Abstract
Soluble HIV-1 envelope glycoprotein (Env) immunogens are a prime constituent of candidate vaccines designed to induce broadly neutralizing antibodies. Several lines of evidence suggest that enhancing Env immunogen thermostability can improve neutralizing antibody (NAb) responses. Here, we generated BG505 SOSIP.v9 trimers, which displayed virtually no reactivity with non-neutralizing antibodies and showed increased global and epitope thermostability, compared to previous BG505 SOSIP versions. Chemical crosslinking of BG505 SOSIP.v9 further increased the melting temperature to 91.3 °C, which is almost 25 °C higher than that of the prototype SOSIP.664 trimer. Next, we compared the immunogenicity of a palette of BG505-based SOSIP trimers with a gradient of thermostabilities in rabbits. We also included SOSIP.v9 proteins in which a strain-specific immunodominant epitope was masked by glycans to redirect the NAb response to other subdominant epitopes. We found that increased trimer thermostability correlated with increased potency and consistency of the autologous NAb response. Furthermore, glycan masking steered the NAb response to subdominant epitopes without decreasing the potency of the autologous NAb response. In summary, SOSIP.v9 trimers and their glycan masked versions represent an improved platform for HIV-1 Env based vaccination strategies.
Similar content being viewed by others
Introduction
One of the main goals of the HIV-1 vaccine field has been the design of soluble envelope glycoprotein (Env)-based immunogens that can induce protective neutralizing antibody (NAb) responses1,2,3,4. However, several intrinsic features of Env result in immune evasion by hindering the development of NAb responses4. For instance, the instability and conformational flexibility of the Env complex results in the exposure of immunodominant non-neutralizing antibody (non-NAb) epitopes that have been traditionally hypothesized to act as immune decoys5. These non-neutralizing epitopes include the immunodominant variable region 3 (V3), which is normally hidden inside the closed structure of native Env6,7, the inner domain of gp1208,9 and part of gp41. Furthermore, much of the protein surface that is exposed on native Env is either hypervariable or covered by a dense glycan shield10,11,12.
Several advances, exemplified by the BG505 SOSIP.664 prototype, allow for the routine generation of soluble Env trimers that closely mimic the prefusion conformation of viral Env6,13,14,15,16,17,18,19. The SOSIP.664 design contains an optimized furin cleavage motif (R6), an Ile-559-Pro (IP) substitution that stabilizes Env in its prefusion trimeric conformation and a disulfide bond (SOS) between positions 501 and 605 that covalently links the gp120 and gp41 subunits13,20,21,22. Incidentally, the “IP” substitution was, in a modified form, also applied to stabilized SARS-CoV-2 Spike protein in many COVID vaccines, termed “2P”23,24. Contrary to gp120 and non-native-like trimers, SOSIP.664 and other first-generation native-like trimers can induce NAb responses against autologous neutralization-resistant (Tier 2) viruses6,25,26. However, they still expose several non-NAb epitopes and induce significant V3-directed responses. Thus, a significant number of studies have focused on improving the performance of Env immunogens by immunosilencing undesired epitopes and reducing their conformational flexibility4,27,28. The stability derived from the reduced conformational flexibility might also result in a prolonged antigen presentation in the prefusion native-like conformation and thus increase the probability of inducing the desired NAb responses. Furthermore, more stable immunogens may facilitate a longer shelf-life and require less stringent cold-chain conditions, thus allowing their deployment under real-world conditions29.
Several stability-enhancing modifications to the BG505 SOSIP.664 prototype have been described. BG505 DS-SOSIP was generated by including an intra-gp120 disulfide bond (201C-433C) to prevent CD4-induced (CD4i) rearrangements that lead to non-NAb epitope exposure30,31. For the design of SOSIP.v4 trimers, we included two mutations (64K/66R and 316W), at positions neighboring the V3 loop, which resulted in reduced exposure of CD4i and V3 non-NAb epitopes and increased thermostability32. The MD39 set of mutations, discovered by directed molecular evolution, stabilize the core of Env trimers and result in improved trimerization and reduced V3 reactivity33. We further stabilized the prefusion sequestered conformation of the V3 loop by hydrophobic interactions induced by a 306L-308L pair of mutations34. Furthermore, we designed SOSIP.v6 trimers by introducing additional intra- (73C-561C) and interprotomer (49C-555C) disulfide bonds. SOSIP.v6 trimers showed significantly increased melting temperatures and induced strong autologous tier 2 NAb responses35. Chemical crosslinking with glutaraldehyde (GLA) and EDC/NHS has also been used to improve the half-life of Env immunogens and reduce their conformational flexibility. Crosslinked trimers induce more potent NAb responses than their non-crosslinked counterparts, indicating that higher stability is related to improved immunogenicity36,37,38.
The extensive glycosylation of the Env protein provides protection from NAbs targeting conserved regions12,39,40. Holes in this glycan shield expose strain-specific protein segments that lure the immune system into generating narrow-neutralizing antibody responses that might compete with responses targeting other, potentially cross-neutralizing, epitopes41,42,43,44. For instance, the strain-specific glycan hole around positions 241 and 289 in BG505 SOSIP trimers is immunodominant in rabbits and to a lesser extent in macaques42,45,46,47. The NAb responses directed to these glycan holes are of narrow specificity and unlikely to develop breadth48. They likely represent dead-end pathways on the route towards broadly neutralizing antibodies (bNAbs) and possibly compete with potential bNAb responses49. However, it has been shown that 241/289 glycan hole responses can be redirected by opening up other glycan holes in BG505 SOSIP trimers50.
Here, we combined several stabilizing mutations to generate a set of BG505 SOSIP trimers, named SOSIP.v9.1-v9.4, that display a high thermal stability and low reactivity to non-NAbs. Next, we immunized rabbits with a palette of BG505-based SOSIP trimers with a gradient of stabilities to determine whether greater immunogen stability improves the magnitude and consistency of the NAb response. Furthermore, we also immunized rabbits with ultrastable SOSIP.v9 trimers in which the 241/289 glycan hole was masked. We observed that this glycan masking efficiently redirected the NAb responses from the immunodominant 241/289 glycan hole to a more subdominant epitope (C3/465 region) without negatively affecting the magnitude of the NAb response. These ultrastable immunogens provide an improved design platform for epitope-focused vaccine strategies.
Results
Design and generation of hyperstable BG505 SOSIP trimers
We previously described the BG505 SOSIP.v6 immunogen35, a thermostable SOSIP trimer that contained additional inter- and intraprotomeric disulfide bonds and folded as a covalent trimer with a remarkably high thermostability. However, SOSIP.v6 still showed some reactivity to non-NAbs and was not produced as efficiently as preceding SOSIP versions. Our initial goal here was to generate hyperstable SOSIP trimers that do not present these drawbacks. Thus, we selected a set of mutations that have been shown to improve the antigenicity, stability, trimerization, and purification yields of HIV-1 Env trimers31,33,34,35 and introduced them in the BG505 SOSIP.v6 design in different combinations to generate BG505 SOSIP.v9.1-v9.4 (Fig. 1a, Fig. S1). A subset of the MD39 mutations (519S, 568D, 570H, and 585H)33 were introduced in all four SOSIP.v9.1 -v9.4 trimers in order to improve trimerization and purification yields. Newly incorporated mutations 306L-308L34, 304V and 319Y33, as well as the 316W mutation previously introduced in SOSIP.v432, sequester the immunodominant V3 loop region by hydrophobic interactions. Due to the proximity of these mutations in the three-dimensional structure of the trimer (Fig. 1b), we included them in three different combinations (316W+306L-308L in SOSIP.v9.1 and SOSIP.9.4, 304V+319Y in SOSIP.v9.2 and 316W+304V+319Y in SOSIP.v9.3) to avoid undesired interactions or interferences. The disulfide bond 201C-433C, which reduces sampling of the CD4-induced state31, was introduced in SOSIP.v9.4 only.
a Table showing the combinations of stabilizing mutations used for the generation of SOSIP.v9 trimers. SOSIP.v5 mutations refer to 64K, 315Q, 501C-605C, R6, 535M, 543N, 559P and the truncation after residue 66413,32. b Location of the newly incorporated mutations in the three-dimensional structure of a BG505 SOSIP trimer. Mutations appear colored according to the color code in Fig. 1A. c SDS-PAGE analysis of PGT145-purified SOSIP.v5, SOSIP.v6 and SOSIP.v9 trimers under non-reducing (-DTT) and reducing (+DTT) conditions. d 2D class averages generated by negative-stain electron microscopy analysis of PGT145-purified SOSIP.v9 trimers. The percentage of native-like and non-native-like trimers are showed in green and red, respectively. e Ni-NTA-capture enzyme-linked immunosorbent assay with PGT145-purified SOSIP.v9 and the control (SOSIP.v5 and SOSIP.v6) proteins against a panel of bNAbs and non-NAbs. The binding index represents the average of duplicate measures and corresponds to the ratio between the 2G12 (loading control antibody)-normalized area under the curve (AUC) values of each protein and the 2G12-normalized AUC of the SOSIP.v5 reference protein. The bars represent the average of the binding indexes calculated for all the bNAbs (light green) and non-NAbs (light red).
We first purified His-tagged versions of BG505 SOSIP.v9.1-v9.4 expressed in HEK293F suspension cells by PGT145 affinity chromatography, as previously described32,51. All SOSIP.v9.1-v9.4 proteins showed increased purification yields compared to their SOSIP.v6 predecessor, with a 3.8 - 4.3-fold increase in the case of SOSIP.v9.1-v9.3 and a more subtle but consistent ~1.3-fold increase for SOSIP.v9.4 (Table 1). These yields were ~1.5-fold higher than those of non-hyperstabilized SOSIP.v4 and SOSIP.v5 trimers31. As expected, after PGT145 purification, all SOSIP versions tested formed trimers, as assessed by the BN-PAGE analysis (Fig. S2). An SDS-PAGE analysis showed that all SOSIP.v9.1-v9.4 purified proteins remain trimeric under non-reducing conditions, while they separate into monomers under reducing conditions (Fig. 1c). This confirms that SOSIP.v9 trimers contain interprotomer disulfide bonds that covalently link the three protomers, similarly to the SOSIP.v6 predecessor35. Furthermore, 2D class averages of negative-stain electron microscopy (NS-EM) images revealed that all four SOSIP.v9 variants assumed a >95% native-like conformation (Fig. 1d, Table 1). A Ni-NTA ELISA experiment with the purified proteins revealed a favorable antigenic profile for all the SOSIP.v9 trimers compared to SOSIP.v5 and SOSIP.v6 (Fig. 1e, Fig. S3), with similar (or higher) binding to bNAbs and decreased binding to non-NAbs. BG505 SOSIP.v9.3 showed the most desirable antigenic profile, with increased binding to all the bNAbs tested, in particular the quaternary-dependent bNAbs PG16, PGT145, and VRC026.25, and decreased binding to all non-NAbs tested (17b + sCD4, 14e, 19b, and F105) (Fig. 1e, Fig. S3). In summary, the four BG505 SOSIP.v9 proteins fold as covalent native-like Env trimers and are improved compared to SOSIP.v6 in terms of production yields and antigenicity. Overall, the SOSIP.v9.3 construct displayed the most promising features, especially regarding its antigenicity profile.
SOSIP.v9 trimers show superior global and epitope-specific thermostability
In order to evaluate the influence of the stabilizing mutations incorporated on the thermostability of SOSIP.v9 trimers, we used Nano Differential Scanning Fluorimetry (nanoDSF), which provides information on the overall thermostability, but does not reveal how stable individual bNAb epitopes are. The SOSIP.v9.1-v9.4 trimers presented superior global thermostability compared to all their precursors, including the hyperstable SOSIP.v6 (Fig. 2a, Table 1). The SOSIP.v9.3 and SOSIP.v9.4 proteins displayed the highest melting temperatures (Tm) among the SOSIP.v9 variants, with Tm values of 80.7 °C and 84 °C, respectively (Fig. 2d, Table 1). These Tm values are 13.1 °C and 16.4 °C higher than the ones of first-generation BG505 SOSIP.664 trimers35.
a–c Data used to determine the temperature of melting (Tm) values of PGT145-purifed SOSIP.v9 trimers by nanoDSF (a), 2G12 thermostability ELISA (b) and PGT145 thermostability ELISA (c) assays. Left graphs show the normalized fluorescence ratio (F350/F330), 2G12 binding and PGT145 binding signals measured at different temperatures. Graphs on the right show the first derivative of the measured signals. Dots represent the Tm value of each protein, calculated as the temperature at which the signal is half of the maximum signal (left) or the value of the first derivative is maximized (right). d Temperatures of melting calculated for each protein and thermostability assay, ordered from lower to higher values. e Correlation between Tm values determined using the different thermostability assays. Pearson r and p-values are indicated for each pair of assays.
To gauge the thermostability of specific bNAb epitopes, we performed thermostability ELISA assays with bNAb 2G12 (Fig. 2b), which depends on conformational structure but not on quaternary structure, and PGT145 (Fig. 2c), which depends on proper quaternary conformation at the trimer apex. All SOSIP.v9 trimers displayed more stable 2G12 and PGT145 epitopes than the previous SOSIP versions (Fig. 2b–d, Table 1). The Tm values of the 2G12 and PGT145 epitopes of SOSIP.v9.3 (78.3 °C and 76.1 °C, respectively) and SOSIP.v9.4 (85.0 °C and 82.0 °C, respectively) were the highest among the SOSIP.v9 proteins.
The results of all three techniques (nanoDSF, 2G12, and PGT145 thermostability ELISA) were strongly correlated (Fig. 2e, Table 1). The Tm values measured by nanoDSF were ~2 °C higher than the ones measured by 2G12 thermostability ELISA, and ~5 °C higher than those measured by PGT145 thermostability ELISA. These trends are expected, as trimer dissociation probably precedes loss of gp120 conformation and, finally, unfolding.
Chemical crosslinking further stabilizes SOSIP.v9.3
To add an additional layer of stabilization, we used glutaraldehyde (GLA) to chemically crosslink the SOSIP.v9.3 immunogen (SOSIP.v9.3.XL)37. After purification of the SOSIP.v9.3.XL protein by PGT151 affinity chromatography, we visualized it by NS-EM, confirming that >95% of the species presented a native-like conformation (Fig. S4). Trimer antigenicity was evaluated in a BLI experiment with a small panel of relevant Abs (Fig. S5). Chemically crosslinked and PGT151-purified SOSIP.v9.3 showed similar 2G12 and PGT151 binding compared to normal SOSIP.v9.3, whereas PGT145 and VRC01 binding was decreased, consistent with previous crosslinking experiments37,52. The decrease in PGT145 binding has previously been hypothesized to be related to the direct modification of the lysine residues 168, 169, and 171, rather than to a loss of native-like apex conformation37. Notably, chemical crosslinking increased the Tm of SOSIP.v9.3 from 80.7 °C to 91.3 °C, which is an increase of 23.7 °C compared to the SOSIP.664 prototype35 (Fig. 2d, Table 1). Thus, we generated a panel of SOSIP trimers with a gradient of stabilities (SOSIP.v9.3.XL > SOSIP.v9.4 > SOSIP.v9.3 > SOSIP.v9.2 ≃ SOSIP.v9.1 > SOSIP.v6 > SOSIP.v5 > SOSIP.v4 > SOSIP.664) (Fig. 2d, Table 1).
Addition of PNGS at position 241 and 289 fills the 241/289 glycan hole on SOSIP.v9 trimers
Considering the immunodominance of the N241/N289 BG505 glycan hole, we aimed to immunosilence this epitope and redirect the NAb responses towards other, potentially non-strain-specific, epitopes. Thus, we created Glycan hole Masked (GM) versions of the SOSIP.v9.3 and SOSIP.v9.4 trimers (SOSIP.v9.3.GM and SOSIP.v9.4.GM) by introducing PNGS at positions 241 and 289 (241N, 291S) together with changes that counteract the destabilizing effects result of the incorporation of such PNGS motifs (240T, 271I, 288L, 290E)45 (Fig. 3a).
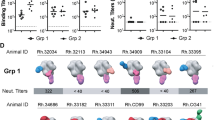
a Representation of the glycan shield of non-glycan masked (non-GM), containing the 241/289 glycan hole (left), and glycan-masked (GM) SOSIP.v9 trimers, which present an intact glycan shield in the 241/289 region (right). Modeling of the glycan shields was performed using the Glycan Shield Mapping tool41. b 2D class averages obtained by negative-stain electron microscopy analysis of PGT145-purified SOSIP.v9 glycan masked trimers. The percentages of native-like and non-native-like trimers are indicated in green and red, respectively. c Reactivity of the 241/289 targeting mAb 10A42 against non-GM and GM SOSIP.v9 proteins measured by a Ni-NTA-capture ELISA. The binding index for each tested protein represents the values of duplicate measures and corresponds to the ratio between the 2G12 (loading control antibody)-normalized area under the curve (AUC) values of each protein and the 2G12-normalized AUC of the SOSIP.v5 reference protein. The bars and error bars represent the average and standard deviation of the ratios obtained for each of the experiments. d Site-specific glycan analysis of PGT145-purified SOSIP.v9.3.GM protein.
We expressed the His-Tagged versions of SOSIP.v9.3.GM and SOSIP.v9.4.GM in 293 F suspension cells and purified them by PGT145 affinity chromatography. SOSIP.v9.3.GM and SOSIP.v9.4.GM showed slightly decreased purification yields (2.4 mg/L and 0.6 mg/L, respectively) as compared to the respective parental proteins (Table 1). They also expressed as covalently linked trimers, as observed on SDS-PAGE (Fig. S6). When imaged by NS-EM, we observed a reduction in the percentage of native-like trimers of SOSIP.v9.4.GM ( > 85% vs >95%), while SOSIP.v9.3.GM conserved a >95% native-like conformation (Fig. 3b, Table 1). Site-specific glycan analysis revealed that PNGS occupancy and glycan composition was similar for SOSIP.v9.3.GM (Fig. 3d, Fig. S7) when compared to previous SOSIP versions. The occupancy at the newly incorporated 241 and 289 PNGS was 90% and 66%, respectively, thus effectively filling the hole with at least one glycan in the majority of molecules (Fig. S7).
Both GM trimers showed an overall similar antigenic profile as SOSIP.v9.3 and SOSIP.v9.4 (Fig. S8). Furthermore, an antibody targeting the 241/289 glycan hole of BG505 (10 A)42 showed negligible reactivity to GM trimers, confirming the effective filling of the 241/289 hole (Fig. 3d).
The inclusion of new PNGS slightly influenced thermostability, with nanoDSF-measured Tm values of 79.9 °C and 83.2 °C for SOSIP.v9.3.GM and SOSIP.v9.4.GM, respectively (i.e. 0.8 °C lower compared to the parental proteins) (Fig. S9).
Ultrastable SOSIP.v9 trimers induce autologous NAb responses
To evaluate the immunogenicity of the SOSIP.v9 trimers and gauge their ability to generate NAb responses, we performed an immunization experiment in rabbits (Fig. 4a). We selected the following immunogens to be tested: (1) the BG505 SOSIP.v5 and SOSIP.v6 predecessors, to serve as control groups; (2) BG505 SOSIP.v9.3 and SOSIP.v9.4, as they showed the best antigenic profile and thermostability, respectively, among the SOSIP.v9 trimers; (3) BG505 SOSIP.v9.3.XL trimer, i.e. the trimer with the highest thermostability; and 4) glycan masked versions of BG505 SOSIP.v9.3 and SOSIP.v9.4 (SOSIP.v9.3.GM and SOSIP.v9.4.GM), to study redirection of responses away from the 241/289 hole. Untagged versions of these immunogens were purified by PGT145 affinity chromatography and an additional size exclusion chromatography (SEC) purification step to remove aggregated species. BN- and SDS-PAGE analyses were performed for quality control (Fig. S10).
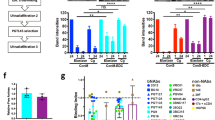
a Rabbits were immunized with different BG505 SOSIP trimer versions adjuvanted with squalene emulsion at weeks 0, 4 and 20. Antibody responses were evaluated at weeks 0, 4, 6, 16, 20, and 22. b Endpoint antibody binding titers over time against BG505 SOSIP.v5 trimer as measured by Ni-NTA ELISA. The mean binding titers and standard deviation of each group are shown. No significant differences were found by a Kruskal–Wallis statistical test between groups at any timepoint tested. c Midpoint neutralization titers (ID50) over time for sera of the immunized rabbits against a virus pseudotyped with a cytoplasmatic tail truncated BG505 Env (BG505.dCT). ID50 values were determined by TZM-bl assay and are detailed in Fig. S13. The geometric mean binding titers and geometric standard deviation of each group are represented d Correlation between the melting temperature of immunogens and the waning of autologous neutralization responses between weeks 6 and 20. The waning index corresponds to the difference between the ID50 values at weeks 6 and 20. The dotted line separates groups that show a reduction (positive waning values) or increase (negative waning values) in titers between these timepoints. Pearson test r and p correlation values are presented. e Midpoint neutralization titers (ID50) for week 22 sera of the immunized rabbits against an autologous BG505.dCT pseudovirus. Geometric means and geometric standard deviations are represented. f–g Effect of the ultrastabilization on the potency and consistency of autologous NAb responses. Correlation plots between geometric mean (Geomean) (f) and geometric standard deviation (GSD) (g) of the ID50 against the BG505 autologous pseudovirus and the melting temperatures (Tm) of the immunogens, determined by nanoDSF. Spearman r and p-values are presented. h–i Effect of the 241/289 glycan hole masking on the autologous NAb responses. h Comparison of the neutralization titers elicited by non-glycan masked (non-GM) and glycan masked (GM) SOSIP.v9.3 and SOSIP.v9.4 immunogens against autologous BG505/T332N and Tier 1A MW965.26 pseudoviruses. ID50 values were determined in a TZM-bl assay and are presented in Fig. S15. No significant differences (N.S.) were found by a Mann–Whitney statistical test. i Reduction of the NAb titers against a panel of mutant (241/289N, 133aN/136aA, and 465N) BG505 pseudoviruses compared to the wild-type BG505/T332N pseudovirus. Relative ID50 (RID50) is calculated as the ratio between the ID50 against each mutant virus and the ID50 of the wild-type virus. Horizontal lines and error bars represent the geometric mean and geometric standard deviations. A dotted line is used to indicate the RID50 value for which there is no reduction or increase of ID50 values for the mutants.
Seven groups of animals (5 animals per group) were immunized at three timepoints (weeks 0, 4 and 20) with 30 µg of each of the test proteins formulated with Squalene Emulsion (SE) adjuvant. Sera were obtained from the rabbits at weeks 0, 4, 6, 16, 20 and 22 to characterize the antibody responses (Fig. 4a).
First, we measured the binding of the sera at the different timepoints to a His-tagged BG505 SOSIP.v5 trimer using a Ni-NTA ELISA assay. As expected, substantial increases in binding titers occurred at week 6 and week 22, two weeks after the second and third immunizations, respectively (Fig. 4b). We observed no significant differences in the binding titers induced by the different immunogens (Fig. S11a) at any timepoint. There were also no differences between the binding titers induced by the GM immunogens and their parental constructs (Fig. S11b).
Next, we measured the development of the NAb response against an autologous BG505 pseudovirus (Fig. 4c, Fig. S12). We observed that these autologous NAb responses evolved differently between weeks 6 and 20 for the different groups (Fig. 4c, Fig. S13). In some groups, in particular, the ones receiving SOSIP.v5, SOSIP.v9.3 and SOSIP.v9.3.GM, NAb responses peaked at week 6 and then declined resulting in lower NAb responses at week 20. In contrast, the NAb titers increased from week 6 to week 20 in the groups receiving SOSIP.v6, SOSIP.v9.4, SOSIP.v9.4.GM and SOSIP.v9.3.XL (Fig. 4c, Fig. S13). As a result, there was a trend that the groups receiving more stable proteins maintained higher NAb titers until week 20 (Fig. 4d). While this trend was not statistically significant, it would be consistent with the hypothesis that more stable immunogens might persist for longer thereby enhancing the durability of the response. Animals vaccinated with SOSIP.v6, SOSIP.v9.4 and SOSIP.v9.4.GM showed delayed responses, with no NAb activity detected at week 6 (two weeks after the second immunization), while a NAb response was detected by that time in the other rabbit groups. Nonetheless, all immunogens showed similar responses at week 16 and developed strong autologous NAb responses at week 22, after the third immunization (Fig. 4c, Fig. S12).
To assess the breadth of the NAb responses, we evaluated the neutralization of a panel of Tier 1 and Tier 2 heterologous viruses, including the viruses from the global HIV-1 panel53, at week 22 (Fig. S14). Except for some low titer responses, no consistent Tier 2 heterologous neutralization was observed. Remarkably, rabbits vaccinated with SOSIP.v9.4 and SOSIP.v9.4.GM immunogens showed significantly decreased NAb titers to the Tier 1 A MW965.26 virus (Fig. S15). Neutralizing responses to this virus are dominated by V3-targeting NAbs that are unable to neutralize Tier 2 primary HIV-1 isolates54, and the V3 might be less accessible in the SOSIP.v9.4 and SOSIP.v9.4.GM trimers as a consequence of the stabilization. Hence, NAb responses induced by these immunogens might be less hindered by competing V3-targeting narrow-NAb responses.
Autologous NAb responses correlate with SOSIP trimer stability
To assess the influence of the thermostability of the immunogen on the induction of autologous NAb responses, we used the neutralization titers determined for the BG505 pseudovirus virus at week 22 (Fig. 4e, Fig. S12). When we plotted the autologous NAb titers, expressed as the geometric mean (Geomean) of the ID50 values of each group, versus the nanoDSF-measured Tm values of the corresponding immunogen, we observed a positive correlation (p = 0.0167), indicating that increased thermostability is associated with the induction of stronger NAb responses (Fig. 4f). We also observed an inverse trend (p = 0.0833) between the nanoDSF-measured Tm values and the spread of the ID50 values within each group of animals, which suggests that higher thermostability increases the consistency of the NAb response (Fig. 4g). These trends were also observed when 2G12 and PGT145 thermostability ELISA-measured Tm values were used (Fig. S16a) and were found irrespective of the presence or absence of the cytoplasmic tail in the BG505 gp160 used for pseudovirus production (Fig. S16b).
Glycan masking redirects immunodominant glycan-hole responses to subdominant neutralizing epitopes
SOSIP.v9.3.GM and SOSIP.v9.4.GM immunogens were included in the rabbit immunization experiment to test the potential of the glycan masking strategy to redirect the NAb responses towards epitopes different than the 241/289 glycan hole. Surprisingly, we found that animals vaccinated with the glycan masked versions of SOSIP.v9.3 and SOSIP.v9.4 displayed autologous NAb responses of similar magnitude compared to those vaccinated with conventional SOSIP.v9.3 and SOSIP.v9.4, despite the lack of the dominant 241/289 NAb epitope (Fig. 4h). To map the neutralizing epitope of these animals we compared the ID50 titers of wild-type BG505 pseudovirus to the ID50 titer of a panel of mutant BG505 variants that contain PNGS sites at positions 241 and 289 (“241N/289N”), in the variable loop 1 at positions 133 and 136 (“133aN/136aA”) and at position 465 in the constant region 3 (465N) (Fig. S17). The relative ID50 titers (RID50) revealed that responses of most BG505 SOSIP-vaccinated rabbits induced a response to the immunodominant 241/289 glycan hole, consistent with previous studies42,55. However, few, if any, of NAbs elicited in recipients of GM-modified SOSIP trimers focused on the 241/289 glycan hole (p < 0.0001). Instead, NAb responses from these animals were significantly more focused on the 465 position than those in the non-GM groups (p = 0.0181) (Fig. 4i, Fig. S17). There was no difference for other epitopes, such as the one involving positions 133 and 136 (Fig. 4i). This indicates that the glycan masking strategy efficiently redirected the responses from the immunodominant 241/289 epitope to the subdominant C3/465 epitope, and possibly to other epitopes.
While the autologous neutralization in most animals could be attributed to Abs targeting either the 241/289 hole or the C3/465 epitope, we used a wider panel of mutant viruses to map the responses in animals in which these epitopes were not the major targets (i.e. animals UA139, UA142, UA155, and UA157). Only rabbit UA142 showed decreased neutralization to one of the mutants in the panel (BG505 G458Y), pointing at a CD4bs-targeting NAb response (Fig. S18). The NAb targets in the other three animals remain unidentified.
Discussion
We created new ultrastable recombinant SOSIP.v9 trimers that display increased thermal stability, increased purification yields and lower non-NAb reactivity compared to earlier SOSIP versions. The most thermostable trimer, SOSIP.v9.3.XL, with a melting temperature of 91.3 °C (a 14 °C increase compared to SOSIP.v6 and a 23.7 °C increase compared to SOSIP.664) induced the most potent and consistent Tier 2 autologous NAb titers after three immunizations. By characterizing the immunogenicity of a palette of SOSIP trimers with a range of melting temperatures, we found a positive correlation between thermal stability and both the magnitude and consistency of Tier 2 autologous NAb responses induced. We also used a glycan masking strategy to redirect the BG505 NAb responses from the immunodominant 241/289 glycan hole epitope to a more subdominant epitope.
It has been previously suggested that a correlation between immunogen thermostability and the potency of NAb responses exists56. However, this observation was based on NAb responses induced by Env trimers from diverse strains, which makes it difficult to ascertain whether the effect on the immunogenicity was due to differences in thermostability, antigenic profile, size of the glycan hole or other virus strain-specific features57. Here, we have demonstrated a correlation between immunogen thermostability and immunogenicity by using a palette of immunogens with different thermal stabilities derived from a single genotype (BG505). Moreover, NAb responses induced by SOSIP trimer immunogens without PNGS at 241/289 were predominantly directed to this 241/289 glycan hole, irrespective of their thermostability. Altogether, this indicates that the improvement in NAb titers is related to differences in thermal stability and not to a different antigenic profile.
Although highly thermostable immunogens from different viruses have been previously generated35,58,59,60, the mechanisms that explain the superior immunogenicity of thermally stabilized immunogens remain poorly understood. The delayed NAb responses we observed in animals vaccinated with SOSIP.v6, SOSIP.v9.4 and SOSIP.v9.4.GM, which were not detectable two weeks after the second immunization, might be a consequence of somewhat reduced epitope exposure. However, these highly stabilized immunogens may retain their native-like conformation in vivo for extended periods of time and thus increase the probability of displaying the proper antigenic determinants when they are encountered by immune cells, while less stable proteins might rapidly deteriorate into subdomains under in vivo conditions and expose epitopes that are irrelevant for neutralization. Furthermore, the rigidity provided by some stabilizing mutations reduces conformational flexibility and might help to induce neutralizing antibodies by removing entropy constraints61.
Previously, we found that the superior thermostability of SOSIP.v6 trimers versus earlier SOSIP forms leads to the induction of low titer heterologous neutralizing responses in some animals35. However, in the present study, none of the immunogens tested, including SOSIP.v6, induced any heterologous NAb responses. This is probably due to the fact that here we used a squalene emulsion-based adjuvant instead of ISCOMATRIX. Squalene emulsion is an adequate adjuvant, but it has been shown to induce less potent responses than ISCOMATRIX62 and thus might not have been potent enough to induce these heterologous responses.
Previous SOSIP versions have been shown to be stable for long periods of time in laboratory-controlled thermal conditions61,63, but ultrastable immunogens such as SOSIP.v9 trimers might be more suitable for situations in which the cold-chain conditions cannot be guaranteed during distribution and storage. Furthermore, although the induction of higher NAb titers is obviously desirable, the most important advantage of ultrastable proteins in terms of immunogenicity might be the increased consistency of the NAb responses. Based on this observation, ultrastable versions of already existing subunit vaccines could be engineered to induce more consistent coverage of the population of vaccinees and facilitate the distribution of vaccines in low-income countries.
We observed lower neutralization of the clade C Tier 1 MW965.26 viral strain in the animals vaccinated with SOSIP.v9.4 and SOSIP.v9.4.GM. These are the most thermostable trimers among the non-crosslinked immunogens we tested and include several mutations that suppress the immunodominant V3 and CD4i epitopes (316W, 306L, 308L, 201C-433C)30,31,32,34 and may result in the occlusion of other non-NAb epitopes due to reduced conformational flexibility. While the suppression of non-NAb epitopes might be beneficial for some specific applications, all evidence so far have shown that a lower induction of Abs targeting such regions is not correlated with an increased subdominant NAb response34. Therefore, the increased NAb titers here described for the ultrastable trimers may be attributed to their rigidity and stability rather than to the suppression of non-NAb epitopes. Recently, it has been shown that some of the antibodies elicited by HIV-1 trimer vaccines induce the degradation of the Env complex into cognate protomers64. Covalently linked Env proteins such as SOSIP.v9 trimers are probably less susceptible to this phenomenon, which might increase their half-life and thus improve the NAb responses elicited.
The 241/289 glycan hole BG505 Env is a highly immunodominant epitope and is targeted by immunized rabbits and, to a lesser extent, also by immunized macaques42,45,46,47. However, NAbs targeting strain-specific glycan holes are difficult to broaden48. Thus, NAbs targeting the BG505 241/289 glycan hole are probably only useful for neutralizing BG505 and at most a few other viruses that lack the 241 and 289 glycans. Furthermore, it has been recently shown that macaque NAbs that target the 241/289 glycan hole are less potent than those targeting other epitopes of the BG505 spike, such as the C3/V5 and V1/V3 regions65.
Ringe et al. previously showed that glycan masking the position 241 (241KI) of BG505 SOSIP.v4 trimers results in substantially decreased NAb responses against the parental BG505 virus, consistent with the removal of a dominant NAb epitope. Only by closing or opening specific artificial glycan holes at a different position were these BG505 SOSIP.v4 241KI trimers able to induce autologous NAb responses comparable to the ones induced by non-241KI trimers. Depending on the animal, these redirected responses targeted either the artificial glycan hole or the C3/465 epitope, or both. In contrast, we show here that glycan masking the immunodominant 241/289 epitope in the context of NAb-enhancing SOSIP.v9 trimers efficiently and consistently redirects responses to the subdominant C3/465 epitope, and possibly other epitopes, while maintaining a potent NAb response against the parental BG505 virus (Fig. 4h) and without the need of opening artificial glycan holes. Additional studies would be needed to determine if the C3/465-targeting NAb responses are likely to be broadened. In any case, the redirection capacity of filling the 241/289 hole in the context of SOSIP.v9 trimers should be useful for epitope-focusing and germline-targeting strategies that use SOSIP trimers as the platform66,67.
While additional studies focused on the generation of Env trimers from different genotypes without strain-specific glycan holes are necessary, it would also be crucial to immunosilence the glycan hole at the bottom of all soluble Env trimers. This region, which is not exposed in the native membrane-attached viral Env and thus generates non-NAb responses, attracts the majority of the early Ab responses targeting SOSIP trimers68. Several strategies, such as the incorporation of heterologous domains or additional PNGS that mask the bottom glycan hole, might further improve immunofocusing on SOSIP trimers towards bNAb epitopes.
In summary, the new SOSIP.v9 design, in particular SOSIP.v9.3.GM and SOSIP.v9.4.GM, might be a suitable platform for the generation of HIV-1 vaccine candidates. SOSIP.v9.4.GM showed higher thermostability and glycan hole immunosilencing than SOSIP.v9.3.GM. However, SOSIP.v9.4.GM produced less efficiently and showed some non-native forms. Therefore, it will be necessary to select the proper SOSIP platform depending on specific objectives or, alternatively, generate new designs that incorporate the advantages of both. Importantly, such highly thermostable and glycan masked designs might help in the enhancement and focusing of the NAb responses to specific epitopes. For instance, germline-targeting immunogens might significantly benefit from the combination of the ultrastabilization and glycan masking approaches, which might help suppress immunodominant strain-specific epitopes and redirect responses to the epitopes of interest.
Methods
Construct design
The BG505 SOSIP.v5 and SOSIP.v6 constructs, derived from HIV-1 subtype A BG505 env sequence, are extensively described and characterized elsewhere35. Briefly, BG505 SOSIP.v4 trimers include a set of amino acid changes to improve the expression and stability of soluble Env proteins: a TPA signal peptide (MDAMKRGLCCVLLLCGAVFVSPSQEIHARFRRGAR); 501C-605C (HxB2 numbering) (gp120-gp41 disulfide bond); 559P, 64K, and 316W trimer-stabilizing mutations; 535M and 543N, which improve trimerization; RRRRRR (R6) motif to enhance furin cleavage and a stop codon after residue 664. To further increase SOSIP proteins stability, we engineered 73C-561C intraprotomeric (SOSIP.v5) and 49C-555C interprotomeric (SOSIP.v6) disulfide bonds. The BG505 SOSIP.v9.1–9.4 constructs were generated by adding a new set of stabilizing mutations in different combinations: 1) 306L–308L mutations that stabilize the v3 loop region by hydrophobic interactions34; 2) MD39 mutations (304V, 319Y, 519S, 568D, 570H and 585H) to improve trimerization33; 3) a 201C-433C disulfide bond to increase stability and reduce V3 exposure and CD4 induction31. To obtain BG505 SOSIP.v9.3.GM and SOSIP.v9.4.GM glycan masked versions, PNGS motifs were introduced at positions 241 (241N) and 289 (291S), together with 240T, 271I, 288L, and 290E compensatory mutations.
Codon-optimized SOSIP.v9.1-v9.4 genes with a C-terminal His-Tag peptide (GSGSGGSGHHHHHHHH) were synthesized by Integrated DNA Technologies (Coralville, USA) and cloned by restriction-ligation into the pPPI4 vector. Point mutations to generate the glycan masked and untagged proteins were introduced by Quickchange site-directed mutagenesis (Agilent Technologies, La Jolla, CA, USA) and verified by sequencing.
Protein expression
SOSIP proteins were expressed in transiently transfected HEK293F suspension cells (Invitrogen, cat no. R79009), maintained in FreeStyle Expression Medium (Gibco). For transfection, HIV-1 Env and furin protease-encoding plasmids mixed in a 4:1 Env to furin ratio (w/w) were incubated with PEImax (Polysciences Europe GmBH, Eppelheim, Germany) in a 3:1 (w/w) PEImax to DNA ratio and then added in the supernatant of cells at a density of 0.8–1.2 million cells/mL. Five to seven days post-transfection, supernatants were harvested, centrifuged, and filtered using Steritops (0.22 µm pore size; Millipore, Amsterdam, The Netherlands) before protein purification.
Protein purification
Both non-crosslinked and crosslinked SOSIP proteins were purified by immunoaffinity chromatography with PGT145 or PGT151 as selecting agents, respectively32,51. Unpurified proteins contained in HEK293F filtered supernatants or Tris-buffered saline (TBS) solutions were captured on PGT145- or PGT151-functionalized CNBr-activated sepharose 4B beads (GE Healthcare) by overnight rolling incubation at 4 °C. Subsequently, the mixes of supernatant and beads were passed over Econo-Column chromatography columns (Biorad). The columns were then washed with three column volumes of a 0.5 M NaCl and 20 mM Tris HCl pH 8.0 solution. After elution with 3 M MgCl2 pH 7.5, proteins were buffer-exchanged into TN75 (75 mM NaCl, 20 mM Tris HCl pH 8.0) or PBS buffers by ultrafiltration with Vivaspin20 filters (Sartorius, Gӧttingen, Germany) of MWCO 100 kDa. Protein concentrations were determined from the A280 values measured on a NanoDrop2000 device (Thermo Fisher Scientific) and the molecular weight and extinction coefficient values calculated by the ProtParam Expasy webtool.
Proteins used in immunization experiments were first PGT145-purified and subsequently run through a Superdex 200 Increase 10/300 GL (GE Healthcare Life Sciences) column integrated into an NGC chromatography system (Bio-Rad). The fractions corresponding to trimers were pooled, concentrated, and filter-sterilized to avoid adverse reactions in immunized animals.
Protein chemical crosslinking
To obtain the BG505 SOSIP.v9.3.XL protein, GLA crosslinking was performed essentially as described elsewhere37. Briefly, the PGT145-purified BG505 SOSIP.v9.3 protein in PBS was mixed with glutaraldehyde (GLA, Agar Scientific) to a final concentration of 7.5 mM. After a 5 min incubation at room temperature, the crosslinking reaction was stopped by the addition of 75 mM Tris buffer (pH 7.4) and the resulting protein was buffer-exchanged to TBS and further purified by PGT151 affinity chromatography, as above described.
SDS-PAGE AND BN-PAGE
Purified proteins (2–3 µg) were run over NuPAGE 4–12% Bis-Tris and Novex Wedgewell 4–12% Tris-Glycine (both from Invitrogen) polyacrylamide gels for SDS-PAGE and BN-PAGE analysis, respectively32. Subsequently, gels were stained with PageBlue Protein Staining Solution (Thermo Scientific) or the Colloidal Blue Staining Kit (Life Technologies), respectively. Proteins presented in each gel were processed in parallel.
Enzyme-linked immunosorbent assay (ELISA)
ELISA experiments with His-tagged SOSIP trimers were performed essentially as described before13,51. Purified proteins (1 µg/mL) were diluted in TBS and immobilized on 96-well Ni-NTA functionalized ELISA plates (Qiagen) by a 2 h incubation at room temperature. Following a double wash step with TBS to remove unbound trimers, serial dilutions of test antibodies in TBS/2% skimmed milk were added and incubated for 2 h. After 3 washes with TBS, HRP-labeled goat anti-human IgG (Jackson Immunoresearch) diluted 1:3000 in TBS/2% skimmed milk was added and incubated for 1 h, followed by 5 washes with TBS/0.05% Tween20. A developing solution (1% 3,3’,5,5’-tetramethylbenzidine (Sigma-Aldrich), 0.01% H2O2, 100 mM sodium acetate and 100 mM citric acid) allowed the colorimetric reaction, which was stopped by the addition of 0.8 M H2SO4. Finally, color development (absorption at 450 nm, OD450) was measured to obtain the different binding curves.
Endpoint antibody titers from rabbit sera samples were determined using Ni-NTA His-Tag-capture ELISA similarly as above described6,51. However, in this case, sera was diluted in TBS/2% skimmed milk/20% sheep serum and the secondary antibody was an HRP-labeled goat antirabbit IgG (Jackson Immunoresearch).
Thermostability ELISA
For assessing the thermostability of 2G12 and PGT145 epitopes on the different SOSIP proteins, we submitted the trimers for 30 min to a set of temperatures ranging from 25 °C to 90.6 °C. Subsequently, they were diluted in TBS and immobilized on Ni-NTA as above described. Antibody binding was assessed by incubation with a single Casein Blocker (Thermo Scientific) solution with a 2G12 or PGT145 concentration close to their priorly determined IC50 (0.1 µg/mL). Subsequent steps to assess the binding of the test antibodies were performed as above described. Temperatures of melting (Tm) were determined by interpolation of the temperature values that corresponded to 50% of the maximum binding signal, using Graphpad Prism 8.4.3
Biolayer interferometry (BLI)
The BLI assay was performed using an Octet K2 (ForteBio) device at 30 °C and 1000 rpm agitation. First, test antibodies diluted in kinetics buffer (PBS/0.1% bovine serum albumin/0.02% Tween20) were loaded on protein A sensors (ForteBio) to an interference pattern shift of 1 nm. Sensors were equilibrated in kinetics buffer for 60 s to obtain a baseline prior to protein association. Subsequently, purified SOSIP trimers diluted in kinetics buffer (100 nM) were allowed to associate and dissociation for 300 s. Binding data was pre-processed and exported using the Octet software.
Nano Differential Scanning Fluorimetry (nanoDSF)
Protein thermostability was evaluated with a Prometheus NT.48 instrument (NanoTemper Technologies). Proteins at a concentration of 1 mg/mL were loaded to the grade capillaries and the intrinsic fluorescence signal was measured while temperature was increased by 1 °C/min, with an excitation power of 40%. The temperature of onset (Tonset) and temperature of melting (Tm) were determined using the Prometheus NT software.
Negative-stain electron microscopy (NS-EM)
SOSIP proteins were imaged by NS-EM essentially as described elsewhere69. Briefly, 3 µL protein aliquots at ∼0.03 mg/mL were applied onto a glow discharged (20 mA for 30 s) carbon-coated 400 Cu mesh grid. After 5 s, they were negatively stained with 2% (w/v) uranyl formate for 60 s. Images were taken using a FEI Tecnai T12 electron microscope operating at 120 keV.
Site-specific glycan analysis using mass spectrometry
The BG505 SOSIP.v9.3.GM was denatured for 1 h in 50 mM Tris/HCl, pH 8.0 containing 6 M of urea and 5 mM dithiothreitol (DTT). Next, the protein was reduced and alkylated by adding 20 mM iodacetamide (IAA) and incubated for 1 h in the dark, followed by a 1 h incubation with 20 mM DTT to eliminate residual IAA. The alkylated protein was buffer-exchanged into 50 mM Tris/HCl, pH 8.0 using a Vivaspin ultrafiltration unit (Sartorius, Gӧttingen, Germany) of MWCO 3 kDa and digested separately using trypsin or chymotrypsin (Mass Spectrometry Grade, Promega) at a ratio of 1:30 (w/w) by overnight incubation. Peptides were dried, extracted using C18 Zip-tip (MerckMilipore) and dried again before being resuspended in 0.1% formic acid and analyzed by nanoLC-ESI MS with an Easy-nLC 1200 (Thermo Fisher Scientific) system coupled to a Fusion mass spectrometer (Thermo Fisher Scientific) using higher energy collision-induced dissociation (HCD) fragmentation. Peptides were separated using an EasySpray PepMap RSLC C18 column (75 µm x 75 cm). The LC conditions were as follows: 275-min linear gradient consisting of 0–32% acetonitrile in 0.1% formic acid over 240 min followed by 35 min of 80% acetonitrile in 0.1% formic acid. The flow rate was set to 200 nL/min, the spray voltage, to 2.7 kV and the temperature of the heated capillary, to 40 °C. The ion transfer tube temperature was set to 275 °C. The scan range was 400–1600 m/z. The HCD collision energy was set to 50%, appropriate for fragmentation of glycopeptide ions. Precursor and fragment detection were performed using an Orbitrap at a resolution MS1 = 100,000. MS2 = 30,000. The AGC target for MS1 = 4e5 and MS2 = 5e4 and injection time were MS1 = 50 and MS2 = 54.
Glycopeptide fragmentation data were extracted from the raw file using ByonicTM (Version 3.4) and ByologicTM software (Version 3.4; Protein Metrics Inc.) and evaluated manually for each glycopeptide. The peptide was scored as true-positive when the correct b and y fragment ions were observed along with oxonium ions corresponding to the glycan identified. The MS data was searched using a standard library. The precursor mass tolerance was set at 4 ppm and 10 ppm for fragments. A 1% false discovery rate (FDR) was applied. The relative amounts of each glycan at each site as well as the unoccupied proportion were determined by comparing the extracted ion chromatographic areas for different glycopeptides with an identical peptide sequence.
Site-specific analysis of low abundance N-glycan sites using mass spectrometry
To obtain data for sites that frequently present low-intensity glycopeptide, the glycans present on the glycopeptides were homogenized to boost the intensity of these peptides. This analysis loses fine processing information but enables the ratio of oligomannose:complex:unoccupied to be determined. The remaining glycopeptides were first digested with Endo H (New England Biolabs) to deplete oligomannose- and hybrid-type glycans and leave a single GlcNAc residue at the corresponding site. The reaction mixture was then dried completely and resuspended in a mixture containing 50 mM ammonium bicarbonate and PNGase F (New England Biolabs) using only H2O18 (Sigma-Aldrich) throughout. This second reaction cleaves the remaining complex-type glycans but leaves the GlcNAc residues remaining after Endo H cleavage intact. The use of H2O18 in this reaction enables complex glycan sites to be differentiated from unoccupied glycan sites as the hydrolysis of the glycosidic bond by PNGaseF leaves a heavy oxygen isotope on the resulting aspartic acid residue. The resultant peptides were purified as outlined above and subjected to reverse-phase (RP) nanoLC-MS. Instead of the extensive N-glycan library used above, two modifications were searched for: +203 Da corresponding to a single GlcNAc, a remnant of an oligomannose/hybrid glycan, and +3 Da corresponding to the O18 deamidation product of a complex glycan. A lower HCD energy of 27% was used as glycan fragmentation was not required. Data analysis was performed as above and the relative amounts of each glycoform determined, including unoccupied peptides.
Immunization experiment
The immunization experiment was performed by Covance Research Products Inc. (Denver, PA, USA) in female and naïve New Zealand rabbits (2.5 – 3 kg) arbitrarily distributed in 7 different groups (5 rabbits per group). Rabbits were immunized at 3 different timepoints (weeks 0, 4, and 20) with 30 µg of Env proteins diluted in PBS and formulated with Squalene emulsion (SE) adjuvant (Polymun, Klosterneuburg, Austria) in a 1:1 Env in PBS to SE ratio (v/v). Each immunization consisted of two intramuscular injections (2 × 250 µL) on both quadriceps of each animal. All immunization procedures were performed under compliance with all relevant ethical regulations for animal testing and research and received ethical approval from the Covance Institutional Animal Care and Use Committee (IACUC), with approval number C0041-19.
Neutralization assays
TZM-bl neutralization assays were performed as described elsewhere70 at two different sites: Amsterdam UMC (AMC), Amsterdam, The Netherlands and Duke University Medical Center (DUMC), Durham, NC, USA. Briefly, 3-fold dilutions of heat-inactivated sera (starting form 1:20 dilution) were mixed with Env-pseudotyped viruses and incubated for 1 h at RT. Subsequently, the mix was added to TZM-bl cells seeded the previous day at a density of 17.000 cells/well. After 72 h, the cells were lysed and luciferase activity was measured using the Bright Glo Luciferase kit (Promega) in a Glomax plate reader. Neutralization titers (ID50 values) were determined as the serum dilution at which infectivity was inhibited by 50%.
BG505/T332N (Fig. 4h, i, Fig. S14, 17, 18) and cytoplasmatic tail truncated BG505 (BG505.dCT) (Fig. 4c–g, Fig. S11, 12, 13, 15, 16) pseudoviruses were used to measure the autologous neutralizing responses. The TZM-bl reporter cell line was obtained from John C. Kappes and Xiaoyun Wu and Tranzyme Inc. through the NIH AIDS Research and Reference Reagents Program, NIAID, NIH.
Data representation and statistical analyses
All data representation and statistical analyses were performed using Graphpad Prism 8.4.3. Groups were compared using unpaired two-tailed Mann–Whitney U-test and multiple comparisons were analyzed with Kruskal–Wallis test, unless otherwise specified. Pearson or Spearman correlation coefficients were used to determine correlations, as specified in each case.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data that support the findings in this study are available from the corresponding author (R.W.S.) upon reasonable request.
References
Kwong, P. D. & Mascola, J. R. HIV-1 vaccines based on antibody identification, B cell ontogeny, and epitope structure. Immunity 48, 855–871 (2018).
McCoy, L. E. The expanding array of HIV broadly neutralizing antibodies. Retrovirology 15, 70 (2018).
Sok, D. & Burton, D. R. Recent progress in broadly neutralizing antibodies to HIV. Nat. Immunol. 19, 1179–1188 (2018).
del Moral-Sánchez, I. & Sliepen, K. Strategies for inducing effective neutralizing antibody responses against HIV-1. Expert Rev. Vaccines 18, 1127–1143 (2019).
Kwong, P. D. et al. HIV-1 evades antibody-mediated neutralization through conformational masking of receptor-binding sites. Nature 420, 678–682 (2002).
Sanders, R. W. et al. HIV-1 VACCINES. HIV-1 neutralizing antibodies induced by native-like envelope trimers. Science 349, aac4223 (2015).
Munro, J. B. et al. Conformational dynamics of single HIV-1 envelope trimers on the surface of native virions. Science 346, 759–763 (2014).
Yasmeen, A. et al. Differential binding of neutralizing and non-neutralizing antibodies to native-like soluble HIV-1 Env trimers, uncleaved Env proteins, and monomeric subunits. Retrovirology 11, 41 (2014).
Wyatt, R. et al. The antigenic structure of the HIV gp120 envelope glycoprotein. Nature 393, 705–711 (1998).
Gaschen, B. et al. Diversity considerations in HIV-1 vaccine selection. Science 296, 2354–2360 (2002).
Wei, X. et al. Antibody neutralization and escape by HIV-1. Nature 423, 197–197 (2003).
Seabright, G. E., Doores, K. J., Burton, D. R. & Crispin, M. Protein and glycan mimicry in HIV vaccine design. J. Mol. Biol. 431, 2223–2247 (2019).
Sanders, R. W. et al. A next-generation cleaved, soluble HIV-1 Env trimer, BG505 SOSIP.664 gp140, expresses multiple epitopes for broadly neutralizing but not non-neutralizing antibodies. PLoS Pathog. 9, e1003618 (2013).
Lee, J. H., Ozorowski, G. & Ward, A. B. Cryo-EM structure of a native, fully glycosylated, cleaved HIV-1 envelope trimer. Science 351, 1043–1048 (2016).
Lyumkis, D. et al. Cryo-EM structure of a fully glycosylated soluble cleaved HIV-1 envelope trimer. Science 342, 1484–1490 (2013).
Georgiev, I. S. et al. Single-chain soluble BG505.SOSIP gp140 trimers as structural and antigenic mimics of mature closed HIV-1 Env. J. Virol. 89, 5318–5329 (2015).
Sharma, S. K. et al. Cleavage-independent HIV-1 Env trimers engineered as soluble native spike mimetics for vaccine design. Cell Rep. 11, 539–550 (2015).
Guenaga, J. et al. Glycine substitution at helix-to-coil transitions facilitates the structural determination of a stabilized subtype C HIV envelope glycoprotein. Immunity 46, 792–803.e3 (2017).
Sarkar, A. et al. Structure of a cleavage-independent HIV Env recapitulates the glycoprotein architecture of the native cleaved trimer. Nat. Commun. 9, 1956 (2018).
Binley, J. M. et al. Enhancing the proteolytic maturation of human immunodeficiency virus type 1 envelope glycoproteins. J. Virol. 76, 2606–2616 (2002).
Binley, J. M. et al. A recombinant human immunodeficiency virus type 1 envelope glycoprotein complex stabilized by an intermolecular disulfide bond between the gp120 and gp41 subunits is an antigenic mimic of the trimeric virion-associated structure. J. Virol. 74, 627–643 (2000).
Sanders, R. W. et al. Stabilization of the soluble, cleaved, trimeric form of the envelope glycoprotein complex of human immunodeficiency virus type 1. J. Virol. 76, 8875–8889 (2002).
Pallesen, J. et al. Immunogenicity and structures of a rationally designed prefusion MERS-CoV spike antigen. Proc. Natl Acad. Sci. 114, E7348–E7357 (2017).
Sanders, R. W. & Moore, J. P. Virus vaccines: proteins prefer prolines. Cell Host Microbe 29, 327–333 (2021).
Martinez-Murillo, P. et al. Particulate array of well-ordered HIV clade C Env trimers elicits neutralizing antibodies that display a unique V2 cap approach. Immunity 46, 804–817.e7 (2017).
Bale, S. et al. Cleavage-Independent HIV-1 Trimers From CHO cell lines elicit robust autologous Tier 2 neutralizing antibodies. Front. Immunol. 9, 1116 (2018).
de la Peña, A. T. & Sanders, R. W. Stabilizing HIV-1 envelope glycoprotein trimers to induce neutralizing antibodies. Retrovirology 15, 63 (2018).
Sliepen, K., van Montfort, T., Melchers, M., Isik, G. & Sanders, R. W. Immunosilencing a highly immunogenic protein trimerization domain. J. Biol. Chem. 290, 7436–7442 (2015).
Karp, C. L. et al. Evaluating the value proposition for improving vaccine thermostability to increase vaccine impact in low and middle-income countries. Vaccine 33, 3471–3479 (2015).
Guenaga, J. et al. Well-ordered trimeric HIV-1 subtype B and C soluble spike mimetics generated by negative selection display native-like properties. PLoS Pathog. 11, e1004570 (2015).
Kwon, Y. D. et al. Crystal structure, conformational fixation and entry-related interactions of mature ligand-free HIV-1 Env. Nat. Struct. Mol. Biol. 22, 522–531 (2015).
de Taeye, S. W. et al. Immunogenicity of Stabilized HIV-1 envelope trimers with reduced exposure of non-neutralizing epitopes. Cell 163, 1702–1715 (2015).
Steichen, J. M. et al. HIV vaccine design to target germline precursors of glycan-dependent broadly neutralizing antibodies. Immunity 45, 483–496 (2016).
de Taeye, S. W. et al. Stabilization of the gp120 V3 loop through hydrophobic interactions reduces the immunodominant V3-directed non-neutralizing response to HIV-1 envelope trimers. J. Biol. Chem. 293, 1688–1701 (2018).
de la Peña, A. T. et al. Improving the immunogenicity of native-like HIV-1 envelope trimers by hyperstabilization. Cell Rep. 20, 1805–1817 (2017).
Schiffner, T. et al. Immune focusing and enhanced neutralization induced by HIV-1 gp140 chemical cross-linking. J. Virol. 87, 10163–10172 (2013).
Schiffner, T. et al. Chemical cross-linking stabilizes native-like HIV-1 envelope glycoprotein trimer antigens. J. Virol. 90, 813–828 (2016).
Schiffner, T. et al. Structural and immunologic correlates of chemically stabilized HIV-1 envelope glycoproteins. PLoS Pathog. 14, e1006986 (2018).
Wei, X. et al. Antibody neutralization and escape by HIV-1. Nature 422, 307–312 (2003).
Sagar, M., Wu, X., Lee, S. & Overbaugh, J. Human immunodeficiency virus type 1 V1-V2 envelope loop sequences expand and add glycosylation sites over the course of infection, and these modifications affect antibody neutralization sensitivity. J. Virol. 80, 9586–9598 (2006).
Wagh, K. et al. Completeness of HIV-1 envelope glycan shield at transmission determines neutralization breadth. Cell Rep. 25, 893–908.e7 (2018).
McCoy, L. E. et al. Holes in the glycan shield of the native HIV envelope are a target of trimer-elicited neutralizing antibodies. Cell Rep. 16, 2327–2338 (2016).
de la Peña, A. T. et al. Immunogenicity in rabbits of HIV-1 SOSIP trimers from clades A, B, and C, given individually, sequentially, or in combination. J. Virol. 92, e01957 (2018).
Crooks, E. T. et al. Vaccine-elicited Tier 2 HIV-1 neutralizing antibodies bind to quaternary epitopes involving glycan-deficient patches proximal to the CD4 binding site. PLoS Pathog. 11, e1004932 (2015).
Klasse, P. J. et al. Epitopes for neutralizing antibodies induced by HIV-1 envelope glycoprotein BG505 SOSIP trimers in rabbits and macaques. PLoS Pathog. 14, e1006913 (2018).
Pauthner, M. et al. Elicitation of robust Tier 2 neutralizing antibody responses in nonhuman primates by HIV envelope trimer immunization using optimized approaches. Immunity 46, 1073–1088.e6 (2017).
Nogal, B. et al. Mapping polyclonal antibody responses in non-human primates vaccinated with HIV Env trimer subunit vaccines. Cell Rep. 30, 3755–3765.e7 (2020).
Yang, Y. R. et al. Autologous antibody responses to an HIV envelope glycan hole are not easily broadened in rabbits. J. Virol. 94, e01861 (2020).
Nogal, B. et al. HIV envelope trimer-elicited autologous neutralizing antibodies bind a region overlapping the N332 glycan supersite. Sci. Adv. 6, eaba0512 (2020).
Ringe, R. P. et al. Closing and opening holes in the glycan shield of HIV-1 envelope glycoprotein SOSIP trimers can redirect the neutralizing antibody response to the newly unmasked epitopes. J. Virol. 93, e01656 (2019).
Sliepen, K. et al. Structure and immunogenicity of a stabilized HIV-1 envelope trimer based on a group-M consensus sequence. Nat. Commun. 10, 2355 (2019).
Soldemo, M. et al. Glutaraldehyde Cross-linking of HIV-1 Env Trimers Skews the Antibody Subclass Response in Mice. Front. Immunol. 8, 1654 (2017).
deCamp, A. et al. Global panel of HIV-1 Env reference strains for standardized assessments of vaccine-elicited neutralizing antibodies. J. Virol. 88, 2489–2507 (2014).
Qualls, Z. M. et al. Identification of novel structural determinants in MW965 Env that regulate the neutralization phenotype and conformational masking potential of primary HIV-1 isolates. J. Virol. 92, e01779 (2018).
Klasse, P. J. et al. Sequential and simultaneous immunization of rabbits with HIV-1 envelope glycoprotein SOSIP.664 trimers from clades A, B and C. PLoS Pathog. 12, e1005864 (2016).
Feng, Y. et al. Thermostability of well-ordered HIV spikes correlates with the elicitation of autologous Tier 2 neutralizing antibodies. PLoS Pathog. 12, e1005767 (2016).
Zhou, T. et al. Quantification of the impact of the HIV-1-glycan shield on antibody elicitation. Cell Rep. 19, 719–732 (2017).
Rostad, C. A. et al. Enhancing the thermostability and immunogenicity of a respiratory syncytial virus (RSV) live-attenuated vaccine by incorporating unique RSV line19F protein residues. J. Virol. 92, e01568 (2018).
Du, P. et al. Biomineralization improves the thermostability of foot-and-mouth disease virus-like particles and the protective immune response induced. Nanoscale 11, 22748–22761 (2019).
Stobart, C. C. et al. A live RSV vaccine with engineered thermostability is immunogenic in cotton rats despite high attenuation. Nat. Commun. 7, 13916 (2016).
Kong, L. et al. Local conformational stability of HIV-1 gp120 in unliganded and CD4-bound states as defined by amide hydrogen/deuterium exchange. J. Virol. 84, 10311–10321 (2010).
Sliepen, K. et al. Interplay of diverse adjuvants and nanoparticle presentation of native-like HIV-1 envelope trimers. NPJ Vaccines 6(1), 103 (2021).
Dey, A. K. et al. cGMP production and analysis of BG505 SOSIP.664, an extensively glycosylated, trimeric HIV-1 envelope glycoprotein vaccine candidate. Biotechnol. Bioeng. 115, 885–899 (2018).
Turner, H. L. et al. Disassembly of HIV envelope glycoprotein trimer immunogens is driven by antibodies elicited via immunization. Sci. Adv. 7 (2021).
Zhao, F. et al. Mapping neutralizing antibody epitope specificities to an HIV Env trimer in immunized and in infected Rhesus Macaques. SSRN Electron. J. https://doi.org/10.2139/ssrn.3578689 (2020).
Medina-Ramírez, M. et al. Design and crystal structure of a native-like HIV-1 envelope trimer that engages multiple broadly neutralizing antibody precursors in vivo. J. Exp. Med. 214, 2573–2590 (2017).
Steichen, J. M. et al. A generalized HIV vaccine design strategy for priming of broadly neutralizing antibody responses. Science 366, eaax4380 (2019).
Bianchi, M. et al. Electron-microscopy-based epitope mapping defines specificities of polyclonal antibodies elicited during HIV-1 BG505 envelope trimer immunization. Immunity 49, 288–300.e8 (2018).
Pritchard, L. K. et al. Structural constraints determine the glycosylation of HIV-1 envelope trimers. Cell Rep. 11, 1604–1613 (2015).
Montefiori, D. C. Measuring HIV neutralization in a luciferase reporter gene assay. Methods Mol. Biol. 485, 395–405 (2009).
Acknowledgements
The authors thank Marit van Gils for donating the 10A monoclonal antibody; and Michel Nussenzweig, Hermann Katinger, Mark Connors, James Robinson, Dennis Burton, John Mascola, Peter Kwong, and William Olson for donating antibodies and reagents directly or through the NIH AIDS Research and Reference Reagent Program. We thank Dietmar Katinger and Ehsan Suleiman for providing the squalene emulsion adjuvant. This project has received funding from the European Union’s Horizon 2020 research and innovation program under grant agreement No. 681137 (to R.W.S., Q.S., and M.C.). This work was also supported by the U.S. National Institutes of Health Grant P01 AI110657 (to A.B.W. and R.W.S.) and NIAID Contract #HHSN27201100016C (to D.C.M.); by the International AIDS Vaccine Initiative (IAVI); by the Bill and Melinda Gates Foundation through the Collaboration for AIDS Vaccine Discovery (CAVD), grants OPP1111923 and OPP1132237 (to R.W.S.) and OPP1115782 (A.B.W.); by the Aids Fonds Netherlands, Grant #2016019 (to R.W.S.); and by the Fondation Dormeur, Vaduz (to R.W.S.). R.W.S. is a recipient of a Vici grant from the Netherlands Organization for Scientific Research (NWO). We thank EMBO for the Short-Term Fellowship (STS-7814) awarded to S.K. The electron microscopy data were collected at Electron Microscopy Facility of The Scripps Research Institute.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: I.d.M.S., R.A.R., E.E.S., C.A.C., J.D.A., A.T.d.l.P., C.C.L., S.K., M.C., A.B.W., D.C.M., Q.J.S., K.S., R.W.S. Performed the experiments: I.d.M.S., R.A.R., E.E.S., C.A.C., J.D.A., C.C.L., S.K., K.S. Analyzed the data: I.d.M.S., R.A.R., E.E.S., C.A.C., J.D.A., C.C.L., S.K., K.S. Wrote the paper: I.d.M.S., J.D.A., K.S., R.W.S. Edited or revised the paper: All authors commented on the manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
del Moral-Sánchez, I., Russell, R.A., Schermer, E.E. et al. High thermostability improves neutralizing antibody responses induced by native-like HIV-1 envelope trimers. npj Vaccines 7, 27 (2022). https://doi.org/10.1038/s41541-022-00446-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41541-022-00446-4
This article is cited by
-
Triple tandem trimer immunogens for HIV-1 and influenza nucleic acid-based vaccines
npj Vaccines (2024)
-
CoPoP liposomes displaying stabilized clade C HIV-1 Env elicit tier 2 multiclade neutralization in rabbits
Nature Communications (2024)
-
Single-component multilayered self-assembling protein nanoparticles presenting glycan-trimmed uncleaved prefusion optimized envelope trimers as HIV-1 vaccine candidates
Nature Communications (2023)
-
Fine-mapping the immunodominant antibody epitopes on consensus sequence-based HIV-1 envelope trimer vaccine candidates
npj Vaccines (2022)