Abstract
Spaceflight causes muscle wasting. The Sarcolab pilot study investigated two astronauts with regards to plantar flexor muscle size, architecture, and function, and to the underlying molecular adaptations in order to further the understanding of muscular responses to spaceflight and exercise countermeasures. Two crew members (A and B) spent 6 months in space. Crew member A trained less vigorously than B. Postflight, A showed substantial decrements in plantar flexor volume, muscle architecture, in strength and in fiber contractility, which was strongly mitigated in B. The difference between these crew members closely reflected FAK-Y397 abundance, a molecular marker of muscle’s loading history. Moreover, crew member A showed downregulation of contractile proteins and enzymes of anaerobic metabolism, as well as of systemic markers of energy and protein metabolism. However, both crew members exhibited decrements in muscular aerobic metabolism and phosphate high energy transfer. We conclude that countermeasures can be effective, particularly when resistive forces are of sufficient magnitude. However, to fully prevent space-related muscular deterioration, intersubject variability must be understood, and intensive exercise countermeasures programs seem mandatory. Finally, proteomic and metabolomic analyses suggest that exercise benefits in space may go beyond mere maintenance of muscle mass, but rather extend to the level of organismic metabolism.
Similar content being viewed by others
Introduction
Physical deconditioning is known to occur during spaceflight since Skylab and Mir missions. Lower limb muscles, in particular, undergo rapid wasting and loss of function.1,2 Most affected is the triceps surae muscle, where 20% fiber atrophy occurs after 6 months of spaceflight.3,4 In ground-based models (e.g. bed rest), muscle atrophy is associated with decreases in fascicle pennation angle and length.5,6 Alterations of muscle architecture are expected to affect the mechanical output, thereby contributing to muscle weakness.7 Moreover, muscle unloading also leads to reductions in the fibers’ specific force and power, and in myosin heavy chain (MCH) concentration.8 All these factors can independently alter the mechanical capabilities of muscles.
Muscle atrophy results from imbalance between protein synthesis and degradation. This imbalance can be caused by enhanced muscle protein breakdown (MPB), controlled by catabolic pathways (ubiquitin proteasome and autophagy), and also by inhibited muscle protein synthesis (MPS), controlled by the Akt/mTOR/p70S6K pathway.9 To date, the relative contribution of MPB and MPS is still unclear.10 The determination of the actual rates of protein synthesis and degradation in humans is challenging. The relative activation of intracellular pathways involved varies across species and disuse conditions. Moreover, it is unclear whether a metabolic program plays a relevant role in causing disuse atrophy in humans. Recently, it has been suggested that mitochondrial dysfunction is a major trigger of MPB and MPS imbalance.11,12 Solving these open issues is important for spaceflight.
Countermeasure exercises are nowadays mandatory on board the International Space Station (ISS). With regards to muscle, they involve exercises with the advanced resistive exercise device (aRED) and a treadmill (T2). Effects of strength training upon skeletal muscle on Earth have been studied extensively, but relatively little is known about the molecular events in disuse or in spaceflight.13 It is therefore an open question how far training in space is helpful to maintain the lower limb musculature.
Costameric proteins can serve as a molecular proxy of the muscle’s loading history.14,15,16,17,18,19,20 They anchor the sarcomeres to extracellular matrix receptors.21,22,23 Among these, the integrin-linked focal adhesion kinase (FAK) is a mechanically regulated costamere component that controls the turnover of focal adhesion in a fiber type-specific manner together with FAK-related non-kinase FRNK, FAK’s natural inhibitor.18,21,24 Herein, the content of post-translation modification of tyrosine 397 (Y397) is a critical event, which is affecting the activity of FAK.25,26,27 Importantly, the FAK pathway likely controls protein synthesis through 70S6K, a component of the Akt/mTOR pathway.
Thus, the Sarcolab study has been designed to: (i) disentangle the various constituents of muscle weakness associated with spaceflight; (ii) elucidate the muscular adaptations to long-term spaceflight; (iii) address the affected molecular and metabolic pathways in skeletal muscle and in the blood in astronauts during long missions on ISS. The main hypothesis of the Sarcolab study is that alterations in the physiological cross-sectional area, fiber length, as well as in single fiber mechanics collectively contribute to the space-related muscle weakness. Moreover, the study aims to screen for muscle proteomic adaptations to spaceflight. Finally, intracellular signaling pathways, namely those controlling muscle mass (ubiquitin proteasome pathway, autophagy, FAK) and metabolism (PGC-1alpha, SREBP-1) are studied, and blood metabolomics screens for systemic consequences-related muscular alterations. The present paper reports results from two astronauts who took part in the Sarcolab pilot study, before the experiment had been enlarged into the currently performed three-agency study (Sarcolab3), supported by ESA, NASA, and Roscosmos.
Results
Onboard exercise training
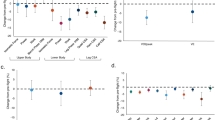
Crew member A performed fewer treadmill sessions than B (90 vs. 114), ran with lower pull-down force (median of 55.9 vs. 85.6% of body weight), ran at slower speed (median 11.3 vs. 12.9 km/h), and covered a shorter distance than B per running session (median 4.7 vs. 5.8 km, Fig. 1). Crew member A also trained less with aRED, performing fewer heel raise sessions (54 vs. 98) with fewer repetitions (median 30 vs. 48) and at lower resistive force (median 122 vs. 221% of body weight). Thus, A was generally training less vigorously than B, in particular with regards to exercise elements related to loading force.
Muscle function
During postflight session 1 (PF-1, performed 0−4 days after return to Earth), plantar flexor muscle strength was reduced by 30.6% in crew member A, whereas B depicted no change from baseline (Fig. 2).
Muscle size and architecture
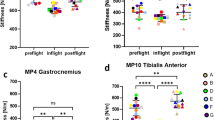
Medial gastrocnemius (GM) muscle volume, pennation angle (PA), and fascicle length (Lf) at PF-1 were all substantially reduced in crew member A, but much less in B (Table 1). Physiological cross-sectional area (pCSA = Vol/Lf) was affected to comparable extent in both crew members at PF-1 (Table 1). At PF-2 (performed at R + 15), pCSA of the GM muscle had recovered in B, but not in A.
Fiber type, fiber size and fiber mechanics
At baseline, soleus (SOL) muscle was almost exclusively composed of the slow isoform of myosin heavy chain (MHC-1) in both crew members. No shift in MHC relative content was observed postflight. Cross-sectional area of single muscle fibers (CSA_SF) at PF-1 declined in crew member A by −45%, but only by −23% in B (Fig. 2).
Functional single fiber analysis could be performed pre- and postflight in crew member A only. All fibers analyzed were type 1 fibers. Specific isometric force (Po/CSA) was lower postflight than preflight (−27%). Unloaded shortening velocity (Vo) as well as actin sliding velocity (Vf) were both comparable pre- vs. postflight, suggesting no alteration in the actomyosin kinetics and no alteration of myosin at molecular level.
Costameric protein expression
All costameric proteins studied were detectable in SOL at baseline (Figures S1 to S3). Gamma-vinculin was more abundant than meta-vinculin (Figure S1). At PF-1, protein concentrations of FAK (normalized to actin) was reduced in crew members A and B by −60 and −44%, respectively, and FRNK by −60 and −67% (Table 2, Figure S2). By contrast, FAK and FRNK concentrations were both increased above baseline levels at PF-2 (Table 2, Figure S2). Importantly, FAK-pY397 concentration was decreased in crew member A at PF-1, but increased in B (Figure S2, Table 2). These findings suggest that musculoskeletal forces in space may have been comparable to those on Earth in crew member B, but substantially reduced in A.
Concentration of the meta-vinculin protein isoform and tenascin-C was increased at PF-1 in crew member A by 786 and 2781%, respectively, but both were slightly reduced in crew member B (Table 2, Figures S1 and S3). At PF-2, these proteins resumed to near baseline levels in crew member A, but were increased in crew member B (Figure S1). The concentration of gamma-vinculin was only marginally affected by spaceflight (Table 2).
Baseline meta-vinculin:gamma-vinculin ratio was comparable in crew members A and B. Crew member A depicted eightfold and fivefold increases at PF-1 and PF-2, respectively, but no changes were observed in B (Table S1). Baseline FRNK:FAK ratio was greater in crew member A than B (2.01 vs. 0.58). It remained unchanged in crew member A after spaceflight, but decreased moderately in B.
Skeletal muscle proteomic analysis
From 1100 spots detected in the two-dimensional difference in gel electrophoresis 2D-DIGE, 900 were included in the base set for statistical analysis. The principal component analysis (PCA) of the muscle tissue from both crew members yielded two components (PCA1 and PCA2) that explain 59.2 and 25.9% of the global variation, respectively (Figure S4). Concordant changes were observed for PCA2 from baseline to PF-1 in both crew members, but changes were discordant from baseline to PF-2, indicating closer similarity between PF-2 and baseline in crew member B.
Proteomic analyses followed by paired one-way ANOVA and Tukey tests (α = 0.01) indicated significant differences between baseline and PF-1 in 32 and 39 spots in crew members A and B, respectively. When comparing baseline to PF-2, 37 spots changed in crew member A and 24 spots in crew member B (Fig. 3 and Table S2).
Proteomic analysis in human skeletal muscle. Histograms of differential protein expression in soleus muscle between baseline vs. PF-1 (colored bars) and baseline vs. PF-2 (striped bars) in crew member A (green bars) and B (red bars), as detected by 2D DIGE analysis. Proteins significantly altered (paired one-way ANOVA and Tukey, α = 0.01) are indicated by their gene name and expressed as a percent of spot volume variation. a Contractile proteins; b Metabolism
Concerning contractile proteins, PF-1 samples revealed alterations in troponin I and troponin T in both crew members, but only crew member B revealed some moderate changes in actin and tropomyosin (Table S2). More specifically, crew member A demonstrated distinct increases in troponin I fast (TNNI2) and in two proteoforms of troponin T fast (TNNT3), and decreases in two proteoforms of troponin T slow (TNNT1). Notably, where PF-1 changes in the troponins were observed in B, they were in the opposite direction in crew member A. At PF-2, some moderate decreases in actin proteoforms (ACTA1) were observed in both crew members.
Proteins involved in anaerobic metabolism were also affected by spaceflight. Crew member A depicted decreases in four different proteoforms of glycogen phosphorylase (PYGM) at PF-1, as well as increases in glycerol-3-phosphate dehydrogenase (GPD1) and in one beta-enolase proteoform (ENO3). By contrast, B depicted increases in glyceraldehyde-3-phosphatase dehydrogenase (GAPDH), phosphoglycerate mutase 2 (PGAM2), alpha-enolase (ENO1), three proteoforms of ENO3 and pyruvate kinase (PKM). At PF-2, two of the PYGM proteoforms were recovered in crew member A, but decreases in two proteoforms of fructose-bisphosphate aldolase A (ALDOA) and l-lactate dehydrogenase A chain (LDHA) occurred at that time. Thus, dysregulation of glycogen metabolism and accumulation of specific enolase proteoforms appeared postflight in crew member A, whereas crew member B adapted metabolically to spaceflight by increasing anaerobic metabolism.
Concerning aerobic metabolism, enzymes involved in malate shuttle, in oxidative phosphorylation and in lipid metabolism were downregulated in both crew members at PF-1 and at PF-2. In crew member A, this encompassed virtually all enzymes, except that NADH dehydrogenase iron-sulfur protein 3 (NDUFS3) was upregulated. Crew member B depicted a very similar response as A, except that NDUFS3 was down- rather than upregulated (Table S1). Thus, aerobic metabolism seemed comparably compromised in both crew members.
Enzymes involved in high energy phosphate transfer likewise depicted a downregulation in both crew members postflight. This was more pronounced in crew member A than B. The enzymes affected involved four proteoforms of creatine kinase M-type (CKM) and creatine kinase S-type (CKMT2). Overall, dysregulation of high energy phosphate production could be a consequence of the severe impairment of aerobic metabolism, or vice versa.
Intracellular signaling pathways controlling muscle mass and metabolism
The two major catabolic systems were studied on mRNA level by assessing expression of MuRF-1 and atrogin-1 (markers of the ubiquitin proteasome activity) and of p62 and Beclin-1 (markers of autophagy). Whereas atrogin-1 expression was upregulated postflight in both crew members, MuRF-1 expression was highly upregulated in crew member A only. Beclin-1 was upregulated in both crew members postflight, whereas p62 was upregulated in crew member A only.
At PF-2, expression of all of these markers was lower than at PF-1. In crew member A, recovery towards normal activation was somewhat less complete, especially for MuRF-1. The results suggest a higher activation of both catabolic systems and a slower recovery towards normal values in crew member A.
PGC-1alpha is a master controller of mitochondrial biosynthesis and oxidative metabolism. It was surprisingly upregulated in both crew members at PF-1, and even at PF-2. SREBP-1 is a transcription factor controlling lipid synthesis. It was upregulated in crew member B only.
NRF2 is a transcription factor involved in the response to injury and inflammation, and thus also in sensing the level of intracellular ROS and stimulates synthesis of antioxidant defense systems. Its increase occurred in crew member B only, where it was still observed at PF-2.
Plasma metabolomics
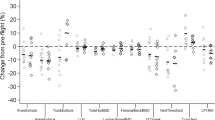
Twenty metabolites were assigned and quantified in the spectra. Levels of significant metabolites (plasma amino acids, glucose, lactate, pyruvate, including the pyruvate/lactate ratio) are reported along with their P values in Fig. 4, compared with a cohort of 79 control subjects. All baseline metabolite levels were comparable with control data, except that isoleucine was elevated and alanine was at the upper control margin at PF-1 in crew member A. Serum alanine was increased in this crew member postflight (in both PF-1 and PF-2 P < 0.01), but conversely from isoleucine was not normalized at PF-2. Moreover, crew member A depicted elevated serum levels of glucose and pyruvate, as well as an elevated pyruvate-lactate ratio on day PF-1 (all P < 0.01).
Metabolomic analysis. Panels a, b show amino acid and energetic metabolites, respectively, in box plot analysis and in arbitrary units concentrations (C.A.U.) of the most significant metabolites. Astronauts A and B are represented by red and green colors, respectively. Circle, square, and triangle represent baseline, PF-1 and PF-2, respectively. Gray circles represent the control cohort (79 volunteers). Significant (P < 0.01) and barely significant (P < 0.05) P values are also reported
In crew member B, serum levels were generally within the control margins. The only exceptions were alanine, phenylalanine, and tyrosine, which all increased at PF-1 (P < 0.01), but not at PF-2.
Discussion
The present study provides a multifaceted account of strongly diverging responses to spaceflight in two astronauts. While crew member A exhibited the expected neuromuscular responses, namely decrements in plantar flexor muscle volume, altered architecture, reductions in contractile protein composition, downgraded fiber contractility, and thus overall muscle strength, crew member B was much less affected after long-term spaceflight. Notably, the loading levels achieved by crew member B are rarely seen in other crew members on ISS. It is therefore tempting to ascribe the salient differences between A and B to the different muscular exercises, in particular with regards to resistive forces. One could therefore ascribe the divergent muscle proteomic responses (contractile proteins and aerobic metabolism affected in crew member A, but almost not in B), as well as differences in the response of the catabolic pathways (greater activation of ubiquitin proteasome system and of autophagy in crew member A), and in systemic metabolic profiles (greater effects upon energy and protein metabolism in A than in B) to the different countermeasure regimens. These findings therefore underline the importance of the onboard exercise. However, the 23% decrement in muscle fiber CSA coupled to the activation of catabolic pathways that was still occurring in crew member B, and the fact that aerobic metabolism and high phosphate energy transfer were also compromised in crew member B may suggest that complete preservation of the musculature in space is not yet achievable.
It is also intriguing that the response of the load-dependent parameters, pY397-FAK and meta-vinculin concentration, but not FAK or FRNK, reflect the diverging responses in muscle mass and strength.17,18 The observed downregulation of FAK and FRNK protein levels at PF-1 replicate earlier findings in antigravity muscles of rats and humans, implicating a requirement of gravitational loading to maintain the gene products of the FAK gene (i.e. PTK2).16,17,18 The present observations confirm that muscle response to unloading involves a net reduction in the capacity for regulation of costamere turnover via Y397 phosphorylated FAK.15 Consistent with this notion, the concentration of both FAK and FRNK increased 2 weeks after return to Earth, indicating reestablishment of adhesion sites in the sarcolemma with resumption of load-bearing muscle activity. Alterations in the proxy of costamere remodeling, FAK-pY397, were inversely related to alterations in the concentration of the costameric protein meta-vinculin and tenascin-C at PF-1 (Table 2; Figures S1−S3). At this time point, and in absolute terms at postflight session 2, expression of meta-vinculin and tenascin-C postflight was considerably less affected in crew member B, while FAK-pY397 concentration was selectively increased in this crew member. This is consistent with the reported upregulation of FAK-pY397 concentration by muscle loading and the greater extent of muscle loading of crew member B than A during the onboard exercise (Fig. 1).28,29 Collectively, the study of costamere-associated proteins suggests protection of the soleus muscle in crew member B inflight, but muscle damage due to unaccustomed mechanical loading after return to Earth.
Importantly, the adaptations in the FAK pathway, modulating the anabolic Akt/mTOR/p70S6K pathway, can link the unloading and muscle atrophy to the imbalance between protein synthesis and degradation.30,31
Interestingly, muscular aerobic metabolism was compromised in both crew members, notwithstanding the extensive treadmill exercise performed by crew member B and the enhanced expression of PGC-1alpha. PCG-1alpha enhances mitochondrial biosynthesis and oxidative metabolism and is a key regulator in the metabolic response to endurance training.32,33 In disuse, PGC-1alpha expression is reduced in soleus muscle of mice and in vastus lateralis muscle of humans.12,34 Reduced expression of PGC-1alpha has been considered a relevant cause of muscle atrophy, suggesting a “metabolic” program of muscle atrophy.11,12,34 The higher expression of PGC-1alpha in this study is surprising. Possibly, inflight countermeasures have prevented PGC-1alpha downregulation. Alternatively, PGC-1alpha upregulation could be an unsuccessful attempt to counteract the decrease in energy levels, namely an increase in AMP and a decrease in ATP due to impaired energy metabolism.
Expression of SREBP-1, a master regulator of lipid metabolism, was increased in the soleus muscle in both crew members after spaceflight. This is consistent with the observation that weightlessness induces intramuscular lipid accumulation.35 Furthermore, it is interesting that NRF2, a transcription factor sensitive to intracellular ROS and simultaneously a stimulator of antioxidant defense systems, was elevated in crew member B, but not in crew member A. Redox imbalance had been observed in human bed rest and in hindlimb suspended mice.12,34,36 However, as crew member B performed more exercise and had less muscle atrophy than A, the higher NRF2 expression in B could be due to an aerobically more intense exercise regimen in B than in A, inasmuch as oxygen demand in running increases with loading forces.37 Given that NRF2 and proteoforms related to mitochondrial and aerobic metabolism were divergently affected by space, one might thus even speculate about a relative overemphasis of aerobic exercise, similar to overtraining on Earth.
Both crew members diverged also in their metabolomic profiles. While postflight changes in crew member A were more pronounced for energy than for amino acid metabolism, the opposite was true for crew member B. Moreover, the increases in glucose, pyruvate, and pyruvate/lactate ratio in crew member A can be understood as the metabolic consequence of muscle wasting.38 Interestingly, both crew members’ metabolic profiles appeared distinct already before launch, as shown by PCA. During spaceflight, they responded in two different ways, moving toward different positions at PF-1, and finally both reverted to their initial metabolic spaces at PF-2 (Figure S5). This behavior is consistent with the observed metabolic resilience previously described in healthy controls.39
Collectively, these data show distinctly diverging responses in two crew members that performed countermeasure exercises at different intensities during their half-year sojourns on ISS, underlining the need of physical exercises in space. The findings also show that running as much as 500 km, with a resistive force close to the body weight, and performing 5000 heel raises at a force of more than twice the body weight was insufficient to entirely prevent muscle atrophy and weakness. Upregulation of the ubiquitin proteasome pathway and increased autophagy likely contributed to muscle wasting, more so in crew member A than B. Upregulation of PGC-1alpha was also found in both crew members, suggesting that the aerobic exercise performed onboard may have been sufficient to prevent disuse-induced decrease in PGC-1alpha, but not the impairment in metabolism and its impact on muscle integrity. The atrophy found both at whole muscle and single fiber level (mostly in crew member A) was associated with a reduction in FAK content and, in crew member A, also in phosphorylated FAK. Instead, in crew member B who exercised very intensively, FAK-pY397 levels actually increased. We have shown in previous ground-based studies that FAK plays an essential role in skeletal muscle remodeling with use, disuse and ageing and the present results confirm its role in regulating muscle mass.31,40,41,42 Adaptations of FAK pathway could, in fact, inhibit protein synthesis and contribute to muscle atrophy. Finally, proteomic assessment of muscle tissue and metabolomic blood analyses indicate that exercise may have benefits in space that go far beyond mere maintenance of muscle mass and strength.
Methods
Subjects and testing schedule
Two crew members of equal sex and comparable age, A and B, were tested before and after their half-year ISS mission (Table S3). The methods were performed in accordance with relevant guidelines and regulations and approved by the Human Research Multilateral Review board, and written informed consent was obtained prior to study inclusion. Inflight countermeasure exercises data were obtained via data sharing with NASA. Heel raise exercises on aRED were computed to yield the total number of repetitions, and peak and average resistive force per session was normalized to body weight. For treadmill data, distance, speed, exercise time, and the bungee force at rest were assessed.
Plantar flexor muscle function
Muscle strength was assessed using a custom-made dynamometer.43 Subjects were seated upright with hip and knee joints at 90°, performing maximum voluntary isometric contractions at ankle joint angles of −20, −10, 0, 10, and 20° (0° representing the neutral position, and positive angles indicating plantarflexed positions).
Muscle volume and architecture
Fiber length and pennation angle of the resting GM and SOL were evaluated with a MyLab25 ultrasonography scanner (Esaote Biomedica), using a 50 mm linear array probe LA523 (13-4 MHz). Scans were taken at 50% of the GM muscle length in the midsagittal plane at 90° knee flexion and −10° ankle dorsiflexion.
Muscle volume was assessed via transversal magnetic resonance images, with 3 mm slice thickness and 3 mm gaps. Before acquisition, subjects laid supine for 30 min.44 For analysis, muscles were segmented manually in every recorded slice.
Muscle biopsy
Approximately 50 mg of tissue were harvested from the soleus muscle with an 11G ACECUT automatic biopsy system (TSK Laboratory, Oisterwijk, the Netherlands). A lateral approach was chosen approximately 2 cm below the distal end of the lateral gastrocnemius muscle. Before incision, the skin was razed and disinfected, and local anesthetic (Lidocain 1%) was injected. Samples were divided and aliquoted within 10 min after the biopsy.
MHC isoform distribution was assessed as described.45,46,47 Single muscle fiber segments were dissolved in Laemmli solution and loaded on 6% SDS-PAGE polyacrylamide gels.48 To assess MHC isoform composition in whole biopsy, frozen portion of biopsy was pulverized with liquid nitrogen and resuspended in Laemmli solution. About 15 µg of proteins were loaded on 6% SDS-PAGE polyacrylamide gels, and electrophoresis was run overnight at 100 V and the gels were stained with Coomassie Blue stain.47,49
Single fiber mechanics
Fiber cross-sectional area (CSA), force, and maximum shortening velocity was assessed as previously described.45,50 Briefly, biopsy samples were divided in small bundles and stored at −20 °C for up to 3 weeks in skinning solution (150 mM K2HPO4, 5 mM KH2PO4, 5 mM magnesium acetate, 1 mM DTT, 5 mM EGTA, 3 mM Na2-ATP, pH 7, leupeptin hydrochloride 20 μg/ml, E64 10 μM) and glycerol 50%. On the day of experiment, segments of single fibers, approximately 1 mm long, were manually isolated under stereomicroscopic control at ×20–×40 magnification in skinning solution. Fibers were immersed for 1 h in skinning solution containing 0.1% Triton X-100 before functional analysis. Each fiber was mounted between a force transducer’s and an electromagnetic puller’s hook. Fiber width and depth were measured with an inverted microscope at ×320 magnification at three different positions along the fiber. CSA was determined assuming an elliptical shape. Isometric force (Po) and unloaded shortening velocity (Vo) were measured by the slack test technique. Activating solution had the following composition: 100 mM KCl, 20 mM imidazole, 5 mM MgCl2, 5 mM Na2-ATP, 0.5 mM EGTA, 25 mM creatine phosphate, 300 U/ml creatine kinase, pCa 8.0. Experiments were performed at 12 °C, in conditions of maximal activation (pCa 4.5) and at optimal sarcomere length (2.5 μm) for force development. At the end of the mechanical experiments, fibers were characterized on the basis of MHC isoform composition.
Costameric protein biochemistry
Biopsies were sectioned and protein was extracted with the help of a rotorstat mixer (Kinematica, Lucerne, Switzerland) using RIPA buffer, and total protein quantified essentially as described.18 Homogenate corresponding to 10 μg total protein was separated on 7.5% SDS-PAGE gels, blotted onto nitrocellulose membrane and subjected to immunological protein detection as described using the following antibodies: 1:1000 dilution of polycolonal FAK antiserum Lulu, 1:500 of monoclonal Tenascin-C serum B28:13, 1:100 of monoclonal antiserum against gamma-vinculin and meta-vinculin (gift from Dr. M.A. Glukhova, Paris, France).18,21,51 Detection of pY397-FAK content was carried out based on immune-precipitation of soluble proteins in homogenate corresponding to 1 mg total protein essentially as described.18 Samples being derived from the three samples of each subject were analyzed in adjacent lanes of the same SDS-PAGE gel. Equal loading and blotting was verified via signal intensity of the actin band on the Ponceau S-stained nitrocellulose membrane; after blotting signal intensity was assessed from background-corrected band intensities using PxI system (Syngene) as described.52 All blots derived from the same experiment and were processed in parallel.
RT-PCR analysis was performed as described.53 Total RNA from muscle samples was extracted using the Promega SV Total RNA isolation kit. Three hundred nanogram of RNA were reverse-transcribed with SuperScript III reverse transcriptase (Life Technologies) to obtain cDNA. The cDNA was analyzed by RT-PCR (see Table S4) using SYBR Green PCR master mix (Life Technologies). Data were normalized to β2-microglobulin expression as housekeeping gene.
Skeletal muscle proteomics
Protein extraction
For two-dimensional difference in gel electrophoresis (2D-DIGE) and immunoblot assays, each sample from each subject was suspended in lysis buffer (7 M urea, 2 M thiourea, 4% CHAPS, 30 mM Tris, and 1 mM PMSF) and solubilized by sonication on ice. Proteins were selectively precipitated using PlusOne 2D-Clean up Kit (GE Healthcare, Little Chalfont, UK) in order to remove nonprotein impurities, and resuspended in lysis buffer. The pH of the protein extracts was adjusted to pH 8.5 by addition of 1 M NaOH. Protein concentrations were determined by PlusOne 2D-Quant Kit (GE Healthcare).
2D-DIGE
Protein minimal labeling with cyanine dyes (Cy3 and Cy5), 2D separation, and analyses were performed as described previously (for crew members A and B; SOL biopsy preflight, immediately postflight and 15 days postflight).20 Briefly, proteins extracted (50 µg) from each individual were labeled with Cy5, while internal standards were generated by pooling (50 µg) individual samples that were Cy3-labeled. Samples were separated on 3–10 nonlinear immobilized pH gradient (IPG) strips; the adopted gradient enables separation of protein isoforms in the first dimension, providing a detailed pattern of the muscle proteome. Each individual sample was run in triplicate (analytical replicates) to minimize intergel variability. Image analysis was performed using DeCyder 7.0 software (GE Healthcare). Statistical analysis was performed using the DeCyder 1.0 extended data analysis (EDA) module. Protein filters were set to select only those protein spots that matched > 90% of the gel images, and these protein spots were included in data analysis. Statistically significant differences of 2D-DIGE data were computed by paired one-way ANOVA (two-sided) coupled to Tukey’s multiple group comparison test; the significance level was set at P < 0.01. In addition, the false discovery rate (FDR) was applied as a multiple testing correction method to keep the overall error rate as low as possible.54 Two independent analyses were performed. Proteins of interest were identified by mass spectrometry.
Protein identification by mass spectrometry
Proteins were identified by peptide mass fingerprinting (PMF) utilizing a matrix-assisted laser desorption/ionization (MALDI) time-of-flight (ToF) mass spectrometer (Ultraflex III ToF/ToF; Bruker Daltonics, Bremen, Germany), as previously described.55 In particular, search was carried out by correlation of uninterpreted spectra to Mammalia entries in NCBInr 20090430 database (8 483 808 sequences; 2 914 572 939 residues). Where this approach was unsuccessful, additional searches were performed using electrospray ionization-MS/MS, as previously described.56
Metabolomics
Blood/plasma samples were obtained both from astronauts and from a control cohort of 79 healthy volunteers. Ethylenediaminetetraacetic acid was always used as anticoagulant, but the effects of its presence were eliminated as previously described.57 In particular, 56 donors were collected in collaboration with the Tuscanian section of the Italian Association of Blood Donors (AVIS) in the Transfusion Service of Pistoia Hospital (Ospedale del Ceppo, AUSL 3—Pistoia, Italy), and 23 samples were collected in Bologna. Specifically, six healthy volunteers were recruited at four different times (up to 7 months, one dropped out at a specific time). No differences in the time series were found (data not shown) and all samples were merged to obtain a more heterogeneous population.
Frozen plasma samples were thawed at room temperature and shaken before use. The NMR samples were prepared according to the standard operating procedures.58 1H-NMR spectra were acquired at 310 K using a Bruker 600 MHz spectrometer (Bruker BioSpin): water suppressed Carr–Purcell–Meiboom–Gill (CPMG) spin echo pulse sequence (RD-90°-(τ-180°-τ)n-acq) to obtain one-dimensional 1H-NMR spectra in which broad signals from high molecular weight metabolites (i.e. proteins and lipoproteins) are attenuated.59 Sixty-four FIDs were collected into 73 728 data points over a spectral width of 12 019 Hz, with a relaxation delay of 4 s and acquisition time of 3.1 s. Free induction decays were multiplied by an exponential function equivalent to a 1.0 Hz line-broadening factor before applying Fourier transformation. Transformed spectra were automatically corrected for phase and baseline distortions and calibrated (anomeric glucose doublet at 5.24 ppm) using TopSpin 3.2 (Bruker Biospin GmbH, Germany).
All metabolomic data analyses were performed using R.60 The spectral regions related to metabolites were assigned in the 1H-NMR profiles by using matching routines of AMIX 3.8.4 in combination with the BBIOREFCODE (Bruker BioSpin GmbH, Germany), and freely available datasets.61,62 Metabolite concentrations in arbitrary units were obtained integrating the related spectral regions. PCA was used as a first unsupervised exploratory analysis to compare metabolic profiles of astronauts and healthy volunteers, and to enable the assessment of the homogeneity or the presence of any outliers in plasma samples of volunteers that come from two different collection centers. The concentrations of each metabolite in the astronauts samples were compared with the healthy volunteers distributions using the Iglewicz and Hoaglin outlier test.63 P values < 0.01 were deemed significant and P values < 0.05 near-significant.
Data availability
The datasets generated and analyzed during the current study are not publicly available due to privacy reasons but are available from the corresponding author on reasonable request.
Change history
24 October 2018
The original version of this Article contained an error in the spelling of the author Claudio Franceschi, which was incorrectly given as Claudio Francheschi. This has now been corrected in both the PDF and HTML versions of the Article.
References
Convertino, V. A. Physiological adaptations to weightlessness: effects on exercise and work performance. Exerc. Sport Sci. Rev. 18, 166 (1990).
Narici, M., Kayser, B., Barattini, P. & Cerretelli, P. Effects of 17-day spaceflight on electrically evoked torque and cross-sectional area of the human triceps surae. Eur. J. Appl. Physiol. 90, 275–282 (2003).
Fitts, R. H., Riley, D. R. & Widrick, J. J. Physiology of a microgravity environment invited review: microgravity and skeletal muscle. J. Appl. Physiol. (1985) 89, 823–839 (2000).
Fitts, R. H. et al. Prolonged space flight-induced alterations in the structure and function of human skeletal muscle fibres. J. Physiol. 588, 3567–3592 (2010).
de Boer, M. D. et al. Effect of 5 weeks horizontal bed rest on human muscle thickness and architecture of weight bearing and non-weight bearing muscles. Eur. J. Appl. Physiol. 104, 401–407 (2008).
de Boer, M. D., Maganaris, C. N., Seynnes, O. R., Rennie, M. J. & Narici, M. V. Time course of muscular, neural and tendinous adaptations to 23 day unilateral lower-limb suspension in young men. J. Physiol. 583, 1079–1091 (2007).
Wilson, A. & Lichtwark, G. The anatomical arrangement of muscle and tendon enhances limb versatility and locomotor performance. Philos. Trans. R. Soc. Lond. Ser. B, Biol. Sci. 366, 1540–1553 (2011).
Hvid, L. G. et al. Myosin content of single muscle fibers following short-term disuse and active recovery in young and old healthy men. Exp. Gerontol. 87, 100–107 (2017).
Wilkinson, S. B. et al. Differential effects of resistance and endurance exercise in the fed state on signalling molecule phosphorylation and protein synthesis in human muscle. J. Physiol. 586, 3701–3017 (2008).
Rudrappa, S. S. et al. Human skeletal muscle disuse atrophy: effects on muscle protein synthesis, breakdown, and insulin resistance—a qualitative review. Front. Physiol. 7, 361 (2016).
Cannavino, J. et al. The role of alterations in mitochondrial dynamics and PGC-1alpha over-expression in fast muscle atrophy following hindlimb unloading. J. Physiol. 593, 1981–1995 (2015).
Cannavino, J., Brocca, L., Sandri, M., Bottinelli, R. & Pellegrino, M. A. PGC1-alpha over-expression prevents metabolic alterations and soleus muscle atrophy in hindlimb unloaded mice. J. Physiol. 592, 4575–4589 (2014).
Hoppeler, H., Baum, O., Lurman, G. & Mueller, M. Molecular mechanisms of muscle plasticity with exercise. Compr. Physiol. 1, 1383–1412 (2011).
Chopard, A., Pons, F. & Marini, J. F. Vinculin and meta-vinculin in fast and slow rat skeletal muscle before and after hindlimb suspension. Pflug. Arch. 444, 627–633 (2002).
Chopard, A., Arrighi, N., Carnino, A. & Marini, J. F. Changes in dysferlin, proteins from dystrophin glycoprotein complex, costameres, and cytoskeleton in human soleus and vastus lateralis muscles after a long-term bedrest with or without exercise. Faseb J. 19, 1722–1724 (2005).
Salanova, M. et al. Disuse deterioration of human skeletal muscle challenged by resistive exercise superimposed with vibration: evidence from structural and proteomic analysis. Faseb J. 28, 4748–4763 (2014).
Li, R. et al. Costamere remodeling with muscle loading and unloading in healthy young men. J. Anat. 223, 525–536 (2013).
Klossner, S., Li, R., Ruoss, S., Durieux, A. C. & Fluck, M. Quantitative changes in focal adhesion kinase and its inhibitor, FRNK, drive load-dependent expression of costamere components. Am. J. Physiol. Regul. Integr. Comp. Physiol. 305, R647–R657 (2013).
Rindom, E. & Vissing, K. Mechanosensitive molecular networks involved in transducing resistance exercise-signals into muscle protein accretion. Front Physiol. 7, 547 (2016).
Moriggi, M. et al. Long term bed rest with and without vibration exercise countermeasures: effects on human muscle protein dysregulation. Proteomics 10, 3756–3774 (2010).
Fluck, M., Ziemiecki, A., Billeter, R. & Muntener, M. Fibre-type specific concentration of focal adhesion kinase at the sarcolemma: influence of fibre innervation and regeneration. J. Exp. Biol. 205, 2337–2348 (2002).
Samarel, A. M. Costameres, focal adhesions, and cardiomyocyte mechanotransduction. Am. J. Physiol. Heart Circ. Physiol. 289, H2291–H2301 (2005).
Grounds, M. D., Sorokin, L. & White, J. Strength at the extracellular matrix-muscle interface. Scand. J. Med Sci. Sports 15, 381–391 (2005).
Miyamoto, S. et al. Integrin function: molecular hierarchies of cytoskeletal and signaling molecules. J. Cell Biol. 131, 791–805 (1995).
Schlaepfer, D. D., Mitra, S. K. & Ilic, D. Control of motile and invasive cell phenotypes by focal adhesion kinase. Biochim. Biophys. Acta 1692, 77–102 (2004).
Li, S. et al. Fluid shear stress activation of focal adhesion kinase. Linking to mitogen-activated protein kinases. J. Biol. Chem. 272, 30455–30462 (1997).
Durieux, A. C. et al. Focal adhesion kinase is a load-dependent governor of the slow contractile and oxidative muscle phenotype. J. Physiol. 587, 3703–3717 (2009).
Fluck, M., Chiquet, M., Schmutz, S., Mayet-Sornay, M. H. & Desplanches, D. Reloading of atrophied rat soleus muscle induces tenascin-C expression around damaged muscle fibers. Am. J. Physiol. Regul. Integr. Comp. Physiol. 284, R792–R801 (2003).
Jarvinen, T. A. et al. Mechanical loading regulates the expression of tenascin-C in the myotendinous junction and tendon but does not induce de novo synthesis in the skeletal muscle. J. Cell Sci. 116, 857–866 (2003).
Klossner, S., Durieux, A. C., Freyssenet, D. & Flueck, M. Mechano-transduction to muscle protein synthesis is modulated by FAK. Eur. J. Appl. Physiol. 106, 389–398 (2009).
de Boer, M. D. et al. The temporal responses of protein synthesis, gene expression and cell signalling in human quadriceps muscle and patellar tendon to disuse. J. Physiol. 585, 241–251 (2007).
Lin, J., Handschin, C. & Spiegelman, B. M. Metabolic control through the PGC-1 family of transcription coactivators. Cell Metab. 1, 361–370 (2005).
Russell, A. P. et al. Endurance training in humans leads to fiber type-specific increases in levels of peroxisome proliferator-activated receptor-gamma coactivator-1 and peroxisome proliferator-activated receptor-alpha in skeletal muscle. Diabetes 52, 2874–2881 (2003).
Brocca, L. et al. The time course of the adaptations of human muscle proteome to bed rest and the underlying mechanisms. J. Physiol. 590, 5211–5230 (2012).
Bergouignan, A., Rudwill, F., Simon, C. & Blanc, S. Physical inactivity as the culprit of metabolic inflexibility: evidence from bed-rest studies. J. Appl. Physiol. (1985) 111, 1201–1210 (2011).
Dalla Libera, L. et al. A transient antioxidant stress response accompanies the onset of disuse atrophy in human skeletal muscle. J. Appl. Physiol. (1985) 107, 549–557 (2009).
Cavagna, G. A., Willems, P. A. & Heglund, N. C. The role of gravity in human walking: pendular energy exchange, external work and optimal speed. J. Physiol. 528, 657 (2000).
Stein, T. P. & Wade, C. E. Metabolic consequences of muscle disuse atrophy. J. Nutr. 135, 1824S–1828S (2005).
Ghini, V., Saccenti, E., Tenori, L., Assfalg, M. & Luchinat, C. Allostasis and resilience of the human individual metabolic phenotype. J. Proteome Res. 14, 2951–2962 (2015).
Fluck, M., Carson, J. A., Gordon, S. E., Ziemiecki, A. & Booth, F. W. Focal adhesion proteins FAK and paxillin increase in hypertrophied skeletal muscle. Am. J. Physiol. 277, C152–C162 (1999).
Li, R. et al. Costamere remodeling with muscle loading and unloading in healthy young men. J. Anat. 223, 525–536 (2013).
Narici, M. V. et al. Skeletal muscle remodeling in response to alpine skiing training in older individuals. Scand. J. Med. Sci. Sports 21(Suppl 1), 23–28 (2011).
Reeves, N. D., Maganaris, C. N., Ferretti, G. & Narici, M. V. Influence of 90-day simulated microgravity on human tendon mechanical properties and the effect of resistive countermeasures. J. Appl. Physiol. (1985) 98, 2278–2286 (2005).
Albracht, K., Arampatzis, A. & Baltzopoulos, V. Assessment of muscle volume and physiological cross-sectional area of the human triceps surae muscle in vivo. J. Biomech. 41, 2211–2218 (2008).
Bottinelli, R., Canepari, M., Pellegrino, M. A. & Reggiani, C. Force-velocity properties of human skeletal muscle fibres: myosin heavy chain isoform and temperature dependence. J. Physiol. 495(Pt 2), 573–586 (1996).
D’Antona, G. et al. The effect of ageing and immobilization on structure and function of human skeletal muscle fibres. J. Physiol. 552, 499–511 (2003).
Pellegrino, M. A. et al. Redox homeostasis, oxidative stress and disuse muscle atrophy. J. Physiol. 589, 2147–2160 (2011).
Soriano, F. X. et al. Evidence for a mitochondrial regulatory pathway defined by peroxisome proliferator-activated receptor-gamma coactivator-1 alpha, estrogen-related receptor-alpha, and mitofusin 2. Diabetes 55, 1783–1791 (2006).
Pellegrino, M. A. et al. Orthologous myosin isoforms and scaling of shortening velocity with body size in mouse, rat, rabbit and human muscles. J. Physiol. 546, 677–689 (2003).
Brocca, L. et al. Structure and function of human muscle fibres and muscle proteome in physically active older men. J. Physiol. 595, 4823–4844 (2017).
Brellier, F. et al. Tenascin-W is a better cancer biomarker than tenascin-C for most human solid tumors. BMC Clin. Pathol. 12, 14 (2012).
Fluck, M. et al. Genomic and lipidomic actions of nandrolone on detached rotator cuff muscle in sheep. J. Steroid Biochem. Mol. Biol. 165, 382–395 (2017).
Brocca, L. et al. Human skeletal muscle fibre contractile properties and proteomic profile: adaptations to 3 weeks of unilateral lower limb suspension and active recovery. J. Physiol. 593, 5361–5385 (2015).
Hochberg, A. M. et al. Sensitivity and specificity of the test kit BAX for screening/E. coli O157:H7 in ground beef: independent laboratory study. J. AOAC Int. 83, 1349–1356 (2000).
Vigano, A. et al. Protein modulation in mouse heart under acute and chronic hypoxia. Proteomics 11, 4202–4217 (2011).
Capitanio, D. et al. Comparative proteomic profile of rat sciatic nerve and gastrocnemius muscle tissues in ageing by 2-D DIGE. Proteomics 9, 2004–2020 (2009).
Bernini, P., Bertini, I., Luchinat, C., Tenori, L. & Tognaccini, A. The cardiovascular risk of healthy individuals studied by NMR metabonomics of plasma samples. J. Proteome Res. 10, 4983–4992 (2011).
Bernini, P. et al. Standard operating procedures for pre-analytical handling of blood and urine for metabolomic studies and biobanks. J. Biomol. NMR 49, 231–243 (2011).
Meiboom, S. & Gill, D. Modified spin-echo method for measuring nuclear relaxation times. Rev. Sci. Instrum. 29, 688–691 (1958).
Ihaka, R. & Gentleman, R. R: a language for data analysis and graphics. J. Comput. Graph. Stat. 5, 299–314 (1996).
Wishart, D. S. et al. HMDB: the Human Metabolome Database. Nucleic Acids Res. 35, D521–D526 (2007).
Bouatra, S. et al. The human urine metabolome. PLoS ONE 8, e73076 (2013).
Iglewicz, B. & Hoaglin, D. C. How to Detect and Handle Outliers Vol. 16 (Asq Press, Universtity of California 1993).
Acknowledgements
The study has been supported by the European Space Agency, with great help from Patrik Sundblad, Simone Thomas, and Marine le-Gouic. Peter Gauger and Wolfram Sies have been instrumental in preparation of human testing devices. Asa Beijer has helped with preprocessing of the muscle samples. The study was supported by internal funding from DLR (cost object 2475 115 “effects of microG”). The study was supported by RFO (Ricerca Fondamentale Orientata) funds to M.C. K.A. received a research grant from the German Space Agency (50WB1728). This work was supported by internal funding from the authors’ institutions.
Author information
Authors and Affiliations
Contributions
R.B., P.C., and M.N. designed the study. Human physiology experiments were performed and data were analysed by J.R., K.A., O.S., I.D.G., and M.N., and testing sessions were organized by J.R. Laboratory analyses were performed by M.F., S.R., L.B., E.L., M.M., L.T., A.V., M.C., C.G, C.L., and C.F. J.R., R.B. and M.N. wrote the manuscript with input and editing from all authors.
Corresponding author
Ethics declarations
Competing interests
Martin Flück reports personal fees and others from Resortho Foundation, grants and personal fees from Swiss National Science Foundation, grants and personal fees from Balgrist Foundation, outside the submitted work. In addition, Dr. Flück has a patent on Regulation of muscle repair (ÐS 20090280103 A1 pending to Martin Flück). All other authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rittweger, J., Albracht, K., Flück, M. et al. Sarcolab pilot study into skeletal muscle’s adaptation to long-term spaceflight. npj Microgravity 4, 18 (2018). https://doi.org/10.1038/s41526-018-0052-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41526-018-0052-1
This article is cited by
-
Repeated Horizontal Jumping is a Feasible Exercise Countermeasure for Microgravity
Microgravity Science and Technology (2022)
-
Contractile behavior of the gastrocnemius medialis muscle during running in simulated hypogravity
npj Microgravity (2021)
-
Effects of microgravity exposure and fructo-oligosaccharide ingestion on the proteome of soleus and extensor digitorum longus muscles in developing mice
npj Microgravity (2021)
-
Daily 30-min exposure to artificial gravity during 60 days of bed rest does not maintain aerobic exercise capacity but mitigates some deteriorations of muscle function: results from the AGBRESA RCT
European Journal of Applied Physiology (2021)
-
Effects of 21 days of bed rest and whey protein supplementation on plantar flexor muscle fatigue resistance during repeated shortening contractions
European Journal of Applied Physiology (2020)