Abstract
A fast, robust pipeline for strain mapping of crystalline materials is important for many technological applications. Scanning electron nanodiffraction allows us to calculate strain maps with high accuracy and spatial resolutions, but this technique is limited when the electron beam undergoes multiple scattering. Deep-learning methods have the potential to invert these complex signals, but require a large number of training examples. We implement a Fourier space, complex-valued deep-neural network, FCU-Net, to invert highly nonlinear electron diffraction patterns into the corresponding quantitative structure factor images. FCU-Net was trained using over 200,000 unique simulated dynamical diffraction patterns from different combinations of crystal structures, orientations, thicknesses, and microscope parameters, which are augmented with experimental artifacts. We evaluated FCU-Net against simulated and experimental datasets, where it substantially outperforms conventional analysis methods. Our code, models, and training library are open-source and may be adapted to different diffraction measurement problems.
Similar content being viewed by others
Introduction
Scanning transmission electron microscopy (STEM) has emerged as one of the primary nanoscale materials characterization tools1. A STEM experiment focuses an electron beam on to a sample, with the probe dimensions ranging from tens of nanometers down to the atomic scale, which is made possible by hardware aberration correction2,3. STEM experiments have successfully measured the 2D position of atomic columns with picometer-precision4, measured the vibrational spectra of single-atom defects5, mapped solid-liquid interfaces in lithium-metal batteries6, and determined the 3D position and chemical species of each atom in a nanoparticle7. Atomic-resolution STEM methods provide extremely high resolution for both spatial and spectroscopic mapping, but have a limited field of view (FOV) because of the necessary minimum sampling rate required to resolve atoms8.
An alternative to real space imaging in STEM is to instead record a converged beam electron diffraction (CBED) pattern at each probe position, resulting in a four-dimensional (4D-STEM) dataset9. 4D-STEM experiments are gaining popularity among electron microscopists because they can collect atomic-scale information from each probe over a nearly arbitrary sized field-of-view10, and can measure a broad spectrum of quantities of physical interest including: 3D structural determination11, ferroelectric polarization12, imaging of lithium in cathode materials13, ptychographic atomic imaging14, correlation of local strain with composition from X-ray ptychography15,16, distinguishing between chemical and structural interfacial roughness17, strain in 2D material bilayers18,19, and many others. The ability to extract quantitative information with atomic-scale resolution is, however, frequently limited by the size and complexity of experimental 4D-STEM data. Open source computational tools such as pyxem in hyperSpy20, liberTEM21, AtomAI22, and py4DSTEM23 provide high-throughput multimodal data analysis tools to the community.
Computational analysis of diffraction images from crystalline materials typically begins with localizing any Bragg scattering. A standard approach to this problem is matching a template—usually an image of the electron beam over vacuum—to each diffraction pattern using cross-correlation.
However, the Bragg disk intensities can oscillate with changing sample thickness, bias asymmetrically due to mistilt of the crystal zone axis relative to the electron beam, form interference effects between overlapping disks, and generally display highly nonlinear signals in all but the very thinnest of samples due to dynamical/multiple scattering24,25,26,27. While the physics of these phenomena are understood and the effects may be readily recognizable to a human observer, writing classical algorithms, which can accommodate them is challenging. These effects lead to uneven illumination of the Bragg disks, and consequently could cause errors in position-finding algorithms. Various approaches have been implemented, including cross, phase, and hybrid correlations28, edge filtering29, circular Hough transforms30, and radial gradient maximization31. Zeltmann et al. fabricated patterned apertures, which result in bullseye shaped electron probes that improve the precision of disk position measurements27. Other authors use Fourier-space methods to pool information about the disk spacing, such as the cepstral transform32.
In addition to the challenge of accuracy, traditional approaches often require careful parameter tuning to achieve acceptable results, and may be time consuming33. Moreover, the quantity one is ideally after is not just the disk positions but the structure factors Vg, the positions and amplitudes of which reflect the reciprocal lattice of the scattering crystal.
Once the Bragg disks have been measured, many subsequent analyses become possible, including crystallographic orientation mapping, off-axis virtual imaging modalities, and mapping the local strain9,28,34,35,36. Spatially resolved strain maps of crystalline and semi-crystalline materials systems are important in various engineering and technological applications. For instance, local strain distortions can play an important role in tuning electronic properties of semiconductors37,38, and lattice deformation and distortions due to defects and doping can be characterized from localized strain maps in metals39,40,41.
Artificial intelligence and machine learning (AI/ML) algorithms are increasingly being implemented in materials characterization, including in electron microscopy42. Deep-learning approaches have been been demonstrated to outperform classical algorithms in variety of computer vision problems in microscopy including classification and segmentation problems43,44,45. For instance, deep-convolutional neural networks (CNNs) are implemented in the analysis of images collected with various microscopy techniques such as crystal phase classification from back-scattered diffraction patterns46, structure measurement from electron diffraction and atomic-resolution STEM images47 and from scanning tunneling microscopy48, crystal symmetry identification from X-ray diffraction49, defect analysis from atomic-resolution STEM images50, crystal tilt and thickness detection from position averaged CBED patterns51,52, and orientation and strain mapping from 4D-STEM diffraction datasets53,54. Recently, Yuan et al. demonstrated the possibility of using CNNs to predict high precision orientation and strain maps of crystalline systems using 4D-STEM data, computing strain in field effect transistors with both a CNN and a more traditional Hough transform approach54. Li et al. used manifold learning to directly classify different features in 4D-STEM data53. Similarly, Shi et al. used an unsupervised method to analyze lattice deformations, and classify the resulting material properties such as strain from 4D-STEM datasets55. These works show the potential of both supervised and unsupervised learning (with and without knowledge of the ground truth, respectively) in the analysis of 4D-STEM datasets and motivated towards achieving automated analysis of massive 4D diffraction datasets.
Bragg disk position and the underlying strain field measurement of crystalline and semi-crystalline samples, leveraging supervised machine learning, can be considered as pixel-wise mapping of diffracted disk intensities to the underlying structure factors. Such tasks may be accomplished, for example, by a traditional U-Net architecture consisting of symmetric contracting (encoder) and expansive (decoder) paths, with the crucial addition of skip layer connections enabling the flow of localized contextual information from low-resolution encoded features to higher resolution upsampled layers56. However, while the U-Net seems to be a prudent choice for the Bragg disk measurement problem, using traditional 2D convolutional layers for the network building blocks poses a challenge: for identical samples, changing microscope parameters, such as the probe semiangle, can substantially change the measured diffraction images. We require a method to encode these changing experimental parameters into the signal inversion, which is not possible in the original U-Net architecture. Additionally, small shifts of the disks can be measured using cross-correlation of a probe template, but this signal is most accurately measured as the phase component of the complex-valued Fourier transform of the correlation. To preserve all the relevant signal including the complex phase, we implement a modified U-Net architecture using fully complex 2D convolutional blocks. Historically, complex representations of images and signals have numerous advantages and outperform their non-complex equivalent forms57,58,59,60.
The complex representation is an elegant method to preserve phase information and mimics biological behavior in neurons61. Rippel et al. implemented a Fourier representation of traditional CNNs by parameterizing convolutional kernels in the spectral domain62. In a recent effort, Trabelsi et al. provided building blocks for deep-complex-valued convolution networks and implemented their network on a variety of deep-learning tasks such as image classification, image recognition, and music and speech transcription problems63. Here, we extend these approaches to modify the U-Net architecture to accommodate the complex and nonlinear correlation between the CBED images and the structure factors.
In this work, we implement a Fourier-space complex U-Net (FCU-Net) deep-neural network, which learns the mapping from measured diffraction pattern intensities to a material’s underlying structure factors (Fig. 1). We train our network on a dataset with over 200,000 unique simulated dynamical CBED data spanning thousands of crystal systems with a variety of random zone axes, off-zone tilts, thicknesses, and microscope parameters. The training datasets are extended with physics-informed image augmentation through the addition of a realistic background, noise, and geometric distortions of the CBED patterns. We compare the accuracy of the FCU-Net outputs to the approach of cross-correlation template matching, benchmarking against the ground truth structure factors for simulated data. We further test and compare these two methods by measuring local strain using the structure factor outputs, for both simulated and experimental diffraction data of a SiGe multilayer stack, and with experimental hexagonal-boron nitride 4D-STEM data. We find that FCU-Net significantly improves the accuracy of disk detection, as well as downstream measurements such as strain. The FCU-Net pipeline is fast, highly automated, performant on materials and microscope parameters on which it has not been trained, and is robust against both experimental error and background noise.
a Multislice diffraction simulations of many samples with different crystal structures, compositions, orientations, and thicknesses, using various microscope parameters. b Augmentation of the simulated images by applying elliptic distortion, pattern shift, limited signal-to-noise, and background functions. c Deep-learning training. d Experimental geometry for diffraction pattern measurements. e Dataset preprocessing. f Inversion of experimental diffraction images to predict the structure factors using the FCU-Net trained in c.
Results and discussion
Comparison of traditional and complex U-Net models
To measure the position of Bragg disks from diffraction patterns, we implement supervised learning on a large training dataset consisting of simulated CBED images and structure factor images. To map disk intensities to the structure factors, we implement three variants of CNN architecture: real-valued U-Net, a U-Net with spectral parameterization, and the fully complex variant, FCU-Net. Figure 1 summarizes the overview of this work, where Fig. 1a–c show the methods we use to train the machine learning models from the simulated STEM diffraction pattern and the underlying structure factors. Figure 1d–f show the inference stage to predict structure factors from experimental diffraction patterns. The computational methods implemented to simulate training data, architecture of the CNN models implemented in this work, the training process, and implementation and inference from experimental diffraction patterns can be found in the Methods 4 section.
Once the networks are trained, we predict the structure factors of diffraction patterns from the simulated test dataset and used them to compute the structural similarity index (SSIM), a metric of image similarity measurement64. Table 1 compares the results for different CNN models. We find a significant improvement in the SSIM scores measured on the test dataset for the FCU-Net model, compared to networks without spectral pooling and/or without complex convolutional layers. The improvement in the overall model efficiency for the high-tilt, off-zone samples is more prominent than in the untilted, on-zone samples. We attribute this to the sensitivity of FCU-Net to the phase component of the input signal, as we expect the contribution of the phase to be more significant for high-tilt samples due to the asymmetry of their diffraction images.
Accuracy of diffracted disk position measurements
To evaluate the accuracy of Bragg disk detection using the trained FCU-Net and using cross-correlation, we calculate the intensity-weighted accuracy of the disk locations determined by each method, using the simulated test dataset with different crystal orientations and in-plane rotations. The intensity-weighted accuracy is defined as
where,
TPint, FPint, FNint denote intensity-weighted true-positive peaks, false-positive peaks and false negative peaks detected, respectively, from the predicted structure factor images. We note that the CBED and the structure factor images in our training dataset were generated with a pixel size of 0.0217 Å−1. To measure the intensity-weighted accuracy and the three metrics—TPInt, FPInt, FNInt for predicted structure factor, we use a threshold size of 0.05 Å−1 to match peaks between the predicted and ground truth structure factor images, in order of peak pair distance. Several example diffraction images, sampled randomly from the test dataset, are shown in Fig. 2a. The corresponding computed and ground truth disk positions and amplitudes are shown in Fig. 2b, c, using cross-correlation and our trained FCU-Net, respectively. The accuracy of disk detection using the FCU-Net is significantly better than the correlation-based approach across the board, with the most striking gains occurring in diffraction patterns, which suffer from multiple scattering due to large thickness, or disk overlap when the scattering vectors are small compared to the probe semiangle.
a Examples of simulated diffraction patterns for crystals of different thicknesses and orientations. Scale bar is 0.5 Å−1. b, c The positions of the ground truth structure factor coefficients of the crystal lattice are plotted below as blue circles, with a size proportional to the structure factor amplitudes Vg. The structure factor positions were computed using (b) template matching by cross-correlation with the vacuum probe signal, and (c) the FCU-Net network. Both measurements are overlaid as black crosses, with a size proportion to the estimated disk amplitude (square root of the disk intensity) and Vg amplitudes, for the correlation and FCU-Net predictions, respectively. The total intensity-weighted accuracy is listed above for all measurements.
The leftmost diffraction pattern in Fig. 2a is comparatively simple, with well separated, flat disks and signal well about the background level. Unsurprisingly, both methods do very well. However even here, in this nearly optimal data for cross-correlative template matching, the gains using FCU-Net are remarkable, achieving 100% accuracy. In the middle three patterns, the background signal and disk overlap make visual identification of the disk positions difficult. It is thus again unsurprising that cross-correlation does relatively poorly. In contrast, FCU-Net is extremely accurate for these three cases. The fifth diffraction image in Fig. 2a is an example of an experiment where the sample, which has been tilted away from the low-index zone axis relative to the beam direction, creating complex variation in disk intensities due to tilt of the Ewald sphere. FCU-Net still outperforms cross-correlation in this case, though the gains here are more modest.
We further evaluated the performance of FCU-Net and correlation methods for strain mapping by applying them to 415 unique crystals and orientations in our simulated dataset. These simulations were selected because they produced diffraction patterns with at least two strongly excited non-orthogonal Bragg vectors (i.e., the diffraction pattern was 2D rather than 1D), and had enough separation between the diffraction spots to automatically detect the ground truth lattice from the structure factor images, which was determined by applying a threshold of 5% to the mean strain error. To avoid the introduction of biases, we used a single set of parameters, which generalized well across the entire dataset, rather than tailoring to a specific diffraction pattern series. The performance of each method was evaluated by calculating the mean absolute value of the two principal strains. We initially analyzed low-index zone axis and randomly oriented crystals separately, but found negligible difference between the two. After disk detection and lattice assignment, we calculated the relative strain between the disk positions measured in the diffraction patterns and the positions measured from the structure factor images.
The median of the strain error as a function of sample thickness from 2 to 50 nm is shown in Fig. 3a. We also show the 25th and 75th percentile range. FCU-Net outperforms the correlation method at every thickness, showing an improvement of ~2–3 times across the thickness series. FCU-Net performs best at 20 nm thickness, but remains fairly flat with comparatively small interquartile range for all thicknesses. By contrast, the correlation method performs best at 4 nm and increases with sample thickness, with a much larger interquartile range. For very thin samples (<10 nm), the performance of the correlation method approaches that of FCU-Net, but never surpasses it. We attribute the higher accuracy at low thicknesses to the scattering being more kinematical (less intensity variation in the diffracted disks). Both methods show higher error at 2 nm thickness, which we attribute to the weak diffracted intensities.
Strain error comparison between correlation and FCU-Net strain measurement, a as a function of sample thickness, and b as a function of electron dose per pattern for 20 nm thick samples. Solid line correspond to the median error, and the shaded regions show the interquartile range, for FCU-Net (red) and correlation (blue) methods, respectively.
In Fig. 3b, we compare the performance of both methods for 20 nm thick samples, as a function of electron dose. We were unable to use the correlation method to measure accurate lattice parameters on patterns with less than 1000 electrons. However, the FCU-Net was able to estimate the lattice with reasonable accuracy on patterns with as few as 100 electrons, due to it pooling information across all disks in Fourier space. At up to 104 electron dose, the FCU-Net is ~50% more accurate than the correlation method. At a dose of 104, the strain error of the correlation method reaches a plateau, demonstrating that the accuracy is no longer limited by dose, but rather by the error in disk positions introduced by multiple scattering. However, the strain errors from the FCU-Net lattice measurements continues to decrease until ~106 electrons, reaching an accuracy over four times greater than the correlation method. We ascribe the higher FCU-Net accuracy to both the Fourier-space convolutional layers, which allow information from all lattice vectors to be pooled together, and to the large size of our training dataset. Together, these enable the FCU-Net to correctly estimate the position of structure factor peaks even when the Bragg disks are close together or even overlap, when signal-to-noise is low, or in the presence of nonlinear variation of the signal within the disks. We believe this robustness makes FCU-Net a good candidate for measurements of samples with unknown structures and orientations, where it may not be possible to guarantee non-overlapping disks or thick samples.
Strain maps from simulated Si-SiGe multilayer data
We next compare strain maps generated using both the cross-correlation and FCU-Net approaches for realistic simulated datasets. The sample geometry consists of alternating layers of Si and SiGe on a mixed SiGe substrate. Two datasets are shown in Fig. 4, both containing the same strain profile, which alternates between ±1% strain relative to the substrate. The first, shown in Fig. 4a–e, is perfectly aligned along the [011] zone axis. The second, shown in Fig. 4f–j, has been helically twisted such that all regions of the sample are tilted away from the ideal diffraction condition. The tilt magnitude varies linearly from 0.4∘ to 4.4∘ from the substrate to the left side, and the tilt direction varies linearly from 45∘ to 315∘ relative to the x-axis.
Measurements perform on a crystal a–e without mistilt, and f–j with helical mistilt. a, f Virtual bright field images calculated from the center disk, with the diffraction patterns corresponding to marked probe positions given in b, g. Real space and reciprocal space scale bars are equal to 5 nm and 0.5 Å−1, respectively. Strain maps measured with c, h cross-correlation and d, i FCU-Net. e, j Line profiles of the mean strain perpendicular (left) and parallel (right) to the interfaces.
Figure 4a shows a virtual bright field image constructed from the center disk across all the diffraction patterns in the perfectly aligned sample. Diffraction patterns from the five regions marked in Fig. 4a are shown in Fig. 4b. The strain maps for this sample along the two principal directions, ϵxx and ϵyy, are plotted in Fig. 4c and d using the correlation method and the FCU-Net model, respectively. For both predictions, the reference lattice is set to be the mean lattice measured from the substrate region on the right hand side.
Figure 4e plots line profiles along the x-direction, perpendicular to the interfaces, of the mean strain for each of ϵxx and ϵyy (left and right, respectively). The strain parallel to the layer interfaces should be ϵyy = 0 everywhere (for an epitaxial film). The ϵyy strain estimated from correlation shows significant deviation from the expected zero strain value, varying systematically and periodically from zero strain near the interfaces, producing a RMS error of ~ 0.2% across the multilayer stacks. In contrast, the FCU-Net ϵyy strain shows almost negligible systematic and random errors (RMS error ≤0.02%).
The strain in the normal direction ϵxx should optimally follow the ideal profile plotted in Fig. 4e. Both approaches perform reasonably well, with the correlation method performing better in the positively strain layers (tension) while the FCU-Net underestimates the strain magnitudes at the middle of each layer, and rounds off the sharp interfaces between layers. Importantly, this effect was not present in the simulated distorted sample or the experimental datasets, and will be discussed in subsequent sections. The likely source of the interfacial error is that at the boundaries, where there is a gradient in both the lattice parameter and the local composition, neither of which have been included in the FCU-Net training. The underestimate of the strain values inside the layers might be due to the highly dynamical intensity measurements present when the sample is perfectly aligned on the zone axis. Additional “on-axis” training data may be required to improve the accuracy of the predicted lattice parameters.
Next, we calculate strain maps from the simulated multilayer dataset, which has been twisted off the ideal diffraction condition. Figure 4f shows the virtual bright field image, and Fig. 4g plots the diffraction patterns for selected positions marked in Fig. 4f. The varying stripes of intensity in the bright field image, and the shifting disk intensity envelope function in the five shown diffraction patterns, both result from the helical twisting of the sample. We again calculate strain maps along the principal directions, shown in Fig. 4h (correlation) and Fig. 4i (FCU-Net). Once again, the reference lattice for the calculation was taken to be the mean lattice vectors from the substrate region on the right of the scan.
Figure 4j plots the line profile of mean strain values parallel and perpendicular to the multilayer stacks. The expected strains are again ϵyy = 0, and ϵxx = ± 1% alternating between the Si and SiGe layers. In ϵyy, the estimates from the correlation method deviate significantly from 0 strain, with a RMS error of ~0.6% in the multilayer region. By contrast, the FCU-Net predictions are closer to the expected zero strain value, with a negligibly small RMS error (<0.1%).
In ϵxx, the correlation method is accurate for several of the layers close to the middle of the scan region, where the mistilt is smallest; however, it becomes quite inaccurate on the left half of the image, where it captures the location of the interfaces but systematically and significantly underestimates the true strain values and fabricates variation within individual layers, where the profile should be flat. Similarly, correlation becomes inaccurate on the far right of the image, in the reference substrate, making it challenging to even estimate the reference lattice. We attribute these artifacts to the varying tilt of the sample, which is known to deleteriously affect template matching by shifting the center of mass of disk intensities. In contrast, the FCU-Net ϵxx strain map mirrors the ground truth value with good fidelity, showing only small deviations such as some slight rounding of the interfaces. The effectiveness of FCU-Net in the presence of sample mistilts is important, as this is a common occurrence in experimental data and very often produces significant error when using traditional strain measurement methods.
Strain maps from experimental h-BN films
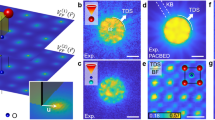
To test the performance of FCU-Net on experimental data, we compute strain maps for hexagonal-boron nitride (h-BN) 4D-STEM datasets using cross-correlation and FCU-Net. Data was collected using four different electron probes, three with circular apertures and convergence semiangles of 0.86, 3.4 and 12 mrads, and one with a bullseye-patterned aperture and 3.4 mrad semiangle27. Figure 5a shows mean diffraction patterns from 20 × 20 different scan positions for each of these probes. Figure 5b, c show strain maps from the correlation and FCU-Net methods, respectively, with the reference lattice set to the average of all positions in the bullseye pattern measurements. The full strain tensor is shown for all positions, consisting of the two principal strain direction ϵxx and ϵyy, the shear strain ϵxy, and the rotation θ. We expect the single crystal h-BN sample to be essentially free of strain and local rotations, suggesting an ideal measurement of 0 for all channels. The mean and standard deviation of the strain values for all probe positions are inset into each panel in Fig. 5b, c. The mean and standard deviations represent the systematic and random errors, respectively. As the field of view is so large, there is some thickness and tilt variation over the field of view.
a Mean diffraction images of 20x20 probe positions, for STEM probes defined by 3.4, 0.86, 3.4, and 12 mrad semiangle apertures, where the leftmost aperture also contains a bullseye pattern. Scale bar is equal to 1 Å−1. b Strain maps measured using cross-correlation template matching for the 4 cases given above. c Strain maps measured using the FCU-Net network predictions. For all maps, the mean and standard deviation strains/angles are inset, relative to the correlation bullseye 3.4 mrads measurement. Scale bar is equal to 200 nm.
The first column of Fig. 5b, c shows results from the 3.4 mrad bullseye probes. Cross-correlation and FCU-Net both perform very well on this data, producing means and standard deviations very close to zero. Some position dependent systematic errors are visible for both methods, possibly due to the sharp edges of the patterned aperture combined with the few pixel shifts of the patterns over the field of view. Interestingly, it is worth noting that FCU-Net does quite well with the bullseye data, despite being trained only on conventional (circular) probes. The surprisingly impressive performance in the strain measurements with completely unseen diffraction images from patterned aperture can be attributed to the introduction of the Fourier-space cross-correlation preprocessing layer as implemented in the FCU-Net model (Fig. 8). While FCU-Net is robust to the patterned probe data, it is possible training on a dataset containing patterned probes may improve accuracy, and is worthy of future research.
Similarly, for the 0.86 mrad probes, shown in the second column of Fig. 5b, c, both correlation and the FCU-Net perform well overall, with means close to 0 in all cases. The standard deviations, indicating the random error, are larger than for the bullseye data, with values as high as ~1% for the correlation ϵxx and ϵxy maps and 0.25% for several of the FCU-Net maps.
These first two columns represent experimental conditions that are well suited to Bragg disk detection using cross-correlation. Bullseye apertures were specifically designed to perform disk detection well using template matching, and this result is borne out here; however, these apertures sacrifice spatial resolution and introduce high-frequency components to the probe shape in real space. Similarly, using a small convergence semiangle improves the disk detection accuracy with cross-correlation by minimizing the chance of disk overlap and the effects of intensity variation within the disks, at the cost of limiting the spatial resolution since reducing the probe size in diffraction space increases its size in real space. The capacity to accurately detect disk position while opening up the aperture size is therefore highly desirable if high-spatial resolution is required.
In the third column of Fig. 5b, c (3.4 mrad probe), the disks begin to show significant intensity gradients within the disks, with higher intensities closer to the origin. This leads to significant positive systematic error in the principal strains (ϵxx and ϵyy) for the correlation estimates. This is likely because the correlation-estimated disk positions are slightly biased towards the origin, leading to a smaller estimated reciprocal lattice and thus positive real space strains. This effect should not modify the results for either shear strain or rotation, and indeed both of these quantities show low error. By contrast, the FCU-Net predictions show low systematic errors for all 4 components of the strain tensor, demonstrating the robustness of the FCU-Net approach to variations in disk intensities. Both methods show fairly low random errors of 0.10% and 0.13% for correlation and FCU-Net, respectively.
In the final column of Fig. 5b, c (12 mrad probe), the disks have expanded to create significant overlap, a condition required for atomic-resolution imaging, but which typically thwarts traditional template matching. The resulting systematic errors are very high, approximately −1.1%, and significant variation over the field of view is visible in all correlation measurements. FCU-Net, in spite of being trained on images with probe semiangles up to a maximum of 4 mrads, performs fairly well on this data, with systematic errors ~5 times lower than the correlation method. We ascribe this to the training dataset containing many crystals and orientations that produce disk overlaps for 4 mrad probes (and below), such that the network has learned to interpret the complex interference patterns formed in the presence of overlapping disks. The random errors are also lower for the FCU-Net compared to the correlation method, and the predicted strains show less variation across the field of view. Overall, the FCU-Net produces more accurate and precise strain predictions over a wider parameter range than the correlation method, including experimental conditions it was not exposed to during training. We also note that the strain measurement accuracy using FCU-Net model may be further improved by fine tuning the pre-trained model with application-specific diffraction data.
Strain maps from experimental SiGe multilayer stacks
Finally, we compare the two strain calculation methods on a thick, non-uniform multilayer stack of alternating layers of Si and a mixture of Si and Ge grown epitaxially. A virtual image constructed from the center disk is shown in Fig. 6a. We observe significant contrast differences over the field of view, corresponding to variation in the sample’s thickness, composition and surface morphology. We have estimated the local composition of the sample by using STEM-EELS, shown in Fig. 6b. The mean composition of the 5 stripes from STEM-EELS is Si0.82Ge0.18. We estimate that the average thickness of the sample is ≈ 110 nm, using the t/λ method65 applied to the pure Si regions and are therefore in the multiple scattering regime66. The local relative thickness is plotted in Fig. 6c, showing a relative thickness variation of about 20%.
a Virtual bright field calculated from center disk. b Composition and c relative thickness, estimated from STEM-EELS. d Diffraction patterns corresponding to the probe positions marked in (a), with estimated Bragg disk positions from e correlation template matching and f FCU-Net. g Strain maps measured from correlation template matching. h Strain maps measured from FCU-Net. i Mean strain values parallel to the multilayer normal direction, for correlation, FCU-Net, and estimated from the STEM-EELS composition. Scale bars are equal to 10 nm.
We plot examples of the diffraction patterns in Fig. 6d, from 5 regions marked in Fig. 6a. We see significant variation in the fine structure of the diffracted disks, especially when comparing regions of different compositions. The round shape of many of the disks are significantly degraded due to the thickness and non-uniformity of the sample. Finally, the center-of-mass of the diffraction pattern intensities changes over the field of view, indicating that bending of the sample had lead to slightly different tilt conditions for different probe positions. We have used both cross-correlation and FCU-Net to estimate the Bragg disk positions, with examples shown in Fig. 6e, f, corresponding to the diffraction patterns shown in Fig. 6d. The resulting disk positions are noticeably less regular for the correlation method, and many disks at higher diffraction angles close to the image edges are too weak to be identified. This is in contrast to the FCU-Net predictions, which returns a highly regular lattice of disk positions, with only a few weak false positives visible at the image boundaries.
The strain maps along the principal directions calculated with the correlation method are shown in Fig. 6g, and those calculated using the FCU-Net predictions are shown in Fig. 6h. In both cases, the reference lattice was taken to be the mean lattice vectors from the substrate region on the right of the field of view. Figure 6i plots line profiles of the mean strain values perpendicular (left) and parallel (right) to the multilayers. In the parallel direction, we expect the strain will be ϵyy = 0 everywhere, due to the epitaxial nature of the layers. The correlation strain shows significant deviation from 0 strain, and moreover, is not flat over the imaged area, with deviations ranging from approximately –0.4% on the left side, to +0.6% in the center, and back down to 0% in the substrate region on the right hand side. The FCU-Net strain ϵyy by contrast is comparatively flat, and ranges from approximately +0.2% on the left side, to 0% strain in the substrate on the right hand side. We note that while the RMS error in strain ϵyy calculation across all the multilayer stacks is ~0.3% with cross-correlation approach, it is ~0.15% from the FCU-Net prediction.
In the normal direction, we can compare the strain ϵxx computed with cross-correlation and with FCU-Net to the strain measured using independent STEM-EELS measurements. The STEM-EELS result is shown as a black line in Fig. 6i. The FCU-Net line profile closely approximates the STEM-EELS profile, capturing most of the sharp transitions at the interfaces, and the roughly flat profiles within each layer. The cross-correlation result fares much worse, capturing the ϵxx structure of the three right-most layers roughly correctly, but then deviating wildly on the left side of the scan region, possibly due to local sample mistilt. The correlation result also deviates from a flat profile in the substrate on the right, making identification of a reference lattice difficult. For the strain ϵxx, FCU-Net produces a RMS error of ~0.25% across the sample leading to almost three-fold increase in the accuracy from cross-correlation, which produced a RMS error of ~0.72%. This example highlights common pitfalls of traditional template matching in the presence of complex, nonlinear electron scattering signals, and the capacity of the FCU-Net model to achieve accurate disk localization measurement in spite of these challenges.
In summary, we have developed a deep-learning network (FCU-Net) for quantitative measurements of Bragg disk positions from electron diffraction patterns. Our networks have been trained with over 200,000 unique, simulated diffraction patterns with thicknesses ranging from 2 to 50 nm thick, covering more than 1000 distinct crystal systems over many orientations and microscope parameters. We found that the resulting Bragg disk position predictions from the FCU-Net network were substantially more accurate than a conventional template matching correlation method. We tested the FCU-Net predictions for crystalline lattice strain mapping, using both simulated and experimental 4D-STEM datasets. In both cases, we found that the FCU-Net predictions were substantially more robust against signal variations due to mistilt of the sample and multiple scattering due to sample thickness. We have integrated FCU-Net into the open-source 4D-STEM analysis python library py4DSTEM, providing free access and use of the network, and a complementary suite of tools for subsequent analysis of the measured structure factors, to the electron microscopy community. All of our simulated and experimental datasets, source codes, and trained networks are freely available in open-source repositories. The improved accuracy and precision of Bragg disk measurements using FCU-Net, even in the presence of complex signals involving thick samples and multiply scattered electrons, can provide widespread benefits in 4D-STEM application such as strain, phase, and orientation mapping, and in quantitative electron crystallography.
Methods
Figure 1 shows a flow chart of the methods we use to invert STEM diffraction patterns into quantitative structure factor positions and amplitudes. First we generate a library of simulated dynamical diffraction data (Fig. 1a). We selected thousands of unique material systems that span a wide variety of crystallographic prototype systems, and simulated the CBED patterns at various thicknesses, tilts, and microscope conditions using the multislice algorithm67,68. The projected structure factors are then computed, including the effect of any excitation error by evaluating the distance of the projected potentials from the Ewald sphere. Simulated data that will be used for training is then augmented with noise profiles, which mimic real experimental conditions. The network is then trained using the noise-augmented simulated data. Figure 1c overviews the input, architecture, and output of the FCU-Net deep-neural network used to predict the (projected) structure factor positions from the input diffraction patterns and electron probe. Figure 1d–f show the typical inference stage, where we use the pre-trained FCU-Net model to predict the underlying structure factor positions and amplitudes from experimental diffraction patterns.
Dynamical diffraction library simulations
To build a dynamical diffraction library for the AI/ML training, we implemented an automated pipeline, which selects the crystal structures, and simulates CBED patterns and the underlying projected structure factors with a variety of experimental parameters. The dynamical diffraction library generation starts with building a materials database. To judiciously select crystal structures of interest for our problem, we initially compare ≈139,000 crystal structures and compositions from the materials project (MP) database69 with more than 500 crystallographic prototypes collected from the AFlow library (Fig. 7)70,71. Crystallographic prototypes are an alternative and popular crystal structure classification paradigm. Figure 7a shows the distribution of the crystal systems from the MP database, grouped according to their structural similarity with crystallographic prototype systems. We presented the first 250 prototype systems, as shown in Fig. 7a, which cumulatively span ~95% of the materials systems from materials project database. We sampled ~1000 unique crystal systems following the distribution, presented as a blue line in Fig. 7a.
Figure 7 b–e plots the distribution of atomic number space of the crystal structures, which are structurally similar to four different example prototype systems—CaTiO3, FeB, Fe3C, and Zn3P2. As evident from the distribution in panel b–e, the selected materials systems have diverse range of constituent atomic elements. Following the crystal system extraction, we simulated the CBED patterns and underlying structure factors using the multislice algorithm67,68, as implemented in the Prismatic code72,73.
From these simulations, the corresponding ground truth structure factors are calculated from the projected atomic potentials for each diffraction pattern. This is achieved by first transforming atomic potentials into 3D Fourier space, applying a 2D Tukey window function in the projection plane, and 2D Fourier downsampling to attain the desired output resolution in x and y. A Gaussian weighted filter is applied along z-axis (the beam direction) with a standard deviation of 0.05 Å−1 to select the structure factors close to the projection slice. Finally, the projection is summed along z-axis to generate the ground truth structure factors. Note that these structure factor images are depend linearly on the thickness of the sample. We simulated CBED patterns and the underlying structure factors for all the 1000 unique crystal systems for thicknesses between 2 to 50 nm with an interval of 2 nm. For each crystal system we simulated diffraction patterns for the crystal orientated along 5 different low-index zone axes, and 5 random orientations. We simulated diffraction patterns for each orientation with probe semiangles of 1, 2, and 4 mrads. In total this yielded diffraction library of 750,000 diffraction patterns, each with a unique combination of crystal system, sample tilt, specimen thickness and probe convergence angle. For each of the 750,000 diffraction patterns the probe and structure factors were also created. We have implemented a parallelized framework for the data simulation, training data generation, and training steps74.
Conventional Bragg disk position measurements
Determining the Bragg disk positions and intensities in each diffraction pattern is an important step, which allows subsequent measurement of parameters such as phase, orientation, and strain in crystalline and semi-crystalline materials. Cross-correlative template matching is one method routinely used to measure the positions of Bragg disks10,28, matching to either raw diffraction patterns or edge-filtered images29. In the template matching approach, the Bragg disk positions are calculated in two steps—first, we collect the undiffracted probe over vacuum to create our template for matching. Next we perform cross-correlation between the diffraction pattern and the probe template in Fourier space to find all disk positions in a given diffraction pattern. In this work, we use the disk detection, lattice fitting, and strain mapping tools implemented in the open-source python package py4DSTEM23.
Bragg disk detection using Fourier-space deep learning
We implement three variants of CNN architecture-U-Net56, and its modified variants with spectral parameterization adapted from Ripple et al.62 and fully complex variant, FCU-Net adapted from Trabelsi et al.63. Figure 8a presents the model architecture of U-Net and its hybrid variants with fully complex convolution and spectral pooling layers. The FCU-Net architecture implemented in this work considers two inputs: the probe template and the CBED diffraction pattern. To make the FCU-Net model aware of the vacuum probe template, we implement a preprocessing layer, which multiplies the Fourier transform of the diffraction pattern with the probe template. Finally, we implement the 2D complex convolutional layer, which is the building blocks for the FCU-Net, to teach the complex space information from the Fourier transformed image from the preprocessing layer. Following a combination of complex convolutions, pooling and upsampling operations the final output from the FCU-Net is transformed using inverse Fourier transform operation, before it is compared with the ground truth atomic potentials.
Complex convolution
We implement complex convolutional layers by independently initializing real and imaginary components of the 2D convolutional kernel (Fig. 8b), that is, we consider the real and imaginary parts of the complex numbers as logically distinct real-valued numbers. Akin to the 2D real-valued convolution operator, we convolve a complex kernel matrix (K = KR + iKI); KR, \({K}_{I}\in {{\mathbb{R}}}^{m/2\times m/2}\) with the complex input feature map (F = FR + iFI); FR, \({F}_{I}\in {{\mathbb{R}}}^{m/2\times N}\), where m/2 is the size of the complex kernel weight and N is the number of pixels in the input image (feature map). The complex convolution operation can be formulated as:
We can use a matrix notation to represent the complex convolution operator:
Out of the variety of options available for activation functions for complex convolutions, we have chosen to use the complex rectified linear unit (\({\mathbb{C}}\)ReLU) function such that for any complex number z:
Trabelsi et al. recently compared different variants of ReLU functions for complex operators, and found that \({\mathbb{C}}\)ReLU(z) had the best performance63. In our tests, we found \({\mathbb{C}}ReLU(z)\) to be the preferred nonlinear activation function, as it can distinguish correlations from the complex convolution operation into four distinct region based on if the \({\mathfrak{Re}}(z)\) and \({\mathfrak{Im}}(z)\) are strictly positive or negative. For deep networks such as FCU-Net, this provides the required flexibility and nonlinearity to the network by allowing complete manipulation of the phase information at each layer of the network.
Spectral pooling
To implement the U-Net with spectral parameterization we replace the max-pooling layers typically used in U-Net models with spectral pooling layers as we find that this reduces the introduction of artifacts and nonlinearity, resulting in a more stable and accurate prediction from the network. Where max-pooling layers down sample the image in real space, spectral pooling operates in the frequency domain. Spectral pooling in its original form as described by Rippel et. al.62, transforms an image to Fourier space by applying a fast Fourier transform operation (FFT), after which it is cropped in Fourier space and transformed back to real space by an inverse FFT such as: \(x\in {{\mathbb{C/R}}}^{M\times M}\mathop{\to }\limits^{{{{\rm{FFT}}}}}\tilde{x}\in {{\mathbb{C}}}^{M\times M}\mathop{\to }\limits^{{{{\rm{Crop}}}}}\tilde{x}\in {{\mathbb{C}}}^{N\times N}\mathop{\to }\limits^{{{{\rm{inv}}}}\,{{{\rm{FFT}}}}}x\in {{\mathbb{C}}}^{N\times N}\), where x and \(\tilde{x}\) are the input and Fourier transformed image, respectively, N and M correspond the number of pixels in the image, with N < M.
Training FCU-Net
We train the fully complex FCU-Net network on the simulated sets of images composed of a vacuum probe, a CBED pattern, and the ground truth structure factors, for different material systems at different sample thicknesses up to 50 nm. To make FCU-Net robust against various experimental conditions, we augment the simulated images with several forms of noise typically found in 4D-STEM data: (i) elliptical distortion and (ii) random translations (x,y pixel shifts) of the diffraction patterns, (iii) incoherent backgrounds modeled as plasmonic signal, (iv) shot (counting) noise using Poisson statistics, and (v) random bright (hot) and dark (dead) pixels to simulate the effect of X-rays and detector pixel errors.
For the final training, we randomly sampled ~200,000 unique training (~20,000 test) triplets from the diffraction pattern library. Each triplet contained a vacuum probe and a CBED pattern, used as the training inputs and the structure factors for the training output. Table 2 summarizes the hyperparameters considered during the FCU-Net training. Before the final training iteration, we implement a high-throughput hyperparameter optimization scheme using RayTune python library for deep learning75. A random subset of the training data was used during hyperparameter tuning, as a compromise between accuracy and the computational overhead. Following the hyperparameter optimization, we perform the final round of training iterations for the FCU-Net on 8 NVIDIA Tesla V-100 (16 GB VRAM) GPU nodes using a distributed Tensorflow strategy to accelerate the training performance76. All training and test runs for this work were performed on the super-computing facility (Cori GPU clusters) at the National Energy Research Scientific Computing Center (NERSC).
Integration with py4DSTEM
Bragg disk detection using the trained FCU-Net model is implemented in the py4DSTEM python data analysis toolkit developed by Savitzky et al.23. The workflow for AI/ML guided disk detection using py4DSTEM starts with loading a 4D dataset and the corresponding vacuum probe. These inputs are passed to a function, which feeds them into the trained FCU-Net model, which returns the predicted disk positions. Currently we host the latest (and previously archived versions) of pre-trained model weights on a cloud location and which is updated periodically with new weights with improved test performance. When called, the py4DSTEM AI/ML disk detection function will search for the latest FCU-Net weights and automatically download them prior to disk detection. Once the prediction is completed, we convert the predicted output (a 2D image-like array of structure factors) to a set of M peaks defined by the values \(({q}_{m}^{x},{q}_{m}^{y},{I}_{m})\), which can be used with any of the existing downstream analysis modalities built into py4DSTEM.
Strain mapping
Strain mapping was performed using py4DSTEM. Using the measured disk positions, either from FCU-Net predictions or cross-correlation, we fit the lattice vectors at each beam position. A reference lattice is chosen, and the difference between the reference and local lattice vectors are then used to calculate the infinitesimal strain tensor
where ϵxx and ϵyy are the strain along the x and y directions, and ϵxy is the shear strain. We additionally calculate θ, the rotation of the local lattice relative to the reference lattice. The selection of reference lattice is specified for each strain map computed. More details can be found in23,28.
Simulated diffraction of SiGe multilayers
In order to test the robustness of our network for realistic samples, we perform simulations of thick samples, which incorporate multiple scattering of the electron beam. The sample geometry we used is a multilayer stack along the [011] direction, composed of alternating Si and Si0.5Ge0.5 layers, on a Si0.75Ge0.25 substrate, where each phase has diamond cubic structure. For ease of comparison of our measured strain values with the ground truth, we used slightly different lattice constants from known experimental values, setting the substrate to have a lattice parameter of 5.6034 Å, and the multilayers to have precisely ±1% strains relative to the substrate.
Experimental diffraction of SiGe multilayers and h-BN films
Experimental 4D-STEM datasets were acquired using the TEAM I instrument at the National Center for Electron Microscopy facility of the Molecular Foundry, a double aberration corrected Thermo Fisher Titan fitted with a Gatan Continuum energy filter and K3 direct electron detector. The K3 detector was operated in electron counting mode. Electron diffraction patterns were acquired in energy-filtered mode with a 15 eV slit centered on the elastic energy to suppress background noise from inelastic scattering.
Hexagonal-boron nitride
In order to obtain a reference dataset from a thin, single crystal material with minimal characteristic strain we used thin a flake mechanically exfoliated from a single crystal of hexagonal-boron nitride. This flake was transferred to a silicon nitride TEM grid for 4D-STEM experiments. Multiple 4D-STEM datasets were acquired at an 80 kV accelerating voltage using four different apertures to compare algorithmic performance under various experimental conditions. Three circular apertures were used, with convergence semiangles of 0.86, 3.4, and 12 mrad, and one bullseye-patterned aperture was used27, with a 3.4 mrad convergence semiangle. For each aperture, data was acquired with a 50 ms dwell time, step size of 100 Å, and scan size of 112 × 108 probe positions. Diffraction patterns were binned 4 x 4 after electron counting.
Si-Si/Ge multilayers
In order to obtain an experimental dataset with a large and known strain, we used a silicon/silicon-germanium “MAG*I*CAL” calibration sample obtained from Ted Pella, Inc. The sample consists of a Si wafer with several layers of ~10 nm of Si/Ge mixture grown epitaxially. The sample is prepared for TEM as a polished cross-section with the [110] zone axis normal to the foil. Data was acquired at a 300 kV accelerating voltage and 1.3 mrad convergence semiangle, with a step size of 10 Å and a scan size of 200 x 50 probe positions.
To obtain an independent measurement of the sample strain, we also acquired an electron energy loss spectrum (EELS) dataset from the same region of the sample. Analysis of the EELS data showed the average thickness to be approximately one inelastic mean free path, corresponding to an estimated thickness of 110 nm. Chemical analysis showed the Si region to be pure Si, and the SiGe alloy region to have an average composition of 18% Ge. From this chemical analysis we can derive the expected strain in the SiGe layers.
First, we use Vegard’s law, which posits that the strain depends linearly on the composition xSi77. The Si0.82Ge0.18 layers have a larger lattice constant, and thus will expand relative to the Si layers in the x direction. As the multilayers are epitaxial, the Si0.82Ge0.18 layers are compressed in the multilayer interfacial plane in two directions, which will lead to an additional expansion given by the Poisson’s ratio multiplied by two. The overall strain profile can therefore be estimated as
which is plotted in Fig. 6i, using literature values for the cubic lattice constants of Si and Ge of aSi = 5.54 and aGe = 5.66 Å, respectively78, and for the Poisson’s ratio ν of Si and Ge of ~0.275 in the (001) direction79.
Data availability
Dynamical diffraction library generation tool and the simulated training dataset are available upon reasonable request.
Code availability
Codes related to FCU-Net model, data preprocessing and augmentation can be found in crystal4D repository and are available as open-source package. Distributed Hyperparameter tuning pipeline using rayTune can be found at https://github.com/AI-ML-4DSTEM/4D-OPTIMIZE/tree/nersc_ray. Disk detection using AI/ML (FCU-Net) is implemented as a new functionality in py4DSTEM 0.12.x. The simulated and experimental strain measurements performed in this paper and the required 4D-STEM dataset are available as tutorial notebooks and can be accessed at https://github.com/py4dstem/py4DSTEM_tutorials/tree/main/notebooks/version_0.12/strain_aiml, and will be updated for future releases of py4DSTEM.
References
Liu, J. J. Advances and applications of atomic-resolution scanning transmission electron microscopy. Microsc. Microanal. 1–53 https://doi.org/10.1017/s1431927621012125 (2021).
Haider, M. et al. Electron microscopy image enhanced. Nature 392, 768–769 (1998).
Krivanek, O., Nellist, P., Dellby, N., Murfitt, M. & Szilagyi, Z. Towards sub-0.5 å electron beams. Ultramicroscopy 96, 229–237 (2003).
Yankovich, A. B. et al. Picometre-precision analysis of scanning transmission electron microscopy images of platinum nanocatalysts. Nat. Commun. 5, 1–7 (2014).
Hage, F., Radtke, G., Kepaptsoglou, D., Lazzeri, M. & Ramasse, Q. Single-atom vibrational spectroscopy in the scanning transmission electron microscope. Science 367, 1124–1127 (2020).
Zachman, M. J., Tu, Z., Choudhury, S., Archer, L. A. & Kourkoutis, L. F. Cryo-stem mapping of solid–liquid interfaces and dendrites in lithium-metal batteries. Nature 560, 345–349 (2018).
Yang, Y. et al. Deciphering chemical order/disorder and material properties at the single-atom level. Nature 542, 75–79 (2017).
Yankovich, A. B., Berkels, B., Dahmen, W., Binev, P. & Voyles, P. M. High-precision scanning transmission electron microscopy at coarse pixel sampling for reduced electron dose. Adv. Struct. Chem. Imaging 1, 1–5 (2015).
Ophus, C. Four-dimensional scanning transmission electron microscopy (4D-STEM): From scanning nanodiffraction to ptychography and beyond. Microsc. Microanal. 25, 563–582 (2019).
Ozdol, V. et al. Strain mapping at nanometer resolution using advanced nano-beam electron diffraction. Appl. Phys. Lett. 106, 253107 (2015).
Nord, M. et al. Three-dimensional subnanoscale imaging of unit cell doubling due to octahedral tilting and cation modulation in strained perovskite thin films. Phys. Rev. Mater. 3, 063605 (2019).
Das, S. et al. Observation of room-temperature polar skyrmions. Nature 568, 368–372 (2019).
Ahmed, S. et al. Visualization of light elements using 4D STEM: the layered-to-rock salt phase transition in linio2 cathode material. Adv. Energy Mater. 10, 2001026 (2020).
Chen, Z. et al. Electron ptychography achieves atomic-resolution limits set by lattice vibrations. Science 372, 826–831 (2021).
Hughes, L. et al. Correlative analysis of structure and chemistry of LixFePO4 platelets using 4D-STEM and x-ray ptychography. Mater. Today https://doi.org/10.48550/arXiv.2107.04218 (2021).
Deng, H. D. et al. Correlative image learning of chemo-mechanics in phase-transforming solids. Nat. Mater. 21, 547–554 (2022).
Oxley, M. P. et al. Deep learning of interface structures from simulated 4d stem data: cation intermixing vs. roughening. Mach. Learn. Sci. Technol. 1, 04LT01 (2020).
Kazmierczak, N. P. et al. Strain fields in twisted bilayer graphene. Nat. Mater. 20, 956–963 (2021).
Zachman, M. J. et al. Interferometric 4D-STEM for lattice distortion and interlayer spacing measurements of bilayer and trilayer 2d materials. Small 2100388 https://doi.org/10.1002/smll.202100388 (2021).
Johnstone, D. N. et al. pyxem/pyxem: pyxem. Zenodo (2021).
Clausen, A. et al. Libertem: Software platform for scalable multidimensional data processing in transmission electron microscopy. J. Open Source Softw. 5, 2006 (2020).
Ziatdinov, M., Ghosh, A., Wong, T. & Kalinin, S. V. Atomai: A deep learning framework for analysis of image and spectroscopy data in (scanning) transmission electron microscopy and beyond. Preprint at https://arxiv.org/abs/2105.07485 (2021).
Savitzky, B. H. et al. py4DSTEM: a software package for four-dimensional scanning transmission electron microscopy data analysis. Microsc. Microanal. 27, 712 (2021).
Mahr, C. et al. Theoretical study of precision and accuracy of strain analysis by nano-beam electron diffraction. Ultramicroscopy 158, 38–48 (2015).
Williamson, M., van Dooren, P. & Flanagan, J. Quantitative analysis of the accuracy and sensitivity of strain measurements from nanobeam electron diffraction. In: 2015 IEEE 22nd International Symposium on the Physical and Failure Analysis of Integrated Circuits. 197–200 (IEEE, 2015).
Grieb, T. et al. Strain analysis from nano-beam electron diffraction: Influence of specimen tilt and beam convergence. Ultramicroscopy 190, 45–57 (2018).
Zeltmann, S. E. et al. Patterned probes for high precision 4D-STEM bragg measurements. Ultramicroscopy 209, 112890 (2020).
Pekin, T. C., Gammer, C., Ciston, J., Minor, A. M. & Ophus, C. Optimizing disk registration algorithms for nanobeam electron diffraction strain mapping. Ultramicroscopy 176, 170–176 (2017).
Mukherjee, D., Gamler, J. T., Skrabalak, S. E. & Unocic, R. R. Lattice strain measurement of core shell electrocatalysts with 4d scanning transmission electron microscopy nanobeam electron diffraction. ACS Catal. 10, 5529–5541 (2020).
Yuan, R., Zhang, J. & Zuo, J.-M. Lattice strain mapping using circular hough transform for electron diffraction disk detection. Ultramicroscopy 207, 112837 (2019).
Müller, K. et al. Scanning transmission electron microscopy strain measurement from millisecond frames of a direct electron charge coupled device. Appl. Phys. Lett. 101, 212110 (2012).
Padgett, E. et al. The exit-wave power-cepstrum transform for scanning nanobeam electron diffraction: robust strain mapping at subnanometer resolution and subpicometer precision. Ultramicroscopy 214, 112994 (2020).
MacLaren, I. et al. Comparing different software packages for the mapping of strain from scanning precession diffraction data. Microsc. Microanal. 27, 2–5 (2021).
Seyring, M., Song, X. & Rettenmayr, M. Advance in orientation microscopy: quantitative analysis of nanocrystalline structures. ACS Nano 5, 2580–2586 (2011).
Shukla, A. K., Ophus, C., Gammer, C. & Ramasse, Q. Study of structure of li-and mn-rich transition metal oxides using 4D-STEM. Microsc. Microanal. 22, 494–495 (2016).
Ophus, C. et al. Automated crystal orientation mapping in py4dstem using sparse correlation matching. Microsc. Microanal. 28, 390–403 (2022).
Bedell, S., Khakifirooz, A. & Sadana, D. Strain scaling for CMOS. MRS Bulletin 39, 131–137 (2014).
Chidambaram, P., Bowen, C., Chakravarthi, S., Machala, C. & Wise, R. Fundamentals of silicon material properties for successful exploitation of strain engineering in modern cmos manufacturing. EEE Trans. Electron Devices 53, 944–964 (2006).
Wang, Z.-J. et al. Sample size effects on the large strain bursts in submicron aluminum pillars. Appl. Phys. Lett. 100, 071906 (2012).
Zhang, J., Liu, G. & Sun, J. Strain rate effects on the mechanical response in multi-and single-crystalline cu micropillars: grain boundary effects. Int. J. Plast. 50, 1–17 (2013).
Chen, W. et al. Bending stress relaxation of microscale single-crystal copper at room temperature: an in situ sem study. Eur. J. Mech. A Solids 90, 104377 (2021).
Ede, J. M. Deep learning in electron microscopy. Mach. Learn. Sci. Technol. 2, 011004 (2021).
George, B. et al. Cassper: a semantic segmentation based particle picking algorithm for single particle cryo-electron microscopy. Commun. Biol. 4, 200 (2020).
Roberts, G. et al. Deep learning for semantic segmentation of defects in advanced stem images of steels. Sci. Rep. 9, 1–12 (2019).
Ziatdinov, M. et al. Building and exploring libraries of atomic defects in graphene: Scanning transmission electron and scanning tunneling microscopy study. Sci. Adv. 5, eaaw8989 (2019).
Kaufmann, K. et al. Crystal symmetry determination in electron diffraction using machine learning. Science 367, 564–568 (2020).
Aguiar, J., Gong, M. L., Unocic, R., Tasdizen, T. & Miller, B. Decoding crystallography from high-resolution electron imaging and diffraction datasets with deep learning. Sci. Adv. 5, eaaw1949 (2019).
Vasudevan, R. K. et al. Mapping mesoscopic phase evolution during e-beam induced transformations via deep learning of atomically resolved images. npj Comput. Mater. 4, 1–9 (2018).
Tiong, L. C. O., Kim, J., Han, S. S. & Kim, D. Identification of crystal symmetry from noisy diffraction patterns by a shape analysis and deep learning. npj Comput. Mater. 6, 1–11 (2020).
Lee, C.-H. et al. Deep learning enabled strain mapping of single-atom defects in two-dimensional transition metal dichalcogenides with sub-picometer precision. Nano Lett. 20, 3369–3377 (2020).
Zhang, C., Feng, J., DaCosta, L. R. & Voyles, P. M. Atomic resolution convergent beam electron diffraction analysis using convolutional neural networks. Ultramicroscopy 210, 112921 (2020).
Xu, W. & LeBeau, J. M. A deep convolutional neural network to analyze position averaged convergent beam electron diffraction patterns. Ultramicroscopy 188, 59–69 (2018).
Li, X. et al. Manifold learning of four-dimensional scanning transmission electron microscopy. npj Comput. Mater. 5, 1–8 (2019).
Yuan, R., Zhang, J., He, L. & Zuo, J.-M. Training artificial neural networks for precision orientation and strain mapping using 4d electron diffraction datasets. Ultramicroscopy 231, 113256 (2021).
Shi, C. et al. Uncovering material deformations via machine learning combined with four-dimensional scanning transmission electron microscopy. npj Comput. Mater. 8, 1–9 (2022).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In: Navab, N., Hornegger, J., Wells, W. & Frangi, A. (eds). Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015. Lect. Notes in Comp. Sci, 9351. https://doi.org/10.1007/978-3-319-24574-4_28 (Springer, Cham, 2015).
Danihelka, I., Wayne, G., Uria, B., Kalchbrenner, N. & Graves, A. Associative Long Short-Term Memory. Proc. of the 33rd Int. Conf. on Mach. Learn. PMLR 48. 1986–1994 (2016).
Arjovsky, M., Shah, A. & Bengio, Y. Unitary evolution recurrent neural networks. In: International Conference on Machine Learning. 1120–1128 (PMLR, 2016).
Wisdom, S., Powers, T., Hershey, J., Le Roux, J. & Atlas, L. Full-capacity unitary recurrent neural networks. Adv. Neural. Inf. Process. Syst. 29, 4880–4888 (2016).
Sampat, M. P., Wang, Z., Gupta, S., Bovik, A. C. & Markey, M. K. Complex wavelet structural similarity: A new image similarity index. IEEE Trans. Image Process. 18, 2385–2401 (2009).
Shi, G., Shanechi, M. M. & Aarabi, P. On the importance of phase in human speech recognition. IEEE Trans. Audio Speech Lang. Process. 14, 1867–1874 (2006).
Rippel, O., Snoek, J. & Adams, R. P. Spectral representations for convolutional neural networks. Adv. in Neural Info. Proc. Sys. 28 (2015).
Trabelsi, C. et al. Deep complex networks. Preprint at https://arxiv.org/abs/1705.09792 (2017).
Wang, Z., Bovik, A. C., Sheikh, H. R. & Simoncelli, E. P. Image quality assessment: from error visibility to structural similarity. IEEE Trans. Image Process. 13, 600–612 (2004).
Malis, T., Cheng, S. & Egerton, R. Eels log-ratio technique for specimen-thickness measurement in the tem. J. Electron Microsc. Tech. 8, 193–200 (1988).
Vallejo, I. G. et al. Observation of large multiple scattering effects in ultrafast electron diffraction on monocrystalline silicon. Phys. Rev. B 97, 054302 (2018).
Cowley, J. M. & Moodie, A. F. The scattering of electrons by atoms and crystals. I. a new theoretical approach. Acta Crystallogr. 10, 609–619 (1957).
Kirkland, E. J. Advanced computing in electron microscopy. 3rd edn (Springer Science & Business Media, 2020).
Jain, A. et al. The materials project: a materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
Mehl, M. J. et al. The aflow library of crystallographic prototypes: part 1. Comput. Mater. Sci. 136, S1–S828 (2017).
Hicks, D. et al. The aflow library of crystallographic prototypes: part 2. Comput. Mater. Sci. 161, S1–S1011 (2019).
Ophus, C. A fast image simulation algorithm for scanning transmission electron microscopy. Adv. Struct. Chem. Imaging 3, 1–11 (2017).
DaCosta, L. R. et al. Prismatic 2.0–Simulation software for scanning and high resolution transmission electron microscopy (STEM and HRTEM). Micron 151, 103141 (2021).
Rakowski, A. et al. A complete pipeline for deep learning workflows in transmission electron microscopy (manuscript in preparation) (2021).
Liaw, R. et al. Tune: a research platform for distributed model selection and training (2018). Preprint at https://arxiv.org/abs/1807.05118 (2018).
Abadi, M. et al. Tensorflow: a system for large-scale machine learning. In: 12th {USENIX} Symposium on Operating Systems Design and Implementation ({OSDI} 16). 265–283 (2016).
Vegard, L. Die konstitution der mischkristalle und die raumfüllung der atome. Z. Phys. 5, 17–26 (1921).
Johnson, E. R. & Christian, S. M. Some properties of germanium-silicon alloys. Phys. Rev. 95, 560 (1954).
Wortman, J. & Evans, R. Young’s modulus, shear modulus, and poisson’s ratio in silicon and germanium. J. Appl. Phys. 36, 153–156 (1965).
Acknowledgements
This work was primarily funded by the US Department of Energy in the program “4D Camera Distillery: From Massive Electron Microscopy Scattering Data to Useful Information with AI/ML.” M.K.Y.C. and C.O. each acknowledge support of a US Department of Energy Early Career Research Award. J.C. acknowledges support from the Presidential Early Career Award for Scientists and Engineers (PECASE) through the U.S. Department of Energy. B.H.S. and py4DSTEM development are supported by the Toyota Research Institute. S.E.Z. was supported by the National Science Foundation under STROBE Grant no. DMR 1548924. Work at the Molecular Foundry was supported by the Office of Science, Office of Basic Energy Sciences, of the US Department of Energy under Contract No. DE-AC02-05CH11231. Use of the Center for Nanoscale Materials, an Office of Science user facility, was supported by the U.S. Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. DE-AC02-06CH11357. This research used resources of the National Energy Research Scientific Computing Center, a DOE Office of Science User Facility supported by the Office of Science of the U.S. Department of Energy under Contract No. DE-AC02-05CH11231. We acknowledge S. Kim, and J. Carlstroem for assistance with sample preparation. We also acknowledge donation of GPU resources by NVIDIA.
Author information
Authors and Affiliations
Contributions
J.M. and M.K.Y.C. selected crystal systems used in the training. J.M. designed the architecture with input from A.R., M.H., S.C., M.K.Y.C., and C.O. J.M. and A.R. performed the simulations, data augmentation, network training, and validation, with computational help from M.H. and S.C. S.E.Z. and J.C. performed the experimental TEM measurements. J.M., A.R., B.H.S., S.E.Z., and C.O. implemented network predictions and strain measurements into py4DSTEM, and analyzed the experimental results. A.M.M., M.K.Y.C., and C.O. supervised this research. J.M. and A.R. contributed equally to the work and co-wrote the manuscript. All authors contributed to writing and editing this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Munshi, J., Rakowski, A., Savitzky, B.H. et al. Disentangling multiple scattering with deep learning: application to strain mapping from electron diffraction patterns. npj Comput Mater 8, 254 (2022). https://doi.org/10.1038/s41524-022-00939-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-022-00939-9
This article is cited by
-
Integrated analysis of X-ray diffraction patterns and pair distribution functions for machine-learned phase identification
npj Computational Materials (2024)
-
Enhanced accuracy through machine learning-based simultaneous evaluation: a case study of RBS analysis of multinary materials
Scientific Reports (2024)
-
A dynamic Bayesian optimized active recommender system for curiosity-driven partially Human-in-the-loop automated experiments
npj Computational Materials (2024)
-
Leveraging generative adversarial networks to create realistic scanning transmission electron microscopy images
npj Computational Materials (2023)
-
Machine learning for automated experimentation in scanning transmission electron microscopy
npj Computational Materials (2023)