Abstract
Force field-based classical molecular dynamics (CMD) is efficient but its potential energy surface (PES) prediction error can be very large. Density functional theory (DFT)-based ab-initio molecular dynamics (AIMD) is accurate but computational cost limits its applications to small systems. Here, we propose a molecular dynamics (MD) methodology which can simultaneously achieve both AIMD-level high accuracy and CMD-level high efficiency. The high accuracy is achieved by exploiting deep neural network (DNN)’s arbitrarily-high precision to fit PES. The high efficiency is achieved by deploying multiplication-less DNN on a carefully-optimized special-purpose non von Neumann (NvN) computer to mitigate the performance-limiting data shuttling (i.e., ‘memory wall bottleneck’). By testing on different molecules and bulk systems, we show that the proposed MD methodology is generally-applicable to various MD tasks. The proposed MD methodology has been deployed on an in-house computing server based on reconfigurable field programmable gate array (FPGA), which is freely available at http://nvnmd.picp.vip.
Similar content being viewed by others
Introduction
As a cornerstone of atomistic-scale analysis, molecular dynamics (MD) is widely used in many fields, such as physics1,2, chemistry3,4, biology5, materials6,7, nanotechnology8,9, drug design10,11, earth science12,13, semiconductor integrated circuit14,15, and so on. Despite its importance, it is well-known that MD simulations suffer from a long-standing dilemma between accuracy and efficiency16,17,18,19,20,21. On one hand, ab-initio MD (AIMD), which is based on the first-principles density functional theory (DFT) evaluation of potential energy surface (PES), is accurate but not efficient enough to simulate large systems16,17,18. On the other hand, classical MD (CMD), which is based on artificially-crafted force fields (FF) approximation of PES, is efficient but not accurate enough in some applications19,20,21,22,23,24.
In recent years, this dilemma is mitigated, to some extent, by the machine-learning (ML) MD (MLMD)25,26,27,28,29,30,31. By evaluating PES using ML models, the efficiency of MLMD is significantly superior than that of AIMD, while keeping the AIMD-level high accuracy. Unfortunately, though several orders of magnitude faster than AIMD, the state-of-the-art MLMD is still about two orders of magnitude slower than CMD27,31,32. Until now, it is still an outstanding problem to develop an MD simulator that can simultaneously achieve AIMD-level high accuracy and CMD-level high efficiency.
It is worth noting that, MD simulations are predominantly deployed on general-purpose von-Neumann (vN) computers, where the data processing hardware (e.g., central processing unit (CPU) and graphics processing unit (GPU)) and the data storage hardware (e.g., dynamic random-access memory (DRAM)) are separate hardware components. It is well known that the vN computers suffer from severe vN bottleneck (vNB)—the majority (e.g., over 90%) of computing time and energy must be spent to repeatedly shuttle data back-and-forth between the data processing hardware and the data storage hardware33,34,35. Consequently, only a very small fraction of calculation time and energy consumption is used to perform the useful arithmetic and logic operations, leading to the overall low efficiency of vN computers33,34,35.
The severity of vNB depends on the characteristics of calculation—the more repeated data shuttling, the more performance-limiting vNB becomes. Given the typical MD duration (e.g., tMD ≈ 10−9–10−3 s) and timestep (e.g., ∆t ≈ 10−15 s), the atomistic data (e.g., positions, velocities, forces, and atomic neighbor data, etc.) must be shuttled repeatedly by a large number (e.g., nMD ≈ tMD/∆t = 106–1012) of times. Furthermore, in each MD timestep, a huge number of additional data shuttling is required to accomplish each PES evaluation36. Hindered by such nested-loop heavy-duty data shuttling, both the time efficiency and the energy efficiency of MD calculations are extremely low on general-purpose vN computers23.
However, since the invention of the first general-purpose electronic computer in the 1940s, general-purpose vN architecture has been the dominating paradigm of the mainstream computers like laptops, desktops, and supercomputers for over 7 decades37,38,39. Researchers nowadays widely use vN computers to run MD, largely because they have no other choice. Though some special-purpose MD computers have been developed22,23,40,41, they are all based on CMD and FF, whose accuracy is questionable in many important applications42,43,44,45,46. Therefore, considering the scientific and technological significance of MD1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47, it deserves serious efforts to develop a special-purpose MD computer beyond the vN paradigm, to enable efficient and accurate MD calculations in various fields.
In order to approach this goal, in this paper, we propose a paradigm shift from the established vN architecture to a non vN (NvN) architecture. By leveraging the technologies in MLMD algorithms26,27,30, artificial intelligence48,49, and NvN architecture50,51, the proposed special-purpose MD computer can simultaneously achieve both the AIMD-level high accuracy and the CMD-level high efficiency. This is achieved by deploying a deeply-revised MLMD algorithm, i.e., DeePMD26,27,28,29,30,31, (to ensure high accuracy) on a carefully-optimized NvN hardware (to ensure high efficiency). In the Section Results, the calculation accuracy and calculation efficiency are quantitatively analyzed. In the Section Discussion, a discussion is briefly made. In the Section Methods, the overall system design of the proposed special-purpose MD computer is introduced and the implementation details of the NvN architecture are presented.
Results
The performance of the proposed special-purpose non von Neumann molecular dynamics (NVNMD) computer (see Methods section for more design and implementation details) is quantitatively analyzed in this section. First, the analysis procedure is introduced (Section Analysis procedure). Then, the calculation accuracy (Section Calculation accuracy), the calculation time efficiency (Section Time efficiency), and the calculation energy efficiency (Section Energy efficiency) are analyzed quantitatively.
Analysis procedure
Any user can follow two consecutive steps to run MD on the proposed NVNMD computer, which has been released online52: (i) to train a machine learning (ML) model that can decently reproduce the PES25,26,27,28,29,30,31; and (ii) to deploy the trained ML model on the proposed NVNMD computer, then run MD there to obtain the atomistic trajectories.
ML training (i.e., step (i)) is performed on traditional vN architecture computers (e.g., CPU/GPU) by using the training codes we open-sourced online53, which are programmed purposefully based on TensorFlow54 to help users train ML models that are compatible with the unique NvN computer proposed here. To accomplish step (i), the training samples should be prepared first. This can be done by using either the active learning tools26,29,30, or the brute-force (i.e., less efficient) DFT-based AIMD sampling55,56. Then, these training samples are used as inputs of our training codes53, which output the ML models. Our training procedure is comparatively speaking more complicated than that of the established MLMD26,27—it consists of not only the continuous neural network (CNN) training of the established MLMD, but also an additional step of quantized neural network (QNN) training (Section Quantized neural network) which uses CNN results as inputs. Typically, the CNN training uses a large number of training steps (e.g., 1 × 106) with a high learning rate (e.g., 2 × 10−2); and the subsequent QNN training uses a small number of training steps (e.g., 1 × 104) and a low learning rate (e.g., 2 × 10−7), as it only needs to minimize the small error induced by quantization from CNN to QNN.
ML inference (i.e., step (ii)) is performed on the proposed NvN architecture computer, after uploading the QNN ML model to our online NVNMD system52. In the online NVNMD system, all MD settings and parameters (e.g., timestep, microcanonical/canonical/isothermal–isobaric ensemble, thermostats, etc.) are controlled by using the same input file interface of the LAMMPS package57, except that the force field is replaced by using the uploaded QNN ML model.
Six systems are used to run testing MD calculations, including three molecule systems (i.e., benzene, naphthalene, and aspirin) and three bulk systems (i.e., Sb, GeTe, and Li10GeP2S12). The training data of molecule systems are from MD17 dataset58,59,60; and those of bulk systems (i.e., Sb, GeTe, and Li10GeP2S12) come from Ref. 61, Ref. 56, and Ref. 62, respectively. The result of test accuracy, speed and energy efficiency are shown in Section Calculation accuracy, Section Time efficiency, and Section Energy efficiency, respectively.
Calculation accuracy
The high accuracy of the proposed NVNMD can be seen obviously in Table 1. The root mean square errors (RMSE) of PES fitting of benzene, naphthalene, aspirin, Sb, GeTe, and Li10GeP2S12 systems are 0.19, 0.39, 0.32, 0.14, 0.09, and 0.14 kcal mol−1, respectively. These are close to the established MLMD values in literature (Table 1), and well below the chemical accuracy threshold (1.0 kcal mol−1)63,64, indicating decent accuracy of the proposed NVNMD. As a direct comparison, in Table 1, we also collected some of the energy prediction errors of the MLMD and CMD from the existing literature, after removing the obvious outliers (e.g., |∆Ei | =2.7 kcal mol−1 for methylamine, and | ∆Ei | =1.4 kcal mol−1 for aqueous LiF pair65,66,67). It is obvious that, while CMD suffers from large PES prediction error, the proposed NVNMD has decent PES prediction accuracy, which is inherited from the highly accurate MLMD.
As shown in Table 1, the μe of MLMD is about 10−2–10−1 kcal mol−1 (i.e., single-digit meV atom−1) different from μe of NVNMD. It’s worth noting that, we copied data of different systems from literature into Table 1, since it is distracting and time-consuming to reproduce MD results of so many systems using various MD tools by ourselves. So, Table 1 only roughly shows that the MLMD and NVNMD have the similar accuracy (μe ≈ 10−1 kcal mol−1) and both of them are much more accurate than CMD (μe ≈ 101 kcal mol−1).
Though it is clear from Table 1 that NVNMD is much more accurate than CMD, it is difficult to tell the subtle accuracy difference between NVNMD and MLMD based on different datasets. Therefore, more rigorous and refined analysis is needed. Instead of directly fetching MLMD RMSE data from literature (as we did in Table 1), hereafter we start the entire procedure (including training, inference, and testing) all the way over by using the identical set of training/testing data on an identical set of systems (Table 2).
Since the proposed NVNMD is revised from DeePMD26,27,28,29,30,31 (details available in the Methods section), we use DeePMD as reference and the starting point (third row of Table 2). Then, the vN-based revisions and the NvN-based revisions are made consecutively to obtain the results of CNN (second row of Table 2) and QNN (first row of Table 2), respectively. QNN results are the final results of NVNMD in Table 1. Here, the CNN includes all quantization-free (thus more appropriate for vN) revisions, e.g., the revision of symmetry-preserving feature calculation (Eq. (11)) and the revision of nonlinear activation function calculation (Section Nonlinear activation function). The QNN further includes the quantization-based (thus more appropriate for NvN) revisions, e.g., the continuous neural network is replaced by using quantized neural network (Section Quantized neural network), the multiplication operations of floating-point numbers are replaced by using shift operations of quantized numbers (Section Multiplication-less neural network), the continuous evaluation is replaced by using discretized look-up table searching (Eq. (6)) and so on.
When applied to all systems under test, the NVNMD shows a single-digit meV atom−1 accuracy difference, compared to the MLMD (last row of Table 2). It’s worth noting that this is about 3 orders of magnitude lower than the typical interatomic bonding energy (at the order of 100 eV atom−1), and 1 order of magnitude smaller than the chemical accuracy threshold, 1.0 kcal mol−1 (i.e., about 43.4 meV atom−1)63,64. Furthermore, this high accuracy is kept while calculating elementary, binary, and quaternary systems. The test systems we chose in Table 2 include complicated crystalline-amorphous phase transitions (Sb and GeTe56,61) and atomic diffusion through quaternary system (Li-Ge-P-S62), which involve repeated chemical bond rapture and re-forming with very sophisticated PES. Therefore, these test calculations can prove that NVNMD would be accurate enough to handle many complicated MD applications.
To further test the accuracy of NVNMD, we also computed atomic forces as shown in Table 3, because atomic forces are vital to reliably obtain the MD trajectories during integration of the Newton equation. Since the molecular systems in MLMD literature in Table 3 are all evaluated based on the same dataset58,59,60, we evaluate NVNMD based on this dataset too. When compared against the ab-initio results, the atomic force mean absolute error (MAE) of NVNMD and that of MLMD show a very small difference (i.e., at the order of 101 meV Å−1), as shown in the last row of Table 3. In all test cases, this difference is even below the default atomic force threshold (e.g., 40.0 meV Å−1 in SIESTA68; 25.7 meV Å−1 in Quantum Espresso69; and 23.1 meV Å−1 in CP2K70) to determine atomic force convergence during atomic lattice relaxation or supercell geometry optimization in the mainstream ab-initio density functional theory tools.
To visually illustrate the high accuracy, the energy and force predicted by the proposed NVNMD are plotted against those predicted by the established DFT-based AIMD, as shown in Fig. 1. The high accuracy of energy and forces laid a solid foundation for reliable calculation of physical properties. As shown in Table 4, the bond length, bond angle, and vibration frequencies of the water molecule calculated by using the proposed NVNMD are very close (<1% different), compared to the those obtained by MLMD. As shown in Fig. 2, the radial distribution function, angle distribution function, and coordination number of amorphous GeTe calculated by NVNMD are also very close to those by MLMD.
To further test accuracy in GeTe system, the canonical (NVT) ensemble MD is performed as shown in Fig. 3a. The crystalline GeTe system of 512 atoms is melted from crystalline to liquid, by increasing temperature from 300 K to 1800 K. Then it is quenched from liquid to amorphous from 1800 K to 300 K. Finally, the system is recrystallized from amorphous back into crystalline by annealing it at 600 K. The entire melt-quench-anneal phase transition processes as measured in experiments71,72 can be successfully reproduced, as shown in Fig. 3a, indicating decent accuracy of the proposed NVNMD.
In panel a, the phase transition processes of GeTe is reproduced, which contains initial crystalline phase (I), liquid phase (II), amorphous phase (III), and recrystallization (IV) shown in the insets. In panel b, the mean square displacement (MSD) of Li10GeP2S12 is computed using NVNMD and the diffusion coefficient is extracted from its fitting line. The components of x, y, and z direction show the anisotropic diffusion.
For the MD test of Li10GeP2S12, the system of 900 atoms is initialized and equilibrated at 500 K using NVT ensemble for 10 ps, then simulated at microcanonical (NVE) ensemble for 100 ps. The trajectory is used to calculate diffusion coefficients of system (shown as Fig. 3b). The mean square displacement (MSD) is calculated from trajectory using following expression
The diffusion coefficient is computed by extracting the slope of MSD in Fig. 3b. We get the diffusion coefficient 2.03 × 10−10 m2 s−1 which is close to the value of Ref. 62 (i.e., 2.00 × 10−10 m2 s−1). Furthermore, according to the NVNMD results (Fig. 3b), the MSD in x and y directions are significantly smaller than that in z direction. This is in line with the anisotropic diffusion properties of Li10GeP2S12 well known in literature73,74,75.
Time efficiency
Using the six test systems, the calculation time efficiency of the proposed NVNMD is shown in Table 5. It is obvious that the time efficiency of the proposed NVNMD is around two orders of magnitude better than that of the MLMD. This means that NVNMD runs at a high speed like CMD, despite calculation complexity of MLMD is much higher than that of CMD. As schematically shown in Fig. 4, the proposed NVNMD simultaneously achieves both the CMD-level high efficiency and the MLMD/AIMD-level high accuracy. It’s worth noting that the same high efficiency is kept, no matter elementary (e.g, Sb), binary (e.g., GeTe), and quaternary (e.g., Li–Ge–P–S) systems are simulated.
The black circle, blue square and red star mark the points of (μt, μe), which are fetched from Table 1 and Table 5; the error bars and gray/blue regions are used to schematically illustrate the variations of data generated by different researchers; the horizontal dashed line is the quantum chemistry accuracy threshold, i.e., 1.0 kcal mol−1,63,64. Note: (1) we use μe values in Table 1, in order to cover more results from literature, which are based on different datasets; (2) for more rigorous accuracy analysis based on identical datasets, please refer to Tables 2 and 3.
Energy efficiency
The energy efficiency η is calculated via the formula η = T × P, where T represents the calculation time efficiency (Section Time efficiency); and P denotes the power consumption, which is measured by using a local power tester (PUUCAI P26A-10PN). The total power of the proposed NVNMD system is measured to be only about 108 W. Therefore, η of the proposed NVNMD system is around 10−5 J step−1 atom−1 (Table 6).
By using T and P of MLMD from Ref. 26,31, it can be estimated that η of the established vN-based MLMD is around 10−3–10−2 J step−1 atom−1. Here, P of MLMD is calculated by using the number of CPU/GPU used in Ref. 26,31; and we use 30 W per CPU and 250 W per GPU for estimation76,77,78. For instance, based on Summit supercomputer, MLMD uses 27.3 thousand CPU cores and 27.3 thousand GPUs31, so P ≈ 27.3 × 103 × (250 + 30) ≈ 7.6 MW (around 50–60% of Summit supercomputer’s total power consumption 13 MW) is used to achieve T ≈ 2.7 × 10−10 s step−1 atom−1.
As shown in Table 6, the calculation energy efficiency of the proposed NVNMD is around 2–3 orders of magnitude better than that of the established vN-based MLMD, with similar calculation accuracy (Table 5 and Fig. 4). Such high energy efficiency is achieved, because in NVNMD there is no repeated data shuttling, which consumes most of the energy in its vN-based counterpart33,34,35. Consequently, the calculation energy efficiency of the proposed NVNMD is comparable to that of CMD, but accuracy of the proposed NVNMD is much superior than that of CMD (Table 1).
Discussion
As an early-stage pilot version, we implemented the NVNMD on an FPGA (details available in the Methods section). It is well known that FPGA has merits of low-cost and field-programmability (i.e., short turnaround time for design revisions and iterations), and the disadvantages of limited hardware resources and low clock frequency. In contrast, the application-specific integrated circuit (ASIC) has merits of more abundant hardware resources and much higher clock frequency, and the disadvantages of high fabrication cost and long development cycle. So, FPGA is typically used as a debugging and testing tool (research phase), before taping out the ASIC (mass-production phase).
It’s worth noting that the proposed NVNMD, which simultaneously achieved high calculation accuracy (Table 1), high calculation time efficiency (Table 5), and high calculation energy efficiency (Table 6), is based on a low-end device (Xilinx xcvu9p) in the Xilinx Virtex UltraScale+ FPGA product family (Section Hardware implementation)79,80. This has three significant technological implications on the future ASIC development scenarios of the proposed paradigm (NvN-based MD).
Firstly, the NVNMD we use here is based on a low clock frequency (i.e., 250 MHz), which is about one order of magnitude lower than ordinary ASIC like the commodity-level vN-based GPU/CPU whose clock frequency can reach several GHz76. This implies that the time efficiency of NVNMD could be enhanced by another order of magnitude (i.e., C1 ≈ 101), in a straightforward fashion by boosting the clock frequency, if we move from the research phase (FPGA) to the production phase (ASIC).
Secondly, the NVNMD we use here is implemented using a rather limited amount of hardware resources (i.e., about 106 logic cells in FPGA as shown in Table 7). It is well known that a single ASIC chip could integrate around 109–1010 transistor devices (e.g., 1.6 × 1010 and 2.1 × 1010 transistor devices in one Apple M1 5 nm chip and one NVIDIA Tesla V100 12 nm chip, respectively76,81). Even though we use about 101 transistor devices to realize 1 logic cell, the ASIC could be 2 to 3 orders of magnitude more resource abundant than the FPGA we are using. By leveraging the decent parallelization scaling property of MLMD (inherited by the proposed NVNMD)82, we anticipate at least two orders of magnitude enhancement of time efficiency (i.e., C2 ≈ 102), by purely increasing the intra-ASIC parallelization.
Thirdly, the logic and arithmetic circuit can be deployed much more freely in ASIC than in FPGA, since FPGA uses more resources to ensure flexibility and programmability. Given much less constraints, the same set of functionalities can be implemented with much fewer transistor devices in ASIC than in FPGA. Therefore, by using ASIC to replace FPGA, the time efficiency can be increased by roughly ten times when the logic and arithmetic circuits are simplified (i.e., C3 ≈ 101)83.
To summarize, the FPGA-based results we presented in this paper is a very early development stage of the proposed NVNMD paradigm. Based on the architecture design we verified using FPGA in this paper, we are working on developing ASIC-based NVNMD computer, which could be around 4 orders of magnitude (i.e., C = C1 × C2 × C3 ≈ 104) more efficient than the results we showed in this paper. In another word, by moving from FPGA to ASIC, the time efficiency of NVNMD could be enhanced from 10−7 s step−1 atom−1 (Table 5) to around 10−11 s step−1 atom−1. This means that NVNMD based on a single ASIC chip (with cm2-level size and 101-102 Watt-level power) could be faster than the MLMD based on the whole Summit supercomputer (around 10−10 s step−1 atom−1, with one entire building size and 106–107 Watt-level power)31,82. Of course, during the implementation of the ASIC-based NVNMD, we need to consider factors other than speed, e.g., the generality to run all kinds of MD and the flexibility to control/dump MD simulation results, etc. All these considerations may compromise the speed somehow, which deserves future research attention. Finally, there are lots of machine learning methods other than DeePMD (e.g., SchNet84, DimeNet85, sGDML86, PaiNN87, SpookyNet88, GemNet89, NewtonNet90, UNiTE91, NequIP92, and so on). The NvN acceleration of these methods deserves future research attention, too.
Compared to the other special-purpose MD computers already existing in literature (e.g., Anton)22,23,40,41, the NVNMD proposed here is different in several aspects. Anton focuses on accelerating biology-related MD simulations and, thus, mainly implements biology-oriented classical force fields. While these classical force fields offer valuable insights to simulate biological MD problems (e.g., protein folding), they suffer serious accuracy problems in many applications in other fields, because it only has the CMD-level accuracy67,93,94,95. The accuracy of the proposed NVNMD, however, is at the AIMD/MLMD-level. Furthermore, Anton is implemented on the ASIC using advanced semiconductor technology nodes (e.g., 7 nm node), which offers much higher speed than the FPGA used in this pilot version of NVNMD.
Methods
In this Section, the overall system design of the proposed special-purpose MD computer is introduced. Section Heterogeneous parallelization describes the heterogeneous parallelization between the proposed NVNMD computer’s two major units—the master processing unit (MPU) and the slave processing unit (SPU). Section Pipeline and high-speed transmission interface discusses the high-speed transmission interface (HTI) between MPU and SPU. Section Master processing unit and Section Slave processing unit introduce the functionalities of MPU and SPU, respectively.
As the most important part of the proposed NVNMD computer, the SPU bears the predominant majority (e.g., over 99%) of the total computational load. To maximize calculation efficiency of SPU, we propose a paradigm shift from the established general-purpose vN architecture (e.g., CPU and GPU)26,27,31,56,58,62,65,66,96,97 to a special-purpose NvN architecture. The proposed NvN architecture efficiently computes the energy (Section Energy calculation) and atomic force and virial (Section Force and virial calculation) by leveraging the processing-in-memory (PIM) technology (Section Processing in memory), based on the algorithms of DeepPot-SE MLMD26,30,98 after three crucial modifications. These three modifications are indispensable to realize high calculation efficiency using very limited amount of hardware resources. Firstly, the traditional continuous neural network (CNN) widely used in MLMD is replaced by using the quantized neural network (QNN) (Section Quantized neural network). Secondly, the resource-consuming multiplication-based neural network is replaced by using a resource-economical multiplication-less neural network deliberately-designed here for the NVNMD (Section Multiplication-less neural network). Thirdly, the widely-used trigonometric function-based nonlinear activation functions are replaced by using the lightweight nonlinear activation functions specially-crafted for the NVNMD (Section Nonlinear activation function). With the help of these three significant modifications, the NvN-based SPU is implemented in a field programmable gate array (FPGA), to quantitatively test the overall performance of the proposed MD computer (Section Hardware implementation).
Heterogeneous parallelization
The MD simulation consists of a certain number of timesteps in a loop. In each timestep, there are two parts of calculations – (i) the evaluation of PES, E = E({Ri}), and atomic forces, Fi = −∇iE({Ri}); and (ii) all other calculations, including numerical integration of the Newton equation to update {Ri} and {vi}. Here, E is the system energy; Ri, vi, and Fi are the Cartesian coordinate, velocity, and force of the atom i (i = 1, 2, …, N), respectively; and N is the total number of atoms in the simulation system99.
In large-scale AIMD/MLMD simulations (i.e., N > 102), the overwhelming majority (e.g., over 99%) of computing time is spent to evaluate PES82. So, we focus on accelerating the calculation of part (i) by using the SPU based on the proposed special-purpose NvN architecture. In contrast, the calculation of part (ii) is much less computationally demanding, so the calculation of part (ii) is based on the traditional general-purpose vN architecture in MPU. This vN/NvN heterogeneous architecture is designed to leverage the flexibility of vN architecture. As a consequence, the proposed MD computer can efficiently run all kinds of MD simulations, e.g., canonical ensemble MD, microcanonical ensemble MD, isothermal–isobaric ensemble MD, enhanced sampling, and so on100,101,102,103,104.
As illustrated in Fig. 5, the calculation of each MD timestep consists of seven consecutive steps (i.e., S1, S2, S3, S4, S5, S6, and S7). In the S1, all atoms j in the vicinity of the atom i are chosen as the neighbor atoms, whose indices are stored in a neighbor list Nc(i) = {j, |Rj − Ri| < Rc} where Rc is a predefined cutoff. In the S2, the neighbor list {Nc(i)}, together with all atoms’ chemical species {Zi} and atomic coordinates {Ri}, is encoded into a compact data format using 16, 2, and 64 bits, respectively. Then, the compact data {Nc(i)}, {Zi}, and {Ri} are transmitted from the MPU to the SPU. In the S3, after receiving these compact data from MPU, the global atomic information is transferred into the ith atom’s local atomic information which includes Zi, {Zj, j ∈ Nc(i)} and {Rji} = {Ri−Rj, j ∈ Nc(i)}. In the S4, the ith atom’s energy component Ei, the atomic force components {Fji} = {∂Ei/∂Rji, j ∈ Nc(i)} and virial components {Vji} = {RjiT × Fji, j ∈ Nc(i)} are evaluated by feeding the local atomic information into the PES computation module. In the S5, the system energy E, atomic forces {Fi}, and virial V are computed by summing the contributions of each atom. The relationship can be represented as \(E = \mathop {\sum }\nolimits_{i{{{\mathrm{ = 1}}}}}^N E_i\),
and
The system consists of master processing unit (MPU), slave processing unit (SPU), and high-speed transmission interface (HTI). Calculation of each timestep of MD in the system consists of seven consecutive steps: building neighbor list (S1); encoding atomic information into compact data (S2); receiving and decoding atomic compact data (S3); evaluating potential energy surface (PES) (S4); encoding and sending data of PES (S5), decoding data of PES (S6), and other calculations, e.g., numerical integration (S7). Here, E and V represent the energy and virial of atomic system, respectively; Zi, Ri, Ei, Fi, and Nc(i) represent the chemical specie, coordinate, energy, force, and neighbor list of atom i, respectively; Rji, Fji, and Vji represent the relative coordinate, force component, and virial component between atom i and atom j.
Here, E, {Fi}, and V are encoded with 64 bits, 32 bits, and 64 bits, respectively, and written into a random-access memory (RAM) ready to be read by MPU. In the S6, MPU reads and decodes compact data from the RAM. In the S7, numerical integration of the Newton’s equation, MD thermostat, and material properties are computed.
While S1, S2, S6 and S7 are executed in the MPU, S3, S4 and S5 are executed in the SPU. During integration of these seven steps into a whole functional MD computer, the high calculation efficiency is ensured by two key design ideas. Firstly, the MPU and SPU are linked by the high-speed transmission interface (HTI), and the time spent in MPU calculation and MPU-SPU communication is minimized by the parallel pipeline computation (Section “Pipeline and high-speed transmission interface”). Secondly, the PES evaluation, which is the most time-consuming part of MD, is significantly accelerated by the processing in memory (PIM) calculations in SPU based on the proposed NvN architecture, under the coordination of MPU (Sections Master processing unit and Slave processing unit). The MPU is implemented by running a revised LAMMPS package57 on a multicore CPU (Section Master processing unit); and the SPU is implemented using an FPGA (Section Slave processing unit).
Pipeline and high-speed transmission interface
To mitigate, as much as possible, the efficiency bottleneck caused by MPU-SPU communication, four high-speed technologies are used together here. Firstly, the MPU-SPU HTI is designed as a full-duplex channel to enable simultaneous data sending and data receiving, by using two separate memory hardware units. For instance, in the FPGA implementation (Section Hardware implementation), one on-chip block random access memory (BRAM)105 of SPU is used to read data, and another on-chip BRAM of SPU is used to write data. Secondly, high-speed peripheral component interconnect express (PCIe) technology is used to send/receive data between MPU and SPU. For instance, here we use 16-lane PCIe 3.0, whose maximum bandwidth is 7.88 Gbit s−1 per lane, so the total bandwidth is as high as 15.75 GByte s−1 (Section Hardware implementation)106,107. Thirdly, the direct memory access (DMA) technology is used to transfer data between MPU and SPU. Using DMA, MPU-SPU data communication can be achieved without MPU control, so that the data transfer latency is minimized and the burden of MPU is alleviated. Fourthly, the seven consecutive steps (Fig. 5) are organized in a carefully-designed pipeline (Fig. 6).
The tS1, tS2, tS3, tS4, tS5, tS6, and tS7 are the calculation time of S1, S2, S3, S4, S5, S6, and S7, respectively, in Fig. 5. Here, tSPU = tS3 + tS4 + tS5; and tWT is the waiting time; ‘#A(B)’ stands for the sub-domain B processed by the core #A of MPU; Np is the number of cores in MPU; NSD is the number of sub-domains decomposed according to the SPU capacity. Enable signals are used to coordinate the calling of SPU.
While the first three high-speed technologies (i.e., full-duplex, PCIe, DMA) minimize the SPU’s idle time (i.e., time other than tSPU in Fig. 6a), the fourth one (i.e., pipeline) tries to vanish the SPU’s idle time. As shown in Fig. 6b, by using a small number (typically less than ten) of CPU cores in the MPU, the NvN-based SPU is always busy performing heavy-duty calculations, which is beneficial to maximize the overall efficiency.
The proposed NVNMD is based on a pipeline, in which the MPU and the SPU work in a complementary manner (Figs. 5 and 6). Thus, to maximize the overall efficiency, it is desirable to keep the SPU always busy. In another word, tSPU in Fig. 6 should be large enough (i.e., N should be large enough) to minimize Ti in Table 5. This trend can be seen in Fig. 7 – the calculation efficiency drops if N is small. Since our focus is the MD simulations of large systems (e.g., N > 104 atoms), this should not be a concern.
Master processing unit
MPU performs S1, S2, S6, and S7, as illustrated in Fig. 5. While using a MPU (e.g., CPU with NP cores) to process the system of N atoms, the whole system is spatially decomposed into NP domains with equal volume, and each core processes one domain for parallel acceleration57. To account for the interaction between atoms located within different domains, neighbor atoms of the domain Ω are copied from the neighbor domains to form a shell domain (referred to as Ωn hereafter)57. The indices of atoms inside Ωn are stored in a list IΩn = {j | RjΩ < Rc and j ∉ IΩ}, where RjΩ denotes the minimum distance between atom j and Ω; IΩ is the list of indices of atoms inside Ω. For notational convenience, the list of indices of all atoms inside Ω and Ωn is denoted as IΩa = (IΩ, IΩn), which is a combined list of IΩ and IΩn.
SPU can only store and process limited amount of data at one time due to hardware resource restriction, so domain Ω is further divided into NSD = ⌈μ × NΩ /NSPU⌉ sub-domains (denoted as ω hereafter) with equal volume, where μ is set as 2 to account for the spatial fluctuation of atom density; NΩ is the number of atoms within Ω; NSPU is set as 4096 to strike a balance between communication efficiency and resource utilization; and ⌈x⌉ is the ceiling function which rounds x to upper integer. While processing one sub-domain ω, a shell of ω (referred to as ωn hereafter) is additionally created to account for the interaction between atoms located within Ωn and sub-domains other than ω. The indices of atoms inside ωn are stored in a list Iωn = {j | Rjω < Rc and j ∉ Iω}, where Rjω denotes the minimum distance between atom j and ω; Iω is the list of indices of atoms inside ω.
Based on the abovementioned two-level decomposition (i.e., ‘A’ and ‘B’ in Fig. 6b), the enable signal is utilized to ensure that the MPU cores call the SPU in a proper order. After running S2, the pth core (p = 1, 2, …, NP) doesn’t call SPU until it receives an enable signal from its previous core (i.e., the NPth core when p = 1 and the (p−1)th core otherwise). After obtaining the results from SPU, the pth core sends an enable signal to its next core (i.e., the 1st core when p = NP and the (p+1)th core otherwise). It’s worth noting that the 1st core doesn’t require an enable signal to process its 1st sub-domain. The steps (i.e., S1, S2, S6, and S7) are discussed in detail below. In the S1, each core builds the neighbor list {Nc(i), i ∈ IΩ} of atoms located within its domain Ω.
In the S2, MPU processes the sub-domain ω’s data, including neighbor list {Nc(i), i ∈ Iω}, chemical species {Zi, i ∈ Iωa}, and coordinates {Ri, i ∈ Iωa}, where Iωa = (Iω, Iωn) is a list obtained by combining Iω and Iωn. First, the {Nc(i)} is recoded as local neighbor list. For example, if one element of {Nc(i)} is 5, and the value 5 is located at the 1st position of Iωa, this element of {Nc(i)} will be encoded as 1. In this step, the encoded {Nc(i)} is compressed from 32 bits to 16 bits. Second, {Zi} is compressed from 32 bits to 2 bits through encoding it as the order of chemical species. Third, {Ri} is encoded by multiplying 248 and rounding into 64-bit integer from 64-bit floating number. The encoded data {Nc(i)}, {Zi}, and {Ri} are stored in the buffer until they are transmitted to SPU by HTI.
In the S6, the cores of MPU decode the data fetched from SPU. Take the sub-domain ω inside domain Ω as an example, the data consists of energy Eω, atomic forces {Fi, i ∈ Iωa}, and virial Vω. Eω is decoded from 64-bit integer to 64-bit floating point number by multiplying a factor of 2−13, and then summed up to obtain the energy EΩ of domain Ω. {Fi} is decoded from 32-bit integer to 64-bit floating point number by multiplying a factor 2-25, and then summed up into the corresponding atomic forces of domain Ω (i.e., {Fi, i ∈ IΩa}) according to the index in Iωa. Vω is decoded from 64-bit integer to 64-bit floating number by multiplying a factor 2−25, and then added up to the virial VΩ of Ω.
In the S7, the cores of MPU perform numerical integration, thermostat, and so on. After the core requests SPU to evaluate PES of its domain Ω, the energy EΩ, atomic forces {Fi, i ∈ IΩa}, and virial VΩ are obtained, but they are incomplete. Therefore, the cores exchange the forces {Fi, i ∈ IΩn} of atoms located within the shell IΩn to obtain the complete atomic forces {Fi, i = 1, 2, …, N}. In addition, EΩ and VΩ are also exchanged to obtain the complete energy E and virial V of the whole system. Afterward, the atomic forces are used for numerical integration and other procedures in parallel. After S7 is finished, one timestep of MD is accomplished. The abovementioned steps repeat until all timesteps in the MD trajectory are accomplished.
Slave processing unit
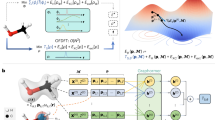
As shown in Fig. 5, the SPU runs S3, S4, and S5 in each MD timestep. Among the three categories of MD (i.e., CMD, AIMD, and MLMD), we choose MLMD to implement the proposed special-purpose MD computer, because the FF-based PES evaluation in CMD is too inaccurate and the DFT-based PES evaluation in AIMD is too sophisticated. We modify the Deep Potential-Smooth Edition (DeepPot-SE)30 and deploy it in the S4 of the SPU. The local atomic information Zi, {Zj}, {Rji} are used to compute the many-body descriptor Di which preserves the translation invariance, rotation invariance, and permutation invariance. Then, Di is used to calculate the ith atom’s energy Ei. Finally, atomic force components {Fji} are obtained by computing the negative derivative of Ei. These steps are discussed in more details below.
In order to preserve the translation invariance, the global coordinates Rj = (xj, yj, zj) are transformed into the relative coordinates Rji = Rj − Ri = (xji, yji, zji). To describe the smooth cutoff, a new coordinate uji is constructed through multiplying Rji by a cutoff function sji, which describes the contribution decay by the increase of Rji until Rc. The new coordinate is expressed as
Here, the cutoff function is defined as
where Rcs is a predefined cutoff parameter26,30. Next, multilayer perceptron (MLP) neural network \(G_{Z_j}\)108, which is called Feature NN (FeaNN) hereafter, is constructed. FeaNN has one input node and M output nodes, which is written as
The output of lth layer of MLP is
where xl, wl, bl, and ξl are input, weight, bias, and nonlinear activation function of the lth layer, respectively. The weights of FeaNN depend on chemical species Zj. Therefore, the output gji can distinguish the contribution of neighbors with different chemical species.
In order to preserve the permutation invariance, matrix Ui of size M × 4 is written as
where
In order to preserve the rotational invariance,
is defined26,30. The subset of \(D_i^\prime\) is extracted as a new M × M2 (1 ≤ M2 ≤ M) matrix Di for reducing unnecessary computational cost.
where l and k are the matrix indexes of row and column, respectively; % is modulo operation.
The total energy is written as \(E = \mathop {\sum }\nolimits_{i{{{\mathrm{ = 1}}}}}^N E_i\). The energy Ei of ith atom is only determined by its chemical species Zi and the symmetry-preserving feature Di26,27. MLP neural network is used to fit the relation between input Di and output Ei (referred to as FitNN hereafter)
Then, the force and virial can be calculated by using Eq. (2) and Eq. (3), respectively.
Energy calculation
The evaluation of PES (S4 in Fig. 5) is realized by using six calculation modules (i.e., M1, M2, M3, M4, M5, and M6) in SPU, and the atomic energy Ei is predicted during forward propagation, as shown in Fig. 8a. In the M1, {Rji2} is computed from the relative coordinate {Rji}. In the M2, {Rji2} is used to calculate the cutoff function {sji} and the outputs {gji} of FeaNN according to Eq. (5) and Eq. (6). The weights and biases of FeaNN are switched according to {Zj}. In the M3, the new coordinate {uji} is obtained by using {sji} and {Rji} (Eq. (4)). In the M4, {uji} is multiplied by {gji} to get {Uji}, and then {Uji} is summed together to get Ui (Eq. (9) and Eq. (8)). In the M5, the many-body descriptor Di is extracted from the subset of the symmetric matrix \(D_i^\prime\) which is the matrix product of Ui and UiT (Eq. (10) and Eq. (11)). In the M6, the FitNN is implemented to evaluate Ei from Di, whose weights and biases are switched according to Zi (Eq. (12)).
Six modules (i.e., M1, M2, M3, M4, M5, and M6) are used to implement the forward propagation calculation (a) from the chemical species Zi and {Zj} and relative coordinates {Rji} to the energy Ei as Section Energy calculation. The backward propagation (b) of modules is employed to calculate the component of force {Fji} and virial {Vji} as Sectioin Force and virial calculation. Each module can be divided into three submodules (c): FP, BP, and FIFO, where FP and BP are used to implement the calculation of forward propagation and backward propagation, respectively; FIFO is employed to transmit data from FP to BP.
In order to simplify the computation complexity of M2, an interpolation method is used to map from {Rji2} to {hji}, where hji is a vector of Rji2 and Zj (e.g., sji and gji). At the beginning, NT mapping tables with NM rows are built to store the data ak and bk of Rji2 and Zj in their kth row (k = 1, 2, …, NM), where NT represents the number of different chemical species; ak and bk stand for the value and derivative value of hji, respectively, when Rji2 = rk (here rk = (k−1)·Rc2 /NM). Then, when Rji2 (rk ≤ Rji2 < rk+1) and Zj is entered, one of the mapping tables is enabled according to Zj and its kth row data (i.e., ak and bk) is fetched. Finally, hji is computed via the formula hji = (Rji2 − rk) · bk + ak. The interpolation method employs a few mapping tables and multiplication to replace the complex computation of trigonometric function (Eq. (5)) and FeaNN (Eq. (6)), and it is utilized to compute {sji} and {gji} from {Rji2} in the M2. The NM is set at 1024 to strike a compromise between accuracy loss and resource usage.
The digital signal processor (DSP)109 resources are used to implement multiplication in M1, M2, M3, M4, and M5. The on-chip memory UltraRAM (URAM)105 is used to implement the mapping table in M2. Look-Up Table (LUT)110 implements FitNN’s matrix multiplication in the M6, and FitNN’s weights and biases are stored in on-chip Look-Up Table RAM (LUTRAM)110 to avoid frequent fetching from off-chip memory. There is no need to temporarily store the intermediate results in the off-chip memory since the output of the former module is the input of the latter.
Force and virial calculation
The atomic force is defined as the negative gradient of energy, so the force component {Fji} is calculated in the backward propagation of the models (i.e., M1, M2, M3, M4, M5, and M6) in SPU (Fig. 8b). In order to perform both FP and BP calculation in these modules, each module consists of three sub-modules: FP, BP, and first-in-first-out (FIFO) (Fig. 8c). FP and BP are used to execute the calculation of forward propagation and backward propagation, respectively. FIFO is employed to transmit the required intermediate results calculated by FP to BP. As illustrated in Fig. 8c, in the FP, the input of kth module is χk, and the output χk+1 is calculated according to the expression corresponding to the module in the forward propagation; in the BP, the input is ∂Ei/∂χk+1, and the output ∂Ei/∂χk is calculated by using chain rule as ∂Ei/∂χk = ∂Ei/∂χk+1 × ∂χk+1/∂χk; in the FIFO, the intermediate results χmk is transmitted in order to calculated ∂χk+1/∂χk.
Using the module structure shown in Fig. 8c, the gradient of each module’s input is computed in the backward propagation (Fig. 8b). More specially, in the M6, FIFO transmits Zi and the input of each layer’s activation function, and BP computes ∂Ei/∂Di. In the M5, FIFO transmits Ui, and BP computes ∂Ei/∂Ui. In the M4, FIFO transmits {uji} and {gji}, and BP computes {∂Ei/∂uji} and {∂Ei/∂gji}. In the M3, FIFO transmits {sji} and {Rji}, and BP computes {∂Ei/∂sji} and {∂Ei/∂uji × ∂uji/∂Rji}. In the M2, FIFO transmits {Rji2} and {Zj}, and BP computes {∂Ei/∂Rji2}. It’s worth noting that {∂sji/∂Rji2} and {∂gji/∂Rji2} are obtained by using the interpolation method proposed in Section Energy calculation. In the M1, FIFO transmits {Rji}, and BP computes {Fji} = {(∂Ei/∂Rji2 × ∂Rji2/∂Rji) + (∂Ei/∂uji × ∂uji/∂Rji)} and {Vji}.
In the BP, the matrix multiplication in the FitNN is implemented by LUT, and other multiplication is realized by DSP resources; on-chip LUTRAM is used to hold the FitNN’s parameters; on-chip URAM is used to implement the mapping tables in the M2. The parameters and the mapping tables only need to be initialized once at the start of NVNMD, and they don’t need to be fetched from off-chip memory on a regular basis. In the FIFO, on-chip URAM is also used to implement the function which transmits the data from FP to BP to avoid communication with off-chip memory. The entire procedure is designed to run in pipeline mode for optimal performance.
Processing in memory
If the energy, force, and virial (Sections Energy calculation and Force and virial calculation) are calculated on traditional vN computers, the efficiency is very low. For instance, to calculate the lth layer of MLP (i.e., Eq. (7)), it needs 11 steps, as shown in Fig. 9. These calculations are not as efficient as they could be, because the data storage unit needs to be accessed 8 times (i.e., steps 1, 2, 4, 5, 6, 8, 9, and 11 in Fig. 9a). Due to the limited size of on-chip memory (e.g., cache) of the vN processing unit (e.g., CPU/GPU), the system has to frequently access the off-chip data storage unit (e.g., main memory), which is typically two orders of magnitude slower than the processing unit (i.e., well known as the vNB).
Equation (7) is used as an example to indicate the difference between traditional vN architecture (a) and proposed NvN architecture (b) based on the processing-in-memory (PIM) technology. Here xl, wl, bl, and ξl are input, weight, bias, and nonlinear activation function of the lth layer, respectively; pl = xl × wl; sl = bl + pl.
To overcome the vNB, the proposed NvN-based SPU leverages the processing-in-memory (PIM) technology to avoid heavy-duty data shuttling111,112,113,114,115. Specifically, the logic devices and memory cells are integrated together, vanishing the data fetching latency of its vN counterparts. In the proposed NvN-based SPU, wl and bl are stored in the local on-chip memory and xl+1 is directly used as the input of the (l+1)th layer of MLP without accessing off-chip memory (Fig. 9), such that the repeated data shuttling from/to the off-chip memory (i.e., vNB) can be avoided. It’s worth noting that the parameters such as wl and bl represent the high-dimensional PES, which are material-dependent. Thus, to compute a long MD trajectory of a particular material, wl and bl are only loaded once from off-chip memory and then kept unchanged in on-chip memory during all timesteps of the MD trajectory. The logic and arithmetic operations (e.g., multipliers, adders, and activation functions) are implemented using reconfigurable circuit, to link on-chip memory cells (e.g., wl and bl). Using PIM, the calculation is pipelined without interruption of data shuttling latency, such that the calculation time is consumed purely for useful logic and arithmetic operations and, thus, the efficiency is maximized.
Quantized neural network
To implement NvN PIM (Fig. 9b), it is very hardware resource-consuming if variables (e.g., xl, wl, bl, pl, etc.) were represented using floating-point numbers116. So, despite continuous neural network (CNN) based on floating-point numbers is adopted in nearly all existing MLMD, we use the quantized neural network (QNN), which has been proposed to replace CNN in hardware devices with limited power supply and computational resources117,118,119. In the QNN, the weights and activations are quantized to save power consumption and computation resources. For example, we use quantization
for floating-point number χ, where χq is quantized value with the precision 2-γ; ⌊x⌋ is floor function which gives the greatest integer less than or equal to x. The quantization parameter γ is determined by the trade-off between accuracy and resources. We found that, by setting γ = 13, there is negligibly small accuracy loss after replacing CNN using QNN in the proposed NVNMD.
Multiplication-less neural network
To implement NvN PIM (Fig. 9b), it is also very hardware resource-consuming if the multiplication operations were realized in the arithmetic circuit directly48,49. So, despite multiplication-based neural network is adopted in nearly all existing MLMD, we propose a multiplication-less neural network, which is specially designed for the proposed NVNMD, in order to reduce the hardware circuit complexity and power consumption. Specifically, to evaluate the (3l-1)th step in Fig. 9b, the multiplication operation ‘×’ is replaced by using the bitwise shift operation ‘>>’, to evaluate
where xq is quantized input of the layer; γ is the quantization parameter (Eq. (13));
is the quantized weights of QNN; sk =−1, 0, or 1 is the sign; nk is a natural number;
is the quantization function;
is used to quantize value to exponent of 2; and ⌈x⌉ is ceiling function which rounds x to upper integer.
Obviously, in the above multiplication-less design, the multiplication operation is replaced by bitwise shift and summation operations, which are much more resource-economical and energy-saving in digital circuit. Our test shows that if K is too small (say, K = 1 or 2), there is serious accuracy loss; if K ≥ 3, the accuracy is decent to fit high-dimensional PES. So, we use K = 3 hereafter.
Nonlinear activation function
To implement NvN PIM (Fig. 9b), it is also very hardware resource-consuming if the trigonometric function-based nonlinear activation functions (e.g., tanh(x)) are implemented directly120. So, despite that these trigonometric function-based nonlinear activation functions are widely-used in existing MLMD, we design a nonlinear activation function (Fig. 10)
without trigonometric functions. In order to implement in NvN-based SPU, we redesign an activation function with continuous value and first derivative, and make it easier to use in training and prediction with fewer calculations. Because its parameters are exponents of 2, the shift operation can be used instead of the relevant multiplication and division. The most complex operation is just multiplication, not exponentiation and division in this activation function. It is easy to implement ϕ(x) in training and testing processes on vN-based and NvN-based computer. The curve of tanh(x) and ϕ(x) are compared in Fig. 10, where ϕ(x) is normalized to the range [−1, 1] by dividing 1.0625 (max value of ϕ(x)). Obviously, at the numerical value and first derivative, the tanh and ϕ(x) are similar.
Hardware implementation
To implement the heterogeneous vN/NvN (Fig. 6), we use vN-based CPU (Intel i7-10700K, 3.80 GHz, 8 cores) and NvN-based FPGA (Xilinx xcvu9p) together. The MPU in Fig. 6 is implemented by using CPU; and the SPU is implemented by using FPGA. For the neural network model deployed in SPU (Section Slave processing unit), the maximum number of neighbor atoms is set to 128; The number of FeaNN output nodes is M = 20; The Di dimension is set as 20 × 10; FitNN contains three hidden layers, each having 20 nodes. The time division multiplexing (TDM) technology is adopted to reduce the number of resources121,122. By optimizing the design, the number of resources is reduced, the timing is improved, and the clock frequency of 250 MHz is achieved. The number of resources consumed by the whole design is shown in Table 7.
Data availability
To reproduce the results in this paper, training and inference calculations are needed. The training codes and data are open-sourced at https://github.com/LiuGroupHNU/nvnmd, for generating the NVNMD-oriented inter-atomic potential models. The inference functionalities (i.e., NVNMD calculations) can be freely accessed at http://nvnmd.picp.vip.
References
Bapst, V. et al. Unveiling the predictive power of static structure in glassy systems. Nat. Phys. 16, 448–454 (2020).
Schott, S. et al. Polaron spin dynamics in high-mobility polymeric semiconductors. Nat. Phys. 15, 814–822 (2019).
Galib, M. & Limmer, D. T. Reactive uptake of N2O5 by atmospheric aerosol is dominated by interfacial processes. Science 371, 921–925 (2021).
Widmer, D. R. & Schwartz, B. J. Solvents can control solute molecular identity. Nat. Chem. 10, 910–916 (2018).
Karplus, M. & Petsko, G. A. Molecular dynamics simulations in biology. Nature 347, 631–639 (1990).
Chen, S. et al. Simultaneously enhancing the ultimate strength and ductility of high-entropy alloys via short-range ordering. Nat. Commun. 12, 4953 (2021).
Ding, W. et al. Prediction of intrinsic two-dimensional ferroelectrics in In2Se3 and other III2-VI3 van der Waals materials. Nat. Commun. 8, 14956 (2017).
Wang, Y. et al. Dynamic deformability of individual PbSe nanocrystals during superlattice phase transitions. Sci. Adv. 5, eaaw5623 (2019).
Lehtinen, O., Kurasch, S., Krasheninnikov, A. V. & Kaiser, U. Atomic scale study of the life cycle of a dislocation in graphene from birth to annihilation. Nat. Commun. 4, 2098 (2013).
Lu, S. et al. Activation pathway of a G protein-coupled receptor uncovers conformational intermediates as targets for allosteric drug design. Nat. Commun. 12, 4721 (2021).
Zhao, Y. et al. Augmenting drug–carrier compatibility improves tumour nanotherapy efficacy. Nat. Commun. 7, 11221 (2016).
Laio, A., Bernard, S., Chiarotti, G. L., Scandolo, S. & Tosatti, E. Physics of iron at Earth’s core conditions. Science 287, 1027–1030 (2000).
Steinle-Neumann, G., Stixrude, L., Cohen, R. E. & Gülseren, O. Elasticity of iron at the temperature of the Earth’s inner core. Nature 413, 57–60 (2001).
Hughes, M. A. et al. n-type chalcogenides by ion implantation. Nat. Commun. 5, 5346 (2014).
Wang, X.-P. et al. Time-dependent density-functional theory molecular-dynamics study on amorphization of Sc-Sb-Te alloy under optical excitation. npj Comput. Mater. 6, 31 (2020).
Kohn, W. & Sham, L. J. Self-consistent equations including exchange and correlation effects. Phys. Rev. 140, A1133–A1138 (1965).
Car, R. & Parrinello, M. Unified approach for molecular dynamics and density-functional theory. Phys. Rev. Lett. 55, 2471–2474 (1985).
Alavi, S. Ab initio molecular dynamics basic theory and advanced methods. By Dominik Marx and Jürg Hutter. Angew. Chem. Int. Ed. 48, 9404–9405 (2009).
Jorgensen, W. L., Maxwell, D. S. & Tirado-Rives, J. Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J. Am. Chem. Soc. 118, 11225–11236 (1996).
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A. & Case, D. A. Development and testing of a general Amber force field. J. Comput. Chem. 25, 1157–1174 (2004).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Shaw, D. E. et al. Anton, a special-purpose machine for molecular dynamics simulation. Commun. ACM 51, 91–97 (2008).
Shaw, D. E. et al. Anton 2: Raising the Bar for Performance and Programmability in a Special-Purpose Molecular Dynamics Supercomputer. in SC14: International Conference for High Performance Computing, Networking, Storage and Analysis 2015-January, 41–53 (IEEE, 2014).
Shaw, D. E. et al. Anton 3: twenty microseconds of molecular dynamics simulation before lunch. in Proceedings of the International Conference for High Performance Computing, Networking, Storage and Analysis 1–11 (ACM, 2021). https://doi.org/10.1145/3458817.3487397.
Behler, J. & Parrinello, M. Generalized neural-network representation of high-dimensional potential-energy surfaces. Phys. Rev. Lett. 98, 146401 (2007).
Wang, H., Zhang, L., Han, J. & E, W. DeePMD-kit: a deep learning package for many-body potential energy representation and molecular dynamics. Comput. Phys. Commun. 228, 178–184 (2018).
Zhang, L., Han, J., Wang, H., Car, R. & E, W. Deep potential molecular dynamics: a scalable model with the accuracy of quantum mechanics. Phys. Rev. Lett. 120, 143001 (2018).
Zhang, L., Lin, D.-Y., Wang, H., Car, R. & E, W. Active learning of uniformly accurate interatomic potentials for materials simulation. Phys. Rev. Mater. 3, 023804 (2019).
Zhang, Y. et al. DP-GEN: a concurrent learning platform for the generation of reliable deep learning based potential energy models. Comput. Phys. Commun. 253, 107206 (2020).
Zhang, L. et al. End-to-end Symmetry Preserving Inter-atomic Potential Energy Model for Finite and Extended Systems. Adv. Neural Inf. Process. Syst. 2018-December, 4436–4446 (2018).
Jia, W. et al. Pushing the Limit of Molecular Dynamics with Ab Initio Accuracy to 100 Million Atoms with Machine Learning. in SC20: International Conference for High Performance Computing, Networking, Storage and Analysis 1–14 (IEEE, 2020). https://doi.org/10.1109/SC41405.2020.00009.
LAMMPS Benchmarks. Available at: https://www.lammps.org/bench.html.
Wulf, W. A. & McKee, S. A. Hitting the memory wall. ACM SIGARCH Comput. Archit. N. 23, 20–24 (1995).
Horowitz, M. 1.1 Computing’s energy problem (and what we can do about it). in 2014 IEEE International Solid-State Circuits Conference Digest of Technical Papers (ISSCC) 57, 10–14 (IEEE, 2014).
Ielmini, D. & Wong, H. S. P. In-memory computing with resistive switching devices. Nat. Electron. 1, 333–343 (2018).
Stegailov, V., Smirnov, G. & Vecher, V. VASP hits the memory wall: processors efficiency comparison. Concurr. Comput. Pract. Exp. 31, e5136 (2019).
John von Neumann. First Draft of a Report on the EDVAC. (1945).
Electronic Numerical Integrator and Computer (ENIAC). Available at: https://en.wikipedia.org/wiki/ENIAC.
Beyond von Neumann. Nat. Nanotechnol. 15, 507–507 (2020).
Taiji, M. et al. Protein Explorer: A Petaflops Special-Purpose Computer System for Molecular Dynamics Simulations. in Proceedings of the 2003 ACM/IEEE conference on Supercomputing - SC ’03 15 (ACM Press, 2003). https://doi.org/10.1145/1048935.1050166.
Harvey, M. J., Giupponi, G. & De Fabritiis, G. ACEMD: Accelerating biomolecular dynamics in the microsecond time scale. J. Chem. Theory Comput. 5, 1632–1639 (2009).
Deringer, V. L. & Csányi, G. Machine learning based interatomic potential for amorphous carbon. Phys. Rev. B 95, 094203 (2017).
Rowe, P., Csányi, G., Alfè, D. & Michaelides, A. Development of a machine learning potential for graphene. Phys. Rev. B 97, 054303 (2018).
Zeng, J., Cao, L., Xu, M., Zhu, T. & Zhang, J. Z. H. Complex reaction processes in combustion unraveled by neural network-based molecular dynamics simulation. Nat. Commun. 11, 5713 (2020).
Li, R., Lee, E. & Luo, T. A unified deep neural network potential capable of predicting thermal conductivity of silicon in different phases. Mater. Today Phys. 12, 100181 (2020).
Rowe, P., Deringer, V. L., Gasparotto, P., Csányi, G. & Michaelides, A. An accurate and transferable machine learning potential for carbon. J. Chem. Phys. 153, 034702 (2020).
Bettini, J. et al. Experimental realization of suspended atomic chains composed of different atomic species. Nat. Nanotechnol. 1, 182–185 (2006).
Wu, B. et al. Shift: A Zero FLOP, Zero Parameter Alternative to Spatial Convolutions. in 2018 IEEE/CVF Conference on Computer Vision and Pattern Recognition 9127–9135 (IEEE, 2018). https://doi.org/10.1109/CVPR.2018.00951
Chen, H. et al. AdderNet: Do We Really Need Multiplications in Deep Learning? in 2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 1465–1474 (IEEE, 2020). https://doi.org/10.1109/CVPR42600.2020.00154
Ahn, J., Yoo, S., Mutlu, O. & Choi, K. PIM-enabled instructions. in Proceedings of the 42nd Annual International Symposium on Computer Architecture 43, 336–348 (ACM, 2015).
Mutlu, O., Ghose, S., Gómez-Luna, J. & Ausavarungnirun, R. Processing data where it makes sense: Enabling in-memory computation. Microprocess. Microsyst. 67, 28–41 (2019).
Liu, J. & Mo, P. The server website of NVNMD. (2021). Available at: http://nvnmd.picp.vip/.
Liu, J. & Mo, P. The training and testing code for NVNMD. (2021). Available at: https://github.com/LiuGroupHNU/nvnmd.
Abadi, M. et al. TensorFlow: A system for large-scale machine learning. Proc. 12th USENIX Symp. Oper. Syst. Des. Implementation, OSDI 2016 265–283 (2016). https://doi.org/10.5555/3026877.3026899
Sosso, G. C., Miceli, G., Caravati, S., Behler, J. & Bernasconi, M. Neural network interatomic potential for the phase change material GeTe. Phys. Rev. B 85, 174103 (2012).
Shi, M., Mo, P. & Liu, J. Deep Neural Network for Accurate and Efficient Atomistic Modeling of Phase Change Memory. IEEE Electron Device Lett. 41, 365–368 (2020).
Plimpton, S. Fast parallel algorithms for short-range molecular dynamics. Journal of Computational Physics 117, (1993).
Chmiela, S. et al. Machine learning of accurate energy-conserving molecular force fields. Sci. Adv. 3, e1603015 (2017).
Chmiela, S., Sauceda, H. E., Müller, K.-R. & Tkatchenko, A. Towards exact molecular dynamics simulations with machine-learned force fields. Nat. Commun. 9, 3887 (2018).
Christensen, A. S. & von Lilienfeld, O. A. On the role of gradients for machine learning of molecular energies and forces. Mach. Learn. Sci. Technol. 1, 045018 (2020).
Shi, M., Li, J., Tao, M., Zhang, X. & Liu, J. Artificial intelligence model for efficient simulation of monatomic phase change material antimony. Mater. Sci. Semicond. Process. 136, 106146 (2021).
Huang, J. et al. Deep potential generation scheme and simulation protocol for the Li10GeP2S12-type superionic conductors. J. Chem. Phys. 154, 094703 (2021).
Bogojeski, M., Vogt-Maranto, L., Tuckerman, M. E., Müller, K.-R. & Burke, K. Quantum chemical accuracy from density functional approximations via machine learning. Nat. Commun. 11, 5223 (2020).
Narayanan, B., Redfern, P. C., Assary, R. S. & Curtiss, L. A. Accurate quantum chemical energies for 133000 organic molecules. Chem. Sci. 10, 7449–7455 (2019).
Morawietz, T. & Artrith, N. Machine learning-accelerated quantum mechanics-based atomistic simulations for industrial applications. J. Comput. -Aided Mol. Des. 35, 557–586 (2021).
Zhang, P., Shen, L. & Yang, W. Solvation Free Energy Calculations with Quantum Mechanics/Molecular Mechanics and Machine Learning Models. J. Phys. Chem. B 123, 901–908 (2019).
Lu, C. et al. OPLS4: Improving force field accuracy on challenging regimes of chemical space. J. Chem. Theory Comput. 17, 4291–4300 (2021).
Soler, J. M. et al. The SIESTA method for ab initio order-N materials simulation. J. Phys. Condens. Matter 14, 2745–2779 (2002).
Giannozzi, P. et al. QUANTUM ESPRESSO: a modular and open-source software project for quantum simulations of materials. J. Phys. Condens. Matter 21, 395502 (2009).
VandeVondele, J. et al. Quickstep: Fast and accurate density functional calculations using a mixed Gaussian and plane waves approach. Comput. Phys. Commun. 167, 103–128 (2005).
Ahn, S. Phase Change Memory. (Springer International Publishing, 2018). https://doi.org/10.1007/978-3-319-69053-7.
Kolobov, A. V., Krbal, M., Fons, P., Tominaga, J. & Uruga, T. Distortion-triggered loss of long-range order in solids with bonding energy hierarchy. Nat. Chem. 3, 311–316 (2011).
Mo, Y., Ong, S. P. & Ceder, G. First principles study of the Li 10GeP 2S 12 lithium super ionic conductor material. Chem. Mater. 24, 15–17 (2012).
Marcolongo, A., Binninger, T., Zipoli, F. & Laino, T. Simulating Diffusion Properties of Solid‐State Electrolytes via a Neural Network Potential: Performance and Training Scheme. ChemSystemsChem 2, e1900031 (2020).
Kamaya, N. et al. A lithium superionic conductor. Nat. Mater. 10, 682–686 (2011).
NVIDIA Corporation. Nvidia Tesla V100 GPU Volta Architecture. White Paper 53 (2017). Available at: https://images.nvidia.cn/content/volta-architecture/pdf/volta-architecture-whitepaper.pdf.
Summit. Available at: https://www.olcf.ornl.gov/olcf-resources/compute-systems/summit/.
NVIDIA. NVIDIA V100. Available at: https://www.nvidia.com/en-us/data-center/v100/.
Xilinx. UltraScale Architecture and Product Data Sheet: Overview. Xilinx.com 1–46 (2020). Available at: https://www.xilinx.com/support/documentation/data_sheets/ds890-ultrascale-overview.pdf.
Xilinx. UltraScale+ FPGAs Product Tables and Product Selection Guide. Xilinx.com 1–11 (2021). Available at: https://www.xilinx.com/support/documentation/selection-guides/ultrascale-plus-fpga-product-selection-guide.pdf.
Ic, S. P., Dube, B., Elisabeth, S. & Scansen, D. Apple M1 System-on-Chip. systemplus.fr 1–36 (2020). Available at: https://www.systemplus.fr/wp-content/uploads/2020/12/SP20608-Apple-M1-System-on-Chip-Sample.pdf.
Lu, D. et al. 86 PFLOPS Deep Potential Molecular Dynamics simulation of 100 million atoms with ab initio accuracy. Comput. Phys. Commun. 259, 107624 (2021).
Samir, N. et al. ASIC and FPGA Comparative Study for IoT lightweight hardware security algorithms. J. Circuits, Syst. Comput. 28, (2019).
Schütt, K. T. et al. SchNet: A continuous-filter convolutional neural network for modeling quantum interactions. Adv. Neural Inf. Process. Syst. 2017-Decem, 992–1002 (2017).
Klicpera, J., Groß, J. & Günnemann, S. Directional Message Passing for Molecular Graphs. Preprint at http://arxiv.org/abs/2003.03123 (2020).
Chmiela, S., Sauceda, H. E., Poltavsky, I., Müller, K. R. & Tkatchenko, A. sGDML: Constructing accurate and data efficient molecular force fields using machine learning. Comput. Phys. Commun. 240, 38–45 (2019).
Schütt, K., Unke, O. & Gastegger, M. Equivariant message passing for the prediction of tensorial properties and molecular spectra. in Proceedings of the 38th International Conference on Machine Learning (Vol. 139 eds. Meila, M. & Zhang, T.) 9377–9388 (PMLR, 2021).
Unke, O. T. et al. SpookyNet: Learning force fields with electronic degrees of freedom and nonlocal effects. Nat. Commun. 12, 7273 (2021).
Klicpera, J., Becker, F. & Günnemann, S. GemNet: Universal Directional Graph Neural Networks for Molecules. Preprint at http://arxiv.org/abs/2106.08903 (2021).
Haghighatlari, M. et al. NewtonNet: A Newtonian message passing network for deep learning of interatomic potentials and forces. Preprint at http://arxiv.org/abs/2108.02913 (2021).
Qiao, Z. et al. UNiTE: Unitary N-body Tensor Equivariant Network with Applications to Quantum Chemistry. Preprint at http://arxiv.org/abs/2105.14655 (2021).
Batzner, S. et al. E(3)-Equivariant Graph Neural Networks for Data-Efficient and Accurate Interatomic Potentials. Preprint at http://arxiv.org/abs/2101.03164 (2021).
Kanal, I. Y., Keith, J. A. & Hutchison, G. R. A sobering assessment of small‐molecule force field methods for low energy conformer predictions. Int. J. Quantum Chem. 118, e25512 (2018).
Zgarbová, M., Otyepka, M., Šponer, J., Hobza, P. & Jurečka, P. Large-scale compensation of errors in pairwise-additive empirical force fields: Comparison of AMBER intermolecular terms with rigorous DFT-SAPT calculations. Phys. Chem. Chem. Phys. 12, 10476–10493 (2010).
Demir, H. et al. DFT-based force field development for noble gas adsorption in metal organic frameworks. J. Mater. Chem. A 3, 23539–23548 (2015).
Shen, L. & Yang, W. Molecular Dynamics Simulations with Quantum Mechanics/Molecular Mechanics and Adaptive Neural Networks. J. Chem. Theory Comput. 14, 1442–1455 (2018).
Jinnouchi, R., Karsai, F. & Kresse, G. Making free-energy calculations routine: combining first principles with machine learning. Phys. Rev. B 101, 060201 (2020).
Han, J., Zhang, L., Car, R. & E, W. Deep potential: a general representation of a many-body potential energy surface. Commun. Comput. Phys. 23, 629–639 (2018).
Allen, M. P. & Tildesley, D. J. Computer Simulation of Liquids. 1, (Oxford University Press, 2017).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Martyna, G. J., Tobias, D. J. & Klein, M. L. Constant pressure molecular dynamics algorithms. J. Chem. Phys. 101, 4177–4189 (1994).
Dullweber, A., Leimkuhler, B. & McLachlan, R. Symplectic splitting methods for rigid body molecular dynamics. J. Chem. Phys. 107, 5840–5851 (1997).
Shinoda, W., Shiga, M. & Mikami, M. Rapid estimation of elastic constants by molecular dynamics simulation under constant stress. Phys. Rev. B 69, 134103 (2004).
Tuckerman, M. E., Alejandre, J., López-Rendón, R., Jochim, A. L. & Martyna, G. J. A Liouville-operator derived measure-preserving integrator for molecular dynamics simulations in the isothermal-isobaric ensemble. J. Phys. A. Math. Gen. 39, 5629–5651 (2006).
Xilinx. UltraScale Architecture: Memory Resources User Guide (UG573). 573, 1–136 (2018).
Goldhammer, A. & Ayer, J. Jr. Understanding performance of PCI express systems. Xilinx WP350 350, 1–18 (2008).
Xilinx, P. C. I. Express for ultrascale architecture-based devices integrated block for PCIe in the ultrascale. Architecture 464, 1–15 (2015).
Hornik, K., Stinchcombe, M. & White, H. Multilayer feedforward networks are universal approximators. Neural Netw. 2, 359–366 (1989).
Xilinx. UltraScale Architecture: DSP Slice User Guide (UG579). Xilinx.com (2020). Available at: https://www.xilinx.com/support/documentation/user_guides/ug579-ultrascale-dsp.pdf.
Xilinx. UltraScale Architecture Configurable Logic Block User Guide (UG574). Xilinx.com (2017). Available at: https://www.xilinx.com/support/documentation/user_guides/ug574-ultrascale-clb.pdf.
Chi, P. et al. PRIME: a novel processing-in-memory architecture for neural network computation in ReRAM-based main memory. Proceedings of the 2016 43rd Int. Symp. Comput. Archit. ISCA 2016 27–39 (2016). https://doi.org/10.1109/ISCA.2016.13
Ghose, S., Boroumand, A., Kim, J. S., Gomez-Luna, J. & Mutlu, O. Processing-in-memory: a workload-driven perspective. IBM J. Res. Dev. 63, 3 (2019).
Sebastian, A., Le Gallo, M., Khaddam-Aljameh, R. & Eleftheriou, E. Memory devices and applications for in-memory computing. Nat. Nanotechnol. 15, 529–544 (2020).
Lu, Z., Arafin, M. T. & Qu, G. RIME: A Scalable and Energy-Efficient Processing-In-Memory Architecture for Floating-Point Operations. Proc. Asia South Pacific Des. Autom. Conf. ASP-DAC 120–125 (2021). https://doi.org/10.1145/3394885.3431524
Bavikadi, S., Sutradhar, P. R., Khasawneh, K. N., Ganguly, A. & Dinakarrao, S. M. P. A review of in-memory computing architectures for machine learning applications. Proc. ACM Gt. Lakes Symp. VLSI, GLSVLSI 89–94 (2020). https://doi.org/10.1145/3386263.3407649
Are, W., Point, F. & Layout, S. IEEE Standard 754 Floating Point Numbers. 1–7 (2011).
Gupta, S., Agrawal, A., Gopalakrishnan, K. & Narayanan, P. Deep learning with limited numerical precision. 32nd Int. Conf. Mach. Learn. ICML 2015 3, 1737–1746 (2015).
Han, S., Mao, H. & Dally, W. J. Deep Compression: Compressing Deep Neural Networks with Pruning, Trained Quantization and Huffman Coding. Int. Conf. Learn. Represent. 1–14 (2016).
Alemdar, H., Leroy, V., Prost-Boucle, A. & Petrot, F. Ternary neural networks for resource-efficient AI applications. Proc. Int. Jt. Conf. Neural Networks 2017-May, 2547–2554 (2017).
Marra, S., Iachino, M. A. & Morabito, F. C. High speed, programmable implementation of a tanh-like activation function and its derivative for digital neural networks. IEEE Int. Conf. Neural Networks - Conf. Proc. 506–511 (2007). https://doi.org/10.1109/IJCNN.2007.4371008
Zheng, D., Zhang, X., Pui, C. W. & Young, E. F. Y. Multi-FPGA Co-optimization: Hybrid Routing and Competitive-based Time Division Multiplexing Assignment. Proc. Asia South Pacific Des. Autom. Conf. ASP-DAC 176–182 (2021). https://doi.org/10.1145/3394885.3431565
Zou, P. et al. Time-Division Multiplexing Based System-Level FPGA Routing for Logic Verification. in 2020 57th ACM/IEEE Design Automation Conference (DAC) 2020-July, 1–6 (IEEE, 2020).
Lee, K., Yoo, D., Jeong, W. & Han, S. SIMPLE-NN: An efficient package for training and executing neural-network interatomic potentials. Comput. Phys. Commun. 242, 95–103 (2019).
Lu, D. et al. DP Train, then DP Compress: Model Compression in Deep Potential Molecular Dynamics. Preprint at http://arxiv.org/abs/2107.02103 (2021).
Sedova, A., Eblen, J. D., Budiardja, R., Tharrington, A. & Smith, J. C. High-performance molecular dynamics simulation for biological and materials sciences: Challenges of performance portability. Proc. P3HPC 2018 Int. Work. Performance, Portability Product. HPC, Held conjunction with SC 2018 Int. Conf. High Perform. Comput. Networking, Storage Anal. 1–13 (2019). https://doi.org/10.1109/P3HPC.2018.00004
Acknowledgements
We thank Han Wang, Linfeng Zhang, Denghui Lu, Wanrun Jiang, Jun Cheng, Yongbin Zhuang, and Jianxing Huang for their precious time to try and test NVNMD, and for their helpful suggestions to improve NVNMD. We thank experts from the DeePMD community for their helpful discussions and technical support. This work is supported by the National Natural Science Foundation of China (#61804049); the Fundamental Research Funds for the Central Universities of P.R. China; Huxiang High Level Talent Gathering Project (#2019RS1023); the Key Research and Development Project of Hunan Province, P.R. China (#2019GK2071); the Technology Innovation and Entrepreneurship Funds of Hunan Province, P.R. China (#2019GK5029); the Fund for Distinguished Young Scholars of Changsha (#kq1905012).
Author information
Authors and Affiliations
Contributions
Pinghui Mo, Chang Li, Dan Zhao, Yujia Zhang, and Jie Liu implemented and tested the NVNMD system; Mengchao Shi and Junhua Li generated the DFT data for training and testing the NVNMD system; Jie Liu proposed the idea and led the research; Pinghui Mo and Jie Liu composed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mo, P., Li, C., Zhao, D. et al. Accurate and efficient molecular dynamics based on machine learning and non von Neumann architecture. npj Comput Mater 8, 107 (2022). https://doi.org/10.1038/s41524-022-00773-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-022-00773-z