-2 virus in wastewater matrices are indicative of positive reactions (Ct values in the range of 30–37)31 i.e. moderate presence of viral RNA in the water matrices, though the studies of Arora et al36. and Hata et al37. indicate an abundant presence of the viral RNA (Ct ≤ 29) in wastewater. In another study, Balboa et al. had reported that no genetic material of the virus was detected in the digested sludge samples of a WWTP in Spain which undergo thermal hydrolysis at high temperatures and experience long residence time in anaerobic digesters38. However, the same might not be true for smaller WWTPs in many developing nations where sludges are disposed of by volume reduction methods instead of thermal treatment. Thus, further research is needed in the safe disposal of sludge containing viral RNA, as viruses have an affinity for adsorbing onto biosolids and organic particles, which increases its persistence in the environment. The disinfection scenario of virus-infected wastewater was studied by Zhang et al. in the septic tanks of a hospital in China39. They detected 0.5 × 103 to 1.87 × 104 copies/L of viral genomes in the hospital wastewater even after disinfection with sodium hypochlorite (800 mg/L for 1.5 h). This has sparked the need for the reevaluation of the existing disinfection strategies, especially in non-centralized WWTPs treating SARS-CoV-2 RNA laden effluents. A comprehensive summary of the recent endeavors in the detection and quantification of the SARS-CoV-2 virus in different water matrices has been presented in Tables 2 and 3. Although, numerous studies have examined the genome concentration of SARS-CoV-2 RNA in different water matrices, but only a couple have conducted the survivability test of the virus at that concentration level by inoculation and reported the cytopathic effects which were mostly negative32,40. However, we do not have such cytopathic information for all the studies pertaining to the persistence of the virus in wastewater matrices. Thus, additional research on different aspects of virus vitality and survivability is needed to draw better conclusions on the persistence and level of infection of the virus in waste and surface waters, especially for areas suffering from inadequate sanitation infrastructure.
Effects of environmental factors
Viruses are obligate intracellular parasites: the viral particles are totally dependent on intracellular resources and can reproduce only in the cytoplasm of the hosts’ cells. If viral particles are present in different water matrices by themselves, they may either die or survive in that environment. Thus, unless host cells are present, the viral population will either remain consistent or may decrease with time in that environment. Hence, it is critical to study the influence of various environmental factors that can impact the persistence and survivability of coronavirus. This examination of the intensity of coronavirus persistence by environmental factors can aid in the development of viral inactivation studies.
Temperature
Temperature is the most critical factor that can influence the persistence and survivability of coronavirus in different water matrices and is the most widely studied one (Table 4).
It has been observed that the viral population decreases with an increase in temperature due to protein denaturation and enhanced activity of exoenzymes41,42,43. Wang et al. studied the persistent nature of SARS-CoV (strain BJ01) by collecting stool and urine samples from SARS patients undergoing treatment in the Xiao Tang Shan Hospital, Beijing26. They also collected hospital wastewater and domestic sewage samples from 309th Hospital of PLA (where SARS 2003 patients were being treated) and a housing estate in Beijing, respectively. They studied the influence of temperature at 20 and 4 °C. At 20 °C, the virus was observed to survive in stool and urine for 3 days and at least 17 days, respectively. However, at 4 °C the virus was able to survive in both the samples for more than 17 days. For hospital wastewater, domestic sewage, and dechlorinated water, about 99.9% reduction of the viral population was observed in 2 days at 20 °C. For the same water matrices, the persistence increased to 14 days at 4 °C. Usually, studies use alternative viral surrogates for experiments as the viruses pertinent to human beings cannot be cultivated efficiently and there is a heightened risk factor linked to it. In this respect, the importance of this study lies in the fact that it examined water, urine, and stool samples collected from SARS infected patients during the 2003 SARS outbreak in China.
Casanova et al. examined the survival of two surrogate coronaviruses, transmissible gastroenteritis (TGEV) and mouse hepatitis (MHV) at room temperature (25 °C) and 4 °C for over 6 weeks44. They examined the viral survivability in three types of water matrices: reagent-grade (RG), pasteurized settled sewage (PSS), and lake water (LW). At 25 °C they observed a progressive decline in viral survivability of both TGEV and MHV over a period of 49 days. At 25 °C, a 99% reduction of the TGEV viral population took place in the first 22, 9, and 13 days for RG, PSS, and LW, respectively. Whereas, for MHV at 25 °C, the same level of reduction in viral infectivity was observed in the first 17, 7, and 10 days respectively, for RG, PSS, and LW. At 4 °C, the time for a 99% reduction was predicted using coefficients from regression analysis for both RG and PSS. Accordingly, at 4 °C TGEV could persist in RG and PSS for predicted values of 220 and 49 days, respectively. For similar conditions, the MHV surrogate could survive for more than 365 days and about 70 days in RG and PSS, respectively. For LW at 4 °C, the TGEV viral population weakened by around 1 log10 (90% reduction) by the first 14 days, whereas the MHV viral infectivity persisted with no significant reduction in the viral titer. However, it should be noted that the virus surrogates used in this study, TGEV and MHV cause viral intestinal infection and liver inflammation or damage in animals, respectively. Hence, it might be possible that the resistance exhibited by these viruses in this study might differ from that displayed by human coronaviruses under similar experimental conditions.
The influence of temperature was also investigated by Gundy et al. by studying the persistence of two representative coronaviruses in filtered tap water at 23 and 4 °C45. These viruses included a feline and a human coronavirus: the feline infectious peritonitis virus (FIPV) and the human coronavirus 229E (HcoV-229E). At 23 °C, both HCoV-229E and FIPV could survive in the water matrix for about 10.1 days exhibiting 3 log10 (99.9%) reduction in the viral population. At 4 °C, the predicted values of viral survivability for HCoV-229E and FIPV was estimated to be more than 588 days and 130 days, respectively. Days of viral persistence were usually predicted by modeling for water samples at low temperatures, as it exceeded the scope of the experiments with respect to time constraints. Recently a similar study had also been conducted for the persistence of zika virus in sewage by Muirhead et al.46.
In a relatively recent study, Ye et al. compared the survivability of MHV in both unpasteurized and pasteurized wastewater47. The experiments were performed at 25 and 10 °C which represent the average summer and winter temperatures for wastewater, respectively. It was observed that the viral inactivation kinetics was more rapid at 25 °C than that at 10 °C for both the types of wastewater. For unpasteurized wastewater, the time needed for 2 log10 (99%) reduction of the viral population was 0.5–0.6 days (12–14.4 h) at 25 °C and that at 10 °C was 1.3–1.7 days (31.2–40.8 h). Whereas, for pasteurized wastewater, a 99% reduction of the viral count was observed to take place in 0.5–1.1 days (12–26.4 h) at 25 °C and 2–10.5 days (48–252 h) at 10 °C. It is to be underlined that for both typical summer and winter temperatures, the decrease in the viral count was faster in unpasteurized wastewater than in pasteurized wastewater. This could be attributed to the presence of indigenous microbial population in unpasteurized wastewater which can compete against the coronavirus in wastewater and weaken it. Additionally, Ye and co-workers state that the typical hydraulic residence time of wastewater in sewage systems is around 24 h. Thus, there is always a significant probability that the coronavirus discharged in the sewage system via feces and urine can reach the WWTPs in an infectious state, more so during winter when temperatures drop substantially.
In a recent study, Chin et al. examined the decline in resistance of the SARS-CoV-2 inoculated in virus transport medium from 22 °C (7 days) to 70 °C (1 min), thus highlighting the effect of temperature on its persistence in environment48. Additionally, Ahmed et al. investigated the persistence of gamma-irradiated SARS-CoV-2 (obtained from the Australian Centre for Disease Preparedness) in untreated wastewater, autoclaved wastewater, and dechlorinated tap water at 4 °C (cold), 15 °C (temperate), 25 °C (sub-tropical), and 37 °C (tropical)49. For all the different water matrices, the persistence of SARS-CoV-2 decreased with increase in temperature with the highest being around 60 days for dechlorinated tap water at 4 °C and the lowest being around 5 days for autoclaved wastewater at 37 °C. Additionally, the persistence was lower in untreated wastewater than autoclaved wastewater due to the presence of competitive microorganisms as well as dissolved detergents that might have decayed the SARS-CoV-2 RNA faster.
In general, for all the different water matrices that were tested, the viral population declined more rapidly at room temperature (20–25 °C) than at 4 or 10 °C. The above-discussed studies reassert the fact that temperature is the most important factor as high temperatures lead to the rapid inactivation of enveloped viruses such as the novel coronavirus and cause increased denaturation of the protein present in the virion structure.
Organic matter
The presence of organic matter may impact the survivability of coronavirus in different water matrices (Table 5).
This is because the viruses can get adsorbed onto the organic matter particles. Consequently, the organic matter particles shield the virus from light as well as alter the settling behavior20,43. The effect of organic matter on the survivability of two strains of coronaviruses (HCoV-229E and FIPV) was investigated by Gundy et al.45 For this assay, they tested unfiltered and filtered tap water as well as unfiltered and filtered primary effluent at a constant temperature of 23 °C. By filtering the water samples, they excluded the interference of organic matter particles present in the samples. Additionally, the concentration of organic matter decreases from primary effluent to secondary effluent to tap water. Hence, an additional sample of secondary effluent was included in the assay for easier comparison. It was observed that for both the strains of coronaviruses and different water matrices, the survivability of the virus was more in unfiltered samples than the filtered ones for a 99.9% reduction of the viral population. For HCoV-229E, the persistence of the virus in unfiltered and filtered tap water was 12.1 and 10.1 days, respectively and that in unfiltered and filtered primary effluent was 3.54 and 2.35 days, respectively. A similar trend was also observed for the FIPV strain: the viral persistence was 12.5 and 10.1 days in unfiltered and filtered tap water, respectively and 2.56 and 2.4 days for unfiltered and filtered primary effluent, respectively. Additionally, the persistence of both HCoV-229E and FIPV in secondary effluent was lower (2.77 and 2.42 days, respectively) than that in primary effluent which concurs with the fact that primary effluent had the highest level of organic matter. It can be observed that the persistence of the virus in tap water was about 4 times more than that in primary and secondary effluent despite the tap water having the lowest level of organic matter particles. This can be attributed to the possible presence of solvents and detergents in wastewater which can weaken the viral envelope of coronavirus thereby inactivating it. This study also concluded that the viral envelope of coronavirus is hydrophobic in nature and thus more prone to adsorb on to the organic particles while maintaining less solubility in water.
Ye et al. also studied the adsorption of MHV onto the solid fraction of medium-strength municipal wastewater that had an average total suspended solids of 235 mg/L47. They conducted the experiments at a constant temperature of 4 °C (the range of municipal wastewater temperatures in the USA is 3–27 °C) to minimize the influence of temperature fluctuation on virus survival. They concluded that around 26% of the viruses were observed to be reversibly adsorbed on to the wastewater solids at equilibrium. However, it is to be noted that the equilibrium adsorption percentage of viruses reported by Ye and co-workers does not represent all wastewater types that have varying levels of solid fraction.
Thus, a high concentration of organic matter or solid fraction in water corresponds to increased survivability of the viral population as the organic matter particles physically protect the virus from disinfectants and other antiviral agents. Due to a similar modus operandi, Okoh et al. had concluded that even sediments pose a threat of being reservoirs of viruses in aqueous systems50. Additionally, a study by Gassiloud and Gantzer reported that organic matter particles in water may also compete with the viruses for getting adsorbed on surfaces51. This is critical because such adsorptions are usually physisorption and hence reversible. Due to the reversible nature of this phenomenon, there is a possibility that the viruses might later be discharged from the aqueous system leading to further contagion that is usually not accounted for.
Antagonistic microorganisms
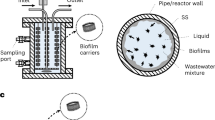
The presence of antagonist microorganisms in the water matrix is yet another factor that can affect the extent of viral inactivation. Naddeo and Liu have suggested that membrane bioreactors (MBRs) which are usually employed in WWTPs can play a crucial role in this respect (Fig. 4)20. Viruses usually remain aggregated in the suspended solids or in the cake that forms on the surface of the membrane with time. Thus, the viruses end up in the company of the indigenous microbial population of the bioreactor which is usually antagonistic in nature. This can lead to the overall decay and inactivation of coronaviruses52,53.
The pasteurization of wastewater also impacts the survivability of coronaviruses. This was highlighted by Ye et al. in their study using pasteurized and unpasteurized wastewater samples47. They pasteurized the wastewater by heating it to 70 °C for 3 h and subsequently tested the infectivity of MHV coronavirus in accordance with protocols established by previous studies27,54. It was observed that MHV lost its infectivity (99% reduction of viral population) at a considerably slower rate in pasteurized wastewater (0.5–1.1 days at 25 °C and 2–10.5 days at 10 °C) compared to unpasteurized wastewater (0.5–0.6 days at 25 °C and 1.3–1.7 days at 10 °C). This could be possibly due to the inactivation of competing or predatory bacteria, protozoans, and metazoans in the pasteurized wastewater samples which would have otherwise competed against the coronaviruses for survival43,55. Additionally, this study highlights the necessity of using unpasteurized wastewater samples for survivability assays of coronavirus if possible.
Disinfectants
A virus maintains its infectivity by performing the following three functions: (i) binding to the host’s cells, (ii) introducing its genome inside the host’s cells, and (iii) replicating itself once the genome is inside the host’s cells. Disinfectants inactivate the virus by inhibiting any of these functions and altering the viral proteins56. Though many studies have investigated the efficiency of disinfectants against coronaviruses on various surfaces, yet studies specific to water matrices are very sparse. Wang and co-workers studied the influence of disinfectant on the survivability of SARS-CoV (BJ01) in wastewater by varying the concentration as well as the time of contact of the disinfectants (chlorine and chlorine dioxide; Table 5)26. They observed that chlorine is more effective than chlorine dioxide in inactivating the virus. Further, for 100% inactivation, the minimum contact time required by 10 and 20 mg/L chlorine was 10 and 1 min, respectively while that required by 40 mg/L of chlorine dioxide was 5 min. A contact time of 10 min for 10 mg/L chlorine dioxide resulted in 68.38% inactivation of the virus. They also calculated the concentration of free residual chlorine in the water matrix after disinfection that can ensure the complete inactivation of the virus, as presented in Table 6.
Overall, chlorine is usually also considered to be the most economic option20. In a recent study, Zhang et al. examined the influent and effluent samples of the septic tank of a COVID-19 hospital in China39. Initially, disinfection was carried out by 800 mg/L of sodium hypochlorite for 1.5 h with residual chlorine more than 6.5 mg/L which was in accordance with the guidelines of China Centers for Disease Control and Prevention (China CDC). However, 12 h later, the free chlorine declined to a nondetectable level and SARS-CoV-2 RNA was detected. Thus, 800 mg/L of sodium hypochlorite was not effective at complete disinfection. Subsequently, a dosage of 6700 mg/L of sodium hypochlorite was added for 1.5 h which resulted in complete disinfection (negative presence of viral RNA) with 21.0–25.0 mg/L of free chlorine after 12 h of disinfection. However, this resulted in higher levels of disinfection by-products residuals in the effluents (like trichloromethane, tribromomethane, bromodichloromethane, and dibromochloromethane) which have substantial environmental risks that need reevaluation.
The reaction mechanism of the inactivation of viruses by disinfectants typically includes a chemical oxidation process. For example, the reaction of chlorine with water leads to the formation of HOCl which further dissociates into hypochlorite (OCl−) and hydrogen (H+) ions:
These highly reactive species, OCl− and HOCl (also termed as free chlorine) create physiological lacerations and penetrate the viral membrane to disrupt protein synthesis. Consequently, this impacts the metabolism processes of the viruses. Similarly, chlorine dioxide undergoes complete oxidation to form chloride ion (Cl−) which is believed to inactivate viruses by interfering with the structure of the membrane proteins57:
The inactivation kinetics of viruses are usually described by the following log-linear relationship:
where, N0 stands for the initial viral population, Nt the viral population at time t, and k the first-order rate constant for inactivation. This relationship yields the survival curve which is employed for the determination of time required to achieve specific levels of inactivation, provided the shape of the curve is indeed log-linear. However, departures from the log-linear shape is quite common: initial shoulder and tailing effect are two such. The former departure is a result of initially stable viral population before the exponential inactivation begins, while the latter corresponding to a decrease in slope results when the last surviving viral population appears to be more resilient (probably due to aggregation or adsorption to organic matter) than the overall population43,58. The tailing effect is better represented by a second-order equation with two rate constants for inactivation. However, the first-order log-linear relationship is quite often used by researchers to summarize the kinetics. This straightforward assumption might result in biased estimates.
Another factor that might hinder the efficacy of viral disinfection is the aggregation of the virus in the medium. A study had observed that only 21% of the viral population existed as isolated particles with the rest forming aggregates of 2–80 particles each59. Thus, it is easier for the disinfectants to reach and inactivate the isolated virus particles than the aggregates.
Ultraviolet exposure
Exposure of coronavirus infected water to ultraviolet (UV; wavelength range 100–400 nm) radiation can lead to the destruction of their nucleic acids and disruption of the DNA and RNA. Viruses are substantially photo-inactivated by processes that involve endogenous direct and exogenous indirect mechanisms. The former process occurs when endogenous chromophores (like nucleic acids and proteins) of the virus absorb photons that lead to modification of the chemical structure of the viral chromophores. The latter mechanism, on the other hand, results when an exogenous chromophore (which is not a part of the virus) absorbs photons and produces photo-produced reactive intermediates (PPRI), which ultimately damage the virus60. Thus, photo-exposure renders the viruses incapable of reproduction. UV-C (100–280 nm) is most effective for germicidal irradiation, though it is most weak on the Earth’s surface as it gets impeded by the ozone layer.
Very limited research is available on the influence of virucidal solar exposure or specifically UV radiation on the survival of coronavirus in aqueous systems. However, Duan and team studied the sensitivity of coronavirus (strain CoV-P9) to UV irradiation by collecting pharyngeal swabs of infected patients during the 2003 SARS pandemic, China61. The isolated viral strain was added to 100 µL of sterilized water and culture medium. It was subsequently exposed to UV irradiation of intensity greater than 90 µW/cm2 with an irradiating distance of 80 cm. Such UV irradiation on the virus medium for 60 min resulted in the destruction of the viral population to an undetectable stage. In another study, Darnell and Taylor examined the effect of UV irradiation on noncellular blood products infected with the Urbani strain of SARS-CoV (obtained from the Centers for Disease Control and Prevention, Atlanta, GA, USA)62. The virus samples were placed 3 cm away from the UV light source. UV-C (254 nm) and UV-A (365 nm) light sources emitting 4016 and 2133 µW/cm2, respectively were used for virucidal inactivation. It was observed that UV-C could efficiently inactive the SARS-CoV virus in 40 min while UV-A when coupled with psoralen compounds (a mutagen that intercalates into the DNA of the virus) could increase the effectiveness of the process.
Currently, limited research exists that examines the influence of UV exposure on aqueous systems infected or spiked with coronavirus. Nonetheless, a review of the existing studies that focus on UV influence on other systems can aid future research developments for water matrices.
pH
The effect of pH on the stability of coronavirus has been addressed by a few studies. According to Daniel and Tablot, MHV is stable over pH ranges of 5–7 and 3–10 at 37 °C and 4 °C, respectively63. Additionally, Pocock and Garwes reported that TGEV remains stable over pH ranges of 5–7 and 5–8 at 37 and 4 °C, respectively64. Casanova et al. observed that the pH declined for MHV with time but did not exceed the pH range of stability44. This suggests that pH (unless it attains very extreme values) might not necessarily be a major factor contributing to the persistence of coronavirus in water.
Future needs
Based on this review of the literature and analysis of current conditions, a multi-pronged approach for future work is suggested:
-
1.
Consideration of updating the existing wastewater infrastructure or the installation of decentralized WWTP for coronavirus hot-spots should be a high priority. Such hot-spots receive hospital and public clinic wastewater which have a significantly high probability of containing viable coronaviruses. A decentralized or in situ treatment facility with an effective septic system, properly sealed floor drains and vent pipes in bathrooms, as well as the use of operational drain traps in flush toilets can reduce the aerosolization of coronavirus infected wastewater65. Such strategies can decrease the environmental loading of the virus as well as secondary transmission in communities.
-
2.
UV based portable devices should be considered for wastewater treatment or for centralized community drinking water treatment to provide water free of the virus. UV based devices for household purposes should be located at the water inlet in conjunction with the water meter, or immediately prior to the location of drinking water outlet(s). This can be followed by a cost-effective chlorine injector at the outlet when the household sewage joins the centralized municipal wastewater lines. This can lead to a community-based approach to inactivate the virus as much as possible and contain its spread.
-
3.
Future research should focus on the development of a possible advanced alarm-based sensing technology that can be used in-line with other systems in WWTPs. Such in situ feedforward systems might include a sensor-based antigen test having a high specificity for viable viral loads in wastewater. This real-time surveillance can lead to early detection of the virus and subsequent rapid intervention before its circulation.
-
4.
Plasma discharge technology should be studied as an alternative to chlorine or other disinfectants. Optimal rates and timings of pulsed high voltages and electrical discharges for treating virus-laden water should be explored in this regard.
-
5.
Open defecation, still practiced in developing countries, should be addressed via public health and extension programs as well as making sanitation facilities accessible to everyone around the globe. For resource-limited geographical areas or coronavirus hot-spots, the construction of pit latrines is a cost-effective alternative. However, lime needs to be added to ensure that such latrines do not pollute the water sources. The vertical distance between the bottom of the pit latrine and the groundwater table is recommended to be a minimum of 1.5 m, while horizontally the pit and the water sources should be separated by a minimum distance of 30 m66. Encouraging residents to utilize facilities not only requires that these facilities themselves be safe and effective at reducing viral loads but should also be a community-led sanitation approach urging all to reduce the practice of open defecation.
-
6.
Research with viruses related to human health (like coronaviruses) has a very high-risk factor associated with it. Thus, further development or cultivation of efficient coronavirus surrogates should be explored to provide a much-needed impetus to its research.
-
7.
In areas where resources are limited, strategies to triage this type of techniques could be further studied. For example, while for individual households, chlorine or UV may be an appropriate technique, treatment of centralized (community) water sources could have positive effects on larger numbers of people. Further studies of combinations of techniques that can most cost-effectively reduce transmission should be considered.
-
8.
Finally, most studies to date have been based on small numbers of patients. Hence, one critical need at this point is to study waterborne and fecal transmission, as well as possible interventions with larger numbers of individuals to enhance the statistical significance of the findings. This would also clarify which techniques may be both biologically relevant and cost-effective in reducing transmission of this and related diseases by waterborne modes.
Conclusions
The probable fecal-oral transmission of SARS-CoV-2 can have serious repercussions on the measures being taken to curb the COVID-19 pandemic. The data-based knowledge reported in many studies and the effect of environmental factors on the survivability of coronaviruses in aqueous environments can aid in risk assessment and planning of sanitation infrastructure. It especially supports mitigation efforts by integrating the urban water cycle with the implications of environmental conditions on the persistence of the virus in water matrices. Apart from adhering to the three vital Ws: washing hands often, wearing a nose and mouth covering to restrict aerosolization of droplets, and waiting 6 ft apart from others, adequate disinfection of potentially virus-laden water and wastewater with best management practices would help us to control this and other future waterborne outbreaks.
References
WHO. Coronavirus disease (COVID-19) situation report – 207. https://www.who.int/docs/default-source/coronaviruse/situation-reports/20200814-covid-19-sitrep-207.pdf?sfvrsn=2f2154e6_2 (2020).
WHO. Naming the coronavirus disease (COVID-19) and the virus that causes it. https://www.who.int/emergencies/diseases/novel-coronavirus-2019/technical-guidance/naming-the-coronavirus-disease-(covid-2019)-and-the-virus-that-causes-it#:~:text=ICTV%20announced%20%E2%80%9Csevere%20acute%20respiratory,the%20two%20viruses%20are%20different (2020).
WHO. WHO Director-General’s remarks at the media briefing on 2019-nCoV on 11 February 2020. https://www.who.int/dg/speeches/detail/who-director-general-s-remarks-at-the-media-briefing-on-2019-ncov-on-11-february-2020 (2020).
Cui, J., Li, F. & Shi, Z. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 17, 181–192 (2018).
Yeo, C., Kaushal, S. & Yeo, D. Enteric involvement of coronaviruses: is faecal–oral transmission of SARS-CoV-2 possible? Lancet Gastroenterol. Hepatol. 5, 335–337 (2020).
Mcintosh, K. Coronaviruses. https://www.uptodate.com/contents/coronaviruses (2020)
La Rosa, G., Bonadonna, L., Lucentini, L., Kenmoe, S. & Suffredini, E. Coronavirus in water environments: Occurrence, persistence and concentration methods - a scoping review. Water Res. 179, 115899 (2020).
Yam, W. C. et al. Evaluation of reverse transcription-PCR assays for rapid diagnosis of severe acute respiratory syndrome associated with a novel coronavirus. J. Clin. Microbiol. 41, 4521–4524 (2003).
Poon, L. L. M. et al. Detection of SARS coronavirus in patients with severe acute respiratory syndrome by conventional and real-time quantitative reverse transcription-PCR assays. Clin. Chem. 50, 67–72 (2004).
Petrich, A. et al. Multicenter comparison of nucleic acid extraction methods for detection of severe acute respiratory syndrome coronavirus RNA in stool specimens. J. Clin. Microbiol. 44, 2681–2688 (2006).
He, Z. et al. Using patient-collected clinical samples and sera to detect and quantify the severe acute respiratory syndrome coronavirus (SARS-CoV). Virol. J. 4, 32 (2007).
Haramoto, E. et al. A review on recent progress in the detection methods and prevalence of human enteric viruses in water. Water Res. 135, 168–186 (2018).
Hamza, I. A. & Bibby, K. Critical issues in application of molecular methods to environmental virology. J. Virol. Methods 266, 11–24 (2019).
Zhang, Y. et al. Isolation of 2019-nCoV from a Stool Specimen of a laboratory-confirmed case of the coronavirus disease 2019 (COVID-19). China CDC Wkly. 2, 123–124 (2020).
Chan, K. H. et al. Detection of SARS Coronavirus in patients with suspected SARS. Emerg. Infect. Dis. 10, 294–299 (2004).
Wang, X. et al. Concentration and detection of SARS coronavirus in sewage from Xiao Tang Shan Hospital and the 309th Hospital. J. Virol. Methods 128, 156–161 (2005).
Watts, J. Report details lessons from SARS outbreak. Lancet 362, 1207 (2003).
Yu, I. T. S. et al. Evidence of airborne transmission of the severe acute respiratory syndrome virus. N. Engl. J. Med. 350, 1731–1739 (2004).
McKinney, K. R., Gong, Y. Y. & Lewis, T. G. Environmental transmission of SARS at Amoy Gardens. J. Environ. Health 68, 26–22 (2006).
Naddeo, V. & Liu, H. Editorial perspectives: 2019 novel coronavirus (SARS-CoV-2): what is its fate in urban water cycle and how can the water research community respond? Environ. Sci. Water Res. Technol. 6, 1213–1216 (2020).
Holshue, M. L. et al. First case of 2019 novel coronavirus in the United States. N. Engl. J. Med. 382, 929–936 (2020).
Regen, H. Experts are investigating if the coronavirus can spread through piping systems. https://edition.cnn.com/asia/live-news/coronavirus-outbreak-02-12-20-intl-hnk/h_f5bafdf84ccb844d1e5b22520f148e74 (2020).
ECDC. Communicable disease threats report week 50, 7–13. https://www.ecdc.europa.eu/en/publications-data/communicable-disease-threats-report-7-13-december-2014-week-50 (2014).
Wigginton, K. R., Ye, Y. & Ellenberg, R. M. Emerging investigators series: the source and fate of pandemic viruses in the urban water cycle. Environ. Sci. Water Res. Technol. 1, 735–746 (2015).
WHO. Sanitation. https://www.who.int/news-room/fact-sheets/detail/sanitation (2019).
Wang, X. et al. Study on the resistance of severe acute respiratory syndrome-associated coronavirus. J. Virol. Methods 126, 171–177 (2005).
Hewitt, J., Greening, G. E., Leonard, M. & Lewis, G. D. Evaluation of human adenovirus and human polyomavirus as indicators of human sewage contamination in the aquatic environment. Water Res. 47, 6750–6761 (2013).
Fumian, T. M. et al. Detection of rotavirus A in sewage samples using multiplex qPCR and an evaluation of the ultracentrifugation and adsorption-elution methods for virus concentration. J. Virol. Methods 170, 42–46 (2010).
Wang, W. et al. Detection of SARS-CoV-2 in different types of clinical specimens. JAMA 323, 1843–1844 (2020).
Xiao, F. et al. Evidence for gastrointestinal infection of SARS-CoV-2. Gastroenterology 158, 1831–1833.e3 (2020).
Medema, G., Heijnen, L., Elsinga, G., Italiaander, R. & Brouwer, A. Presence of SARS-coronavirus-2 RNA in sewage and correlation with reported COVID-19 prevalence in the early stage of the epidemic in the Netherlands. Environ. Sci. Technol. Lett. 7, 511–516 (2020).
Rimoldi, S. G. et al. Presence and infectivity of SARS-CoV-2 virus in wastewaters and rivers. Sci. Total Environ. 744, 140911 (2020).
David Butler, & John W. Davies. Urban Drainage (Spon Press, Taylor & Francis Group, 2004).
United States EPA. What are combined sewer overflows (CSOs)? https://www3.epa.gov/region1/eco/uep/cso.html.
Corpuz, M. V. A. et al. Viruses in wastewater: occurrence, abundance and detection methods. Sci. Total Environ. 745, 140910 (2020).
Arora, S. et al. Sewage surveillance for the presence of SARS-CoV-2 genome as a useful wastewater based epidemiology (WBE) tracking tool in India. Water Sci. Technol. 82, 2823–2836 (2020).
Hata, A., Hara-Yamamura, H., Meuchi, Y., Imai, S. & Honda, R. Detection of SARS-CoV-2 in wastewater in Japan during a COVID-19 outbreak. Sci. Total Environ. 758, 143578 (2020).
Balboa, S. et al. The fate of SARS-COV-2 in WWTPs points out the sludge line as a suitable spot for monitoring. Preprint at https://www.medrxiv.org/content/10.1101/2020.05.25.20112706v1 (2020).
Zhang, D. et al. Potential spreading risks and disinfection challenges of medical wastewater by the presence of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) viral RNA in septic tanks of Fangcang Hospital. Sci. Total Environ. 741, 140445 (2020).
Wang, J. et al. SARS-CoV-2 RNA detection of hospital isolation wards hygiene monitoring during the Coronavirus Disease 2019 outbreak in a Chinese hospital. Int. J. Infect. Dis. 94, 103–106 (2020).
Hurst, C. J., Gerba, C. P. & Cech, I. Effects of environmental variables and soil characteristics on virus survival in soil. Appl. Environ. Microbiol. 40, 1067–1079 (1980).
John, D. E. & Rose, J. B. Review of Factors Affecting microbial survival in groundwater. Environ. Sci. Technol. 39, 7345–7356 (2005).
Pinon, A. & Vialette, M. Survival of viruses in water. Intervirol 61, 214–222 (2019).
Casanova, L., Rutala, W. A., Weber, D. J. & Sobsey, M. D. Survival of surrogate coronaviruses in water. Water Res. 43, 1893–1898 (2009).
Gundy, P. M., Gerba, C. P. & Pepper, I. L. Survival of coronaviruses in water and wastewater. Food Environ. Virol. 1, 10–14 (2008).
Muirhead, A. et al. Zika virus RNA persistence in sewage. Environ. Sci. Technol. Lett. 7, 659–664 (2020).
Ye, Y., Ellenberg, R. M., Graham, K. E. & Wigginton, K. R. Survivability, partitioning, and recovery of enveloped viruses in untreated municipal wastewater. Environ. Sci. Technol. 50, 5077–5085 (2016).
Chin, A. W. H. et al. Stability of SARS-CoV-2 in different environmental conditions. Lancet Microbe 1, e10 (2020).
Ahmed, W. et al. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: a proof of concept for the wastewater surveillance of COVID-19 in the community. Sci. Total Environ. 728, 138764 (2020).
Okoh, A. I., Sibanda, T. & Gusha, S. S. Inadequately treated wastewater as a source of human enteric viruses in the environment. Int. J. Environ. Res. Public Health 7, 2620–2637 (2010).
Gassilloud, B. & Gantzer, C. Adhesion-aggregation and inactivation of poliovirus 1 in groundwater stored in a hydrophobic container. Appl. Environ. Microbiol. 7, 912–920 (2005).
Bodzek, M., Konieczny, K. & Rajca, M. Membranes in water and wastewater disinfection– review. Arch. Environ. Prot. 45, 3–18 (2019).
Chaudhry, R. M., Nelson, K. L. & Drewes, J. E. Mechanisms of pathogenic virus removal in a full-scale membrane bioreactor. Environ. Sci. Technol. 49, 2815–2822 (2015).
Casanova, L. M. & Weaver, S. R. Inactivation of an enveloped surrogate virus in human sewage. Environ. Sci. Technol. Lett. 2, 76–78 (2015).
Rzeżutka, A. & Cook, N. Survival of human enteric viruses in the environment and food. FEMS Microbiol. Rev. 28, 441–453 (2004).
Wigginton, K. R., Pecson, B. M., Sigstam, T., Bosshard, F. & Kohn, T. Virus Inactivation mechanisms: impact of disinfectants on virus function and structural integrity. Environ. Sci. Technol. 46, 12069–12078 (2012).
LeChevallier, M. W. & Au, K. K. Water Treatment and Pathogen Control (Iwa Publishing, London, UK, 2004).
Hiatt, C. W. Kinetics of the inactivation of viruses. Bacteriol. Rev. 28, 150–163 (1964).
Galasso. Effect of particle aggregation on the survival of irradiated vaccinia virus. J. Bacteriol. 90, 1138–1142 (1965).
Nelson, K. L. et al. Sunlight-mediated inactivation of health-relevant microorganisms in water: a review of mechanisms and modeling approaches. Environ. Sci. —Process. Impacts 20, 1089–1122 (2018).
Duan, S. et al. Stability of SARS coronavirus in human specimens and environment and its sensitivity to heating and UV irradiation. Biomed. Environ. Sci. 16, 246 (2003).
Darnell, M. R. & Taylor, D. Evaluation of inactivation methods for severe acute respiratory syndrome coronavirus in noncellular blood products. Transfusion 46, 1770–1777 (2006).
Daniel, C. & Talbot, P. J. Physico-chemical properties of murine hepatitis virus, strain A59. Arch. Virol. 96, 241–248 (1987).
Pocock, D. H. & Garwes, D. J. The influence of pH on the growth and stability of transmissible gastroenteritis virus in vitro. Arch. Virol. 49, 239–247 (1975).
Gormley, M., Aspray, T. J. & Kelly, D. A. COVID-19: mitigating transmission via wastewater plumbing systems. Lancet Glob. Health 8, e643 (2020).
Bhowmick, G. D. et al. Coronavirus disease 2019 (COVID-19) outbreak: some serious consequences with urban and rural water cycle. NPJ Clean. Water 3, 1–8 (2020).
Cai, J. et al. A case series of children with 2019 novel coronavirus infection: clinical and epidemiological features. Clin. Infect. Dis. 71, 1547–1551 (2020).
Chan, J. F. et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet 395, 514–523 (2020).
Jin, X. et al. Epidemiological, clinical and virological characteristics of 74 cases of coronavirus-infected disease 2019 (COVID-19) with gastrointestinal symptoms. Gut 69, 1002–1009 (2020).
Lescure, F. et al. Clinical and virological data of the first cases of COVID-19 in Europe: a case series. Lancet Infect. Dis. 20, 697–706 (2020).
Kam, K. et al. A well infant with coronavirus disease 2019 with high viral load. Clin. Infect. Dis. 71, 847–849 (2020).
Lo, I. L. et al. Evaluation of SARS-CoV-2 RNA shedding in clinical specimens and clinical characteristics of 10 patients with COVID-19 in Macau. Int. J. Biol. Sci. 16, 1698–1707 (2020).
Ling, Y. et al. Persistence and clearance of viral RNA in 2019 novel coronavirus disease rehabilitation patients. Chin. Med. J. 133, 1039–1043 (2020).
Pan, Y., Zhang, D., Yang, P., Poon, L. L. M. & Wang, Q. Viral load of SARS-CoV-2 in clinical samples. Lancet Infect. Dis. 20, 411–412 (2020).
Wang, Y., Kang, H., Liu, X. & Tong, Z. Asymptomatic cases with SARS‐CoV‐2 infection. J. Med. Virol. 92, 1401–1403 (2020).
Young, B. E. et al. Epidemiologic features and clinical course of patients infected with SARS-CoV-2 in Singapore. JAMA 323, 1488–1494 (2020).
Tang, A. et al. Detection of novel coronavirus by RT-PCR in stool specimen from asymptomatic child, China. Emerg. Infect. Dis. 26, 1337–1339 (2020).
Kujawski, S. A. et al. Clinical and virologic characteristics of the first 12 patients with coronavirus disease 2019 (COVID-19) in the United States. Nat. Med. 26, 861–868 (2020).
Wu, Y. et al. Prolonged presence of SARS-CoV-2 viral RNA in faecal samples. Lancet Gastroenterol. Hepatol. 5, 434–435 (2020).
Xie, C. et al. Comparison of different samples for 2019 novel coronavirus detection by nucleic acid amplification tests. Int. J. Infect. Dis. 93, 264–267 (2020).
Xing, Y. et al. Prolonged presence of SARS-CoV-2 in feces of pediatric patients during the convalescent phase. J. Microbiol. Immunol. Infect. 53, 473–480 (2020).
Zhang, N. et al. Comparative study on virus shedding patterns in nasopharyngeal and fecal specimens of COVID-19 patients. Sci. China Life Sci. 1–3 (2020).
Bar-Or, I. et al. Regressing SARS-CoV-2 sewage measurements onto COVID-19 burden in the population: a proof-of-concept for quantitative environmental surveillance. Preprint at https://www.medrxiv.org/content/10.1101/2020.04.26.20073569v1 (2020).
Green, H. et al. Quantification of SARS-CoV-2 and cross-assembly phage (crAssphage) from wastewater to monitor coronavirus transmission within communities. Preprint at https://www.medrxiv.org/content/10.1101/2020.05.21.20109181v1 (2020).
Haramoto, E. et al. First environmental surveillance for the presence of SARS-CoV-2 RNA in wastewater and river water in Japan. Sci. Total Environ. 737, 140405 (2020).
La Rosa, G. et al. First detection of SARS-CoV-2 in untreated wastewaters in Italy. Sci. Total Environ. 736, 139652 (2020).
Kumar, M. et al. First proof of the capability of wastewater surveillance for COVID-19 in India through detection of genetic material of SARS-CoV-2. Sci. Total Environ. 746, 141326 (2020).
Kocamemi, B. A. et al. First data-set on SARS-CoV-2 detection for Istanbul wastewaters in Turkey. Preprint at https://www.medrxiv.org/content/10.1101/2020.05.03.20089417v1 (2020).
Kocamemi, B. A. et al. SARS-CoV-2 detection in Istanbul wastewater treatment plant sludges. Preprint at https://www.medrxiv.org/content/10.1101/2020.05.12.20099358v1 (2020).
Nemudryi, A. et al. Temporal detection and phylogenetic assessment of SARS-CoV-2 in municipal wastewater. Cell Rep. Med. 100098 (2020).
Peccia, J. et al. Measurement of SARS-CoV-2 RNA in wastewater tracks community infection dynamics. Nat. Biotech. 38, 1164–1167 (2020).
Randazzo, W. et al. SARS-CoV-2 RNA in wastewater anticipated COVID-19 occurrence in a low prevalence area. Water Res. 181, 115942 (2020).
Sharif, S. et al. Detection of SARs-CoV-2 in wastewater, using the existing environmental surveillance network: an epidemiological gateway to an early warning for COVID-19 in communities. Preprint at https://www.medrxiv.org/content/10.1101/2020.06.03.20121426v3 (2020).
Wurtzer, S. et al. Evaluation of lockdown impact on SARS-CoV-2 dynamics through viral genome quantification in wastewater, Greater Paris, France, 5 March to 23 April 2020. Euro Surveill. 25, 2000776 (2020).
Wu, F. et al. SARS-CoV-2 titers in wastewater are higher than expected from clinically confirmed cases. mSystems. 5, 00614–20 (2020).
Acknowledgements
The funding received from the Department of Biological and Agricultural Engineering, North Carolina State University, William White Endowment of the North Carolina Agricultural Foundation, and the College of Agriculture and Life Sciences of North Carolina State University is duly acknowledged.
Author information
Authors and Affiliations
Contributions
D.P.: Conceptualization, Data analysis, Editing, Writing. P.K.: Conceptualization, Supervision, Editing, Writing. S.G.H.: Conceptualization, Funding Acquisition, Supervision, Editing, Writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Paul, D., Kolar, P. & Hall, S.G. A review of the impact of environmental factors on the fate and transport of coronaviruses in aqueous environments. npj Clean Water 4, 7 (2021). https://doi.org/10.1038/s41545-020-00096-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41545-020-00096-w
This article is cited by
-
The response of aquatic ecosystems to the interactive effects of stratospheric ozone depletion, UV radiation, and climate change
Photochemical & Photobiological Sciences (2023)
-
Environmental impacts of the widespread use of chlorine-based disinfectants during the COVID-19 pandemic
Environmental Science and Pollution Research (2022)
-
Study on infection behavior and characteristics of poplar wood dyed by Lasiodiplodia theobromae
European Journal of Wood and Wood Products (2022)