Abstract
Kohn–Sham density functional theory (DFT) is the basis of modern computational approaches to electronic structures. Their accuracy heavily relies on the exchange-correlation energy functional, which encapsulates electron–electron interaction beyond the classical model. As its universal form remains undiscovered, approximated functionals constructed with heuristic approaches are used for practical studies. However, there are problems in their accuracy and transferability, while any systematic approach to improve them is yet obscure. In this study, we demonstrate that the functional can be systematically constructed using accurate density distributions and energies in reference molecules via machine learning. Surprisingly, a trial functional machine learned from only a few molecules is already applicable to hundreds of molecules comprising various first- and second-row elements with the same accuracy as the standard functionals. This is achieved by relating density and energy using a flexible feed-forward neural network, which allows us to take a functional derivative via the back-propagation algorithm. In addition, simply by introducing a nonlocal density descriptor, the nonlocal effect is included to improve accuracy, which has hitherto been impractical. Our approach thus will help enrich the DFT framework by utilizing the rapidly advancing machine-learning technique.
Similar content being viewed by others
Introduction
Machine learning (ML) is a method to numerically implement any mapping, relationship, or function that is difficult to formulate theoretically, only from a sampled dataset. In the past decade, it has rapidly been proven to be effective for many practical problems. In studies on materials, the ML scheme is often applied to predict material properties from basic information, such as atomic configurations, by bypassing the heavy calculation required by electronic structure theory, as is done in the material informatics or the construction of atomic forcefields1,2. However, the trained ML model thus obtained often fails to be applicable for materials whose structures or component elements are not included in the training dataset. Meanwhile, ML schemes treating electron density are shown to have large transferability even with a limited training dataset3,4,5. This transferability originates from the fact that the spatial distribution of the density has more information about the intrinsic physical principles than the scalar quantities such as energy. Thus, various physical or chemical properties are expected to be predicted more accurately by considering electron density than by directly predicting them from atomic positions. Furthermore, ML has also been applied to more fundamental physical concepts: density functional theory (DFT).
Kohn–Sham (KS) DFT6,7 is the standard method for theoretical studies on the electronic structures of materials. In this theory, the solution of the KS equation
with density n(r) calculated by summing |φi(r)|2 for all the occupied states yields the TE and the density distribution of an interacting electron system under ionic potential Vion. The exchange-correlation (xc) potential Vxc[n] is ideally a functional of density; its value at r is affected by the entire density distribution {n}. However, its explicit form remains undiscovered. The mapping n to Vxc has been so far established locally as Vxc(n(r)) or semilocally as Vxc(n(r), ∇n(r), …) with or without a nonlocal augmentation by the Hartree–Fock exchange and the linear response theory. The functionals have thus been given increasingly complex analytical forms with gradually climbing the Jacob’s ladder8, but there remains the transferability issue for most existing functionals. “Problematic materials” are well-known, whose accurate DFT-based description is yet to be accomplished9,10,11. Some modern functionals were criticized for being biased toward the energy accuracy than density accuracy12, despite the fact that both are important. The functionals have been formulated so that the physical conditions, such as asymptotic behaviors and scaling properties, are satisfied, but their detailed forms rely on human heuristics, especially in the intermediate regime where the asymptotes do not apply. On the other hand, there are many accurate densities and energies available, thanks to the theoretical and experimental development, which should help us to augment the functionals toward the ideal unbiased and transferable form. In this paper, we demonstrate the development of Vxc utilizing such accurate reference data with the ML.
The pioneering studies on ML application of the density functionals have been conducted by Burke and coworkers13,14,15, where the universal Hohenberg–Kohn functional FHK[n] as a sum of the kinetic energy T[n] and the interaction energy functionals Vee[n] was constructed for orbital-free DFT, whose framework avoids the heavy calculation to solve the KS equation. Our approach contrasts to theirs, as we target Vxc and adopt the KS framework. In our previous study16, we performed the ML mapping n → Vxc for a two-body model system in one dimension trained using the accurate reference data {n, Vxc} generated by the exact diagonalization and subsequent inversion of the KS equation with varying Vion. Therein, the neural network (NN) form was adopted because of its ability to represent any well-behaved functions with arbitrary accuracy17,18. We have found that, when applied to Vion not referenced in the training, the explicit treatment of the kinetic energy suppresses the effect from spurious oscillation in the predicted Vxc, and it reduces the error of finally obtained n(r). This result suggests that the machine-learning approach to Vxc with the KS equation is a promising route. The challenge is then to make the ML of Vxc feasible for real materials.
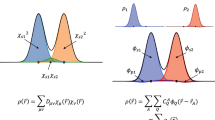
Our strategy is to restrict the functional form to the (semi-)local one, as adopted in most existing functionals for KS-DFT. Specifically, we assume the following form for the xc-energy Exc[n] to obtain the xc potential by Vxc(r) = δExc/δn(r)
where g[n](r) represents any local or nonlocal variables (descriptors) to include the effect of the density distribution around r. Most of the existing functionals follow local spin-density approximation (LSDA)19,20, generalized gradient approximation (GGA)21,22,23, or meta-GGA24,25,26, by defining g[n](r) as (n(r), ζ(r) ≡ (n↑(r)−n↓(r))/n(r))T, (n(r), ζ(r), s(r) ≡ |∇n(r)|/[2(3π2)1/3n4/3(r)])T, or \(\left( {n\left( {\mathbf{r}} \right),\zeta \left( {\mathbf{r}} \right),s\left( {\mathbf{r}} \right),\tau \left( {\mathbf{r}} \right) \equiv 1/2\mathop {\sum}\nolimits_i^{{\mathrm{occ}}} {\left| {\nabla \varphi _i\left( {\mathbf{r}} \right)} \right|^2} } \right)\)T, respectively. In this study, the xc-energy density εxc(r) is formulated using a feed-forward NN with H layers, which is a vector-to-vector mapping u → v represented by
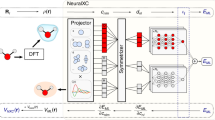
where hi represents the ith layer of the NN, and the input vector x is nonlinearly transformed by the activation function f after being linearly transformed by the weight parameters Wi and bi. To evaluate the functional derivative of δExc/δn(r) for the xc potential, we utilize the back-propagation technique27, which is an efficient algorithm to differentiate an NN applying the chain rule. This NN form thus relates {n(r)} and {Vxc(r)} to be incorporated into the KS equation. In this case, we define u as the local density descriptors g[n](r) and v as a one-dimensional vector εxc(r) (Fig. 1). The “Methods” section contains further details.
This (semi-)local NN form has practical advantages compared with the fully nonlocal form, which is adopted in the previous studies. First, the local mapping g(r) → εxc(r) is obviously transferable to any system with different size, while the fully nonlocal one is not. Second, even a few systems can provide a large amount of training data since every grid point r yields different pair of values {g(r), εxc(r)}, which can be sufficient to determine the NN parameters. As we demonstrate later, those features enable us to construct functionals whose performance is comparable to standard functionals by training with data of only a few of molecules.
We applied this ML construction approach to four types of approximation by setting the input vector g: LSDA, GGA, meta-GGA, as well as a new formulation that we call “near region approximation” (NRA) by defining g[n](r) = (n(r), ζ(r), s(r), τ(r), R(r))T, where \(R\left( {\mathbf{r}} \right) \equiv {\int} {d{\mathbf{r}}^\prime n\left( {{\mathbf{r}}^\prime } \right){\mathrm{exp}}\left( { - \left| {{\mathbf{r}} - {\mathbf{r}}^\prime } \right|/\sigma } \right)}\). Gunnarsson et al.28 demonstrated that such an averaged density around r describes εxc(r) efficiently; therefore, we added it into g of meta-GGA. Construction in such a nonlocal form has been uncommon, except for the van der Waals systems, because of the absence of appropriate physical conditions.
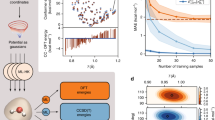
To test the performance of this approach, we constructed a functional using a few molecules to train the NN. We selected three molecules according to the following criteria: (i) the structures of the molecules should be distinct from each other and have low symmetry. (ii) Electrically polarized molecules are preferred to include to deal with cases where optimized orbitals are highly distorted from the atomic orbitals. (iii) It is most important to include at least one spin-polarized molecule, which is necessary for determining the dependency on spin-polarization ζ. Following those criteria, H2O, NH3, and NO are selected as the reference molecules. Note that the NO radical is spin-polarized. The functionals are trained to reproduce the atomization energy (AE) and the density distribution (DD) of them. We generated the training data using accurate quantum chemical calculations, i.e., the Gaussian-2 method (G2)29 for the AE and the coupled-cluster method with single and double excitations (CCSD)30,31 for the DD, which are more accurate methods than DFT. We adopt the AE for training instead of the total energy (TE), considering that typical errors by existing functionals for the TE (~hartree) are much larger than those for the AE (~eV or kcal/mol. See Table 1). The larger error implies the difficulty of reproducing TE within the (semi-)local approximations, whereas the relative energy such as the AE can be predicted more accurately due to the error cancellation. It is also worth emphasizing that DD contains abundant information of the electronic structure all over the three-dimensional space, which is expected to contribute to determining a large number of NN parameters. We selected the above conditions simply for demonstration, though how the accuracy depends on the choice of the training dataset remains a target for future studies. Ultimately, the training dataset comprised the AE and DD of H2O, NH3, and NO.
We trained the NN parameters so that εxc optimally reproduces the values of AE and DD for the training molecules through the self-consistent solution of the KS equation—Eq. 1. For this purpose, we designed a Metropolis-type Monte Carlo update method for the NN parameters. At each step, the KS equation was self-consistently solved for the three molecules to obtain their densities and total energies. Subsequently, errors from the reference values of the AE and DD were evaluated for the update of parameters. The energies of the component atoms (H, N, and O in their isolated form) were also calculated with KS-DFT using the same εxc to calculate the AE. This procedure was repeated until the error was minimized. See the “Methods” section for the exact definition of the error function and computational details.
Results and discussion
Using the trained NN-based functionals, we calculated the AE, DD, barrier heights (BH) of chemical reactions, ionization potentials (IP), and TE of the hundreds of molecular benchmark systems32,33,34, which comprise first- to third-row elements (Table 1). The performances are compared with those calculated with the existing analytic functionals. Among the various functionals developed to date, we selected the representative (semi-)local and hybrid functionals: Slater–Vosko–Wilk–Nusair (SVWN)19,20 is LSDA, Becke–Lee–Yang–Parr (BLYP)21,22 and Perdew–Burke–Ernzerhof (PBE)23 are GGAs, Tao–Perdew–Staroverov–Scuseria (TPSS)24, “Strongly Constrained and Appropriately Normed” (SCAN)25, and M06-L26 are meta-GGAs, and PBE035, B3LYP36, and M0637 are hybrid functionals.
For the wide range of unreferenced molecular systems and unreferenced quantities (BH, IP, and TE), the NN-based functionals exhibit comparable or superior performance to existing functionals in every approximation level. In particular, the nonlocal NRA-type functional is comparable to the hybrid functionals, which partly contain nonlocal effects. It is also noteworthy that the NN-based functionals are comparable to M06-L, B3LYP, or M06, which were implemented with the parameter fitting referring to more than 100 systems. This remarkable transferability with the small training dataset is nontrivial in the context of conventional ML methods predicting material properties. It reflects the advantage of our method when using electron density, which is common to any material, as the input for ML mapping. Even for unreferenced molecules, this NN-based functional would work if its local DD is similar to the one included in the reference molecules. Actually, the NN-based functional shows comparable accuracies for the AE of hydrocarbons (AEHC28) to other molecules, even though no carbon element is included in the reference molecules. Furthermore, some hydrocarbons such as benzene and butadiene have delocalized electrons owing to their conjugated structures. In LSDA or GGA, the error for them is relatively large, whereas the error becomes much smaller in meta-GGA and NRA (see Supplementary Table for detailed values). This means that, as the descriptor of DD increases, the NN gains the ability to distinguish whether the electrons are localized or delocalized.
The NN-based functionals also tend to be accurate for the unreferenced properties TE and IP, as well as for the trained property DD. The accuracy for TE and DD should be related because the Hohenberg–Kohn theorem proves their one-to-one correspondence. Accuracy for IP is also closely related to that of DD, as Perdew et al. showed that IP can be calculated accurately using potential generated from accurate density with reproducing an accurate HOMO orbital energy38. This improvement for those basic quantities would increase the accuracy of all other properties. Thus, training using density is effective not only for determining a large number of NN parameters, but also for improving their accuracy.
It is also remarkable that NN-LSDA achieves far better accuracy than SVWN. Tozer et al.39 showed that, within the local approximation, the energy density functional cannot be determined uniquely because the xc potential takes multiple values for the same local n(r), as it is actually nonlocal. From various dependencies on n(r), the conventional LSDA has been adjusted for uniform electron gas, while our functional can be contrasted as “LSDA adjusted to molecular systems”. In addition, as the approximation level increases (GGA, meta-GGA, and NRA), the multivaluedness of the mapping g → εxc reduces; thus, the accuracy tends to improve as depicted in Fig. 2. These results suggest that systematic improvement of the functional is realized by adding further descriptors to g, and by training with DDs.
The panels represent accuracy for atomization energy (AE147), reaction barrier height (BH76), and total energy (TE147) against accuracy for density distribution (DD147), corresponding to Table 1. The closed and open markers represent the accuracy of existing and the NN-based functionals, respectively.
Figure 3 also represents the improvement along with the approximation level for each benchmark molecule. For example, for the AE of HCN shown in panel (b), the NN-based functional becomes more accurate as the approximation level increases. However, for AEs of SiH4 and CCl4, the accuracy does not improve systematically. This implies that their electron DD cannot be trained sufficiently with the current reference molecules. Actually, they have tetrahedrally coordinated structures, which do not appear in the reference molecules. Large parts of their DD are considered to not appear in the reference molecules, leading to inaccurate prediction of the functional value. When we attempt further improvement by expanding the training dataset, we can find molecules that are “out of training” by this analysis and add them to the dataset.
We also applied the NN-based meta-GGA functional to the bond dissociation of C2H2 and N2, comparing them to the existing meta-GGA functionals as shown in panels (a) and (b) of Fig. 4. They agree very well, even though the NN-based functional is trained only for the molecules in equilibrium structures. This transferability for unreferenced structures is nontrivial in typical ML applications that predict the material properties directly from atomic configurations with skipping basic physical theories. This indicates the advantage of explicitly solving the KS equation, where the kinetic energy operator mitigates nonphysical noises of ML xc potential that may appear when the ML prediction is used for unreferenced inputs, thereby enhancing the transferability of the functional out of the training dataset, as shown in ref. 16.
Dissociation curves of a C2H2 and b N2 are calculated using the NN-based meta-GGA and other existing meta-GGA functionals. For C2H2, the two C–H bonds are dissociated symmetrically along the original bond direction. The horizontal axis shows the bond length, and the vertical axis shows energy relative to the atomized limit (Eatomized). The magenta dashed lines and “x” marks show the bond lengths and the atomization energies from experiments32. The “o" marks show the peaks of each curve. Density transformations of c C2H2 and d N2 due to binding are calculated using the CCSD method and DFT with the NN-based meta-GGA functional. The densities of the single CH radical and N atom are plotted as blue lines, and the densities of the bonded molecules are plotted as red lines. They are plotted along the 1D coordinate x penetrating the centers of those molecules.
Panels (c) and (d) of Fig. 4 compare the density of the molecules and their component radical or atom. The difference in density between binding and unbinding structures is well reproduced with the NN-based functional. This transferability is also nontrivial compared with conventional ML methods to predict density from nuclear coordinates, as they usually have to account for the change in environment around each nucleus in their ML models. On the other hand, as our ML method is incorporated into the KS equation, bonding can be easily reproduced, similarly to ordinary DFT studies.
In Fig. 5, the NN-based meta-GGA functional and other existing meta-GGA functionals are analyzed by plotting enhancement factors relative to the xc functional of LSDA19,20, which corresponds to the energy density in the uniform electron gas (UEG) limit:
The vertical coordinates are defined using Eq. (5). As meta-GGA-type functionals have four variables, the panels show the dependency on one of them while the others are fixed. rs ≡ (3/4πn)1/3 represents the average distance between electrons. rs is about 1 bohr in typical metals. τunif ≡ (3/10)(3π2)2/3n5/3, τW ≡ |∇n|2/8n represents τ at the UEG and single-orbital limits, respectively24.
The fixed parameters in panels (a)–(d) are set to reproduce the UEG limit, whereas the generalized gradient s is set to 0.05 in panels (a), (b), and (d) to avoid the divergence of NN-meta-GGA at s = 0. This divergence does not affect the calculation of molecules because s is greater than 0.05 in 99.8% of integration grids appearing in the training dataset. However, this divergence is recently found to cause ill-convergence when implemented in periodic boundary codes; therefore, it should be suppressed in future work. Except for this divergence, the NN-meta-GGA functional behaves similarly to the other analytical functionals. In particular, at the UEG limit shown in panels (a) and (b), we observe a trend to converge to the exact asymptotic forms as the functions of rs and ζ, which the other functionals are analytically enforced to satisfy. This result suggests that some physical conditions can be automatically reproduced through this ML approach; this property would contribute to the development of DFT with unconventional nonlocal descriptors such as those in our NRA, for which the exact asymptotic behavior is not straightforwardly derived.
In summary, we propose a systematic ML approach to the accurate and transferable xc functional for the KS-DFT. Our results suggest that improvements can be made by following a simple strategy: preparing a maximally flexible NN-based functional form and then training it with the electron DDs and energy-related properties of appropriate reference materials. The NN-based functionals trained using only a few reference molecules exhibit comparable or superior performance to the representative standard. We have revealed that employing the (semi-)local form and including DDs in the training dataset contribute to this transferability, as well as the determination of a large number of NN parameters. Furthermore, this approach enables the systematic construction of a functional with minimum assumptions, as demonstrated by the NRA functional with a nonlocal variable R, which is difficult to construct using conventional methods because of the lack of physical conditions. In Jacob’s ladder8, an approximated functional becomes accurate when including nonlocality in orbital-dependent ways, such as hybrid functionals or the random-phase approximation40,41; however, its computational cost simultaneously increases. On the other hand, our approach of introducing nonlocality retains the classical framework of solving the KS equation with the explicit functional of density, which makes the calculation more feasible for larger systems. In future studies, our approach is expected to be applied to systems that cannot be completely treated with existing functionals, such as those with dispersion interaction usually treated with van der Waals functionals, those with self-interaction error usually treated with range-separated hybrid functionals, or those with strongly correlated systems treated with DFT+U approaches42. For those problems with complicated nonlocality, our approach seems effective as it can systematically construct a maximally flexible functional form.
Methods
Structure of the NN-based functional
We formulate the xc-energy density as
The first factor \(n^{1/3}\) corresponds to the Slater exchange energy density19, and the second is from the dependency of the exchange energy of the uniformly spin-polarized electron gas on spin polarization43. They comprise the minimal physical conditions introduced to make the initial state of the NN close to the goal. The remaining correction \(G_{{\mathrm{xc}}}^{{\mathrm{NN}}}\) is constructed using the fully connected NN defined in Eq. 3 with four layers
Before applying the NN, each included element of g is preprocessed as follows:
These transformations are introduced to facilitate the optimization of NN by making g dimensionless, suppressing the change in magnitude, and regularizing the variance ranges of all input elements. For activation function f, we adopted the smooth nonlinear activation function named “exponential linear units”44, which is defined as f(x) = max(0, x) + min(0, ex − 1). The last layer hH is designed to keep the value of εxc nonpositive. The dimensions of the parameter matrices and bias vectors are as follows: dim W1 = 100 × N, dim W2 = dim W3 = 100 × 100, dim W4 = 1 × 100, dim b1 = dim b2 = dim b3 = 100, dim b4 = 1, where N represents the number of elements in g.
Functional with nonlocal DD
We suggest a functional form treating nonlocality by introducing a nonlocal descriptor:
glocal(r) represents (semi-)local descriptors such as n(r), s(r), or τ(r), while R(r) includes the weighted DD around r, with the weight function \(d\left( {{\mathbf{r}},{\mathbf{r}}^\prime } \right)\) vanishing at the \(\left| {{\mathbf{r}} - {\mathbf{r}}^\prime } \right| \to \infty\) limit. As a result of the nonlocality, the functional derivative contains an integration over the whole space:
We implemented those integrations numerically on the same grid points to those used in the exchange-correlation integration. The cost of evaluating the xc potential is proportional to the square of the system size. In this work, we defined d(r) as \({\mathrm{exp}}\left( { - \left| {{\mathbf{r}} - {\mathbf{r}}^\prime } \right|/\sigma } \right)\). σ was fixed to 0.2 bohr. This is derived from the inverse of the Fermi wavenumber, which is known to be the typical distance at which the contribution to the exchange-correlation hole at r from \({\mathbf{r}}^\prime\) decays28, in the H2O molecule estimated from the DD calculated by the CCSD calculation (averaged with respect to the number of electrons).
Training the NN-based functional
We used the Monte Carlo method by repeating the following steps to train the NN:
-
1.
At the tth iteration, add a perturbation δwt to weights wt in NN. w represents both elements in the matrices {Wi} and the vectors {bi}. Each element in δwt is generated randomly from normal distribution N(0, δw).
-
2.
Conduct the KS-DFT calculation for the target molecules and atoms to evaluate the cost function \({\mathrm{\Delta }}_{{\mathrm{err}}}^i\)in Eq. (14) using the NN-based functional with the weight parameters wt + δwt.
-
3.
According to a random number p generated from uniform distribution in (0,1) and the acceptance ratio P defined as follows, decide whether to accept or reject the weight perturbation δwt.
If P < 1: Set wt+1 = wt + δw and \({\mathrm{\Delta }}_{{\mathrm{err}}}^{{\mathrm{old}}} = {\mathrm{\Delta }}_{{\mathrm{err}}}^t\). Restart from step 2.
If p < P < 1: Set wt+1 = wt + δw. Restart from step 1.
If P < p: Set wt+1 = wt. Restart from step 1.
We repeated those steps while decreasing δw and T. The cost function Δerr is defined as
where ΔG2AE represents the absolute deviation of the AE in hartree from the G2 calculation, and E0 was set to 1 hartree. ΔCCSDn represents the error between n obtained by DFT and CCSD calculation
where Ne represents the number of electrons in molecule M. The integrations were conducted numerically on the same grid points as those used in exchange-correlation integration of the KS equation (see the “Computational details” section). E0 is adjusted such that the magnitudes of the contributions from the two terms become similar at the initial step of the training. c2/c1 determines the balance of the two terms. In this study, E0 and c2/c1 were fixed to 1 hartree and 10, respectively, for the training of any type of functional.
When training each NN-based functional, all steps were repeated for approximately 300 times. The initial T and δW were set to 0.1 and 0.01, respectively, and they were linearly reduced to 0.06 and 0.005, respectively. All whole steps were conducted in parallel with 160 threads by ISSP System C, and the weight parameters, which minimized the cost function, were ultimately adopted.
NN-size dependency of performance
To find the optimum NN size, we compared the performance among the NN-based meta-GGA functionals trained in the same way as described above with three different sizes: (α) H = 3, Nhidden = 50, (β) H = 4, Nhidden = 100, and (γ) H = 5, Nhidden = 200. Nhidden represents the size of hidden layers, h1, ..., hH−1, in Eq. 3. As shown in Fig. 6, the performance improves to a certain extent as the NN size increases, while it does not improve anymore when the size is larger than β. Therefore, we decided to use the NN with size β throughout this work for the balance of computational cost and accuracy. For the development of a further accurate functional, finer tuning should be done.
AE147 MAE and DD147 ME (see the caption of Table 1 for their definitions) of NN-based meta-GGA functionals implemented with different NN sizes are plotted. The definition of each NN size is shown in the main text.
Computational details
All DFT and CCSD calculations in our work were implemented using PySCF version 1.6.245, and the 6–311++G(3df,3pd) basis set was used both in training the NN-based functionals and in testing the accuracies of the functionals. For the DFT calculations, the default settings of PySCF were used throughout. For the integration of xc potentials and energy densities, we used the angular grids of Lebedev et al.46 and the radial grids of Treutler et al.47. The numbers of radial and angular grids were set to (50, 302) for H, (75, 302) for second-row elements, and (80–105, 434) for third-row elements. For molecules, Becke partitioning48 was used. The NN-based functionals could cause a convergence issue owing to poor extrapolation when they are applied to density far from that included in the training dataset; therefore, the initial density guess of self-consistent DFT should be sufficiently close to the final destination. In this work, initial guesses of KS density were given by a superposition of atomic density, which successfully made the calculation converge.
We used the Pytorch version 1.1.0 for the NN implementation and took its derivative via the back-propagation technique49.
Data availability
The individual values for all benchmark systems in Table 1 are listed in Supplementary Table.
Code availability
The trained NN parameters are available at https://github.com/ml-electron-project/NNfunctional with usages implemented in PySCF codes.
References
Xie, T. & Grossman, J. C. Crystal graph convolutional neural networks for an accurate and interpretable prediction of material properties. Phys. Rev. Lett. 120, 145301 (2018).
Behler, J. & Parrinello, M. Generalized neural-network representation of high-dimensional potential-energy surfaces. Phys. Rev. Lett. 98, 146401 (2007).
Brockherde, F. et al. Bypassing the kohn-sham equations with machine learning. Nat. Commun. 8, 872 (2017).
Grisafi, A. et al. Transferable machine-learning model of the electron density. ACS Cent. 5, 57–64 (2018).
Chandrasekaran, A. et al. Solving the electronic structure problem with machine learning. Npj Comput. Mater. 5, 22 (2019).
Hohenberg, P. & Kohn, W. Inhomogeneous electron gas. Phys. Rev. 136, B864–B871 (1964).
Kohn, W. & Sham, L. J. Self-consistent equations including exchange and correlation effects. Phys. Rev. 140, A1133–A1138 (1965).
Perdew, J. P. & Schmidt, K. Jacob’s ladder of density functional approximations for the exchange-correlation energy. AIP Conf. Proc. 577, 1–20 (2001).
Mardirossian, N. & Head-Gordon, M. Thirty years of density functional theory in computational chemistry: an overview and extensive assessment of 200 density functionals. Mol. Phys. 115, 2315–2372 (2017).
Gillan, M. J., Alfè, D. & Michaelides, A. Perspective: how good is dft for water? J. Chem. Phys. 144, 130901 (2016).
Ekholm, M. et al. Assessing the scan functional for itinerant electron ferromagnets. Phys. Rev. B 98, 094413 (2018).
Medvedev, M. G., Bushmarinov, I. S., Sun, J., Perdew, J. P. & Lyssenko, K. A. Density functional theory is straying from the path toward the exact functional. Science 355, 49–52 (2017).
Snyder, J. C., Rupp, M., Hansen, K., Müller, K.-R. & Burke, K. Finding density functionals with machine learning. Phys. Rev. Lett. 108, 253002 (2012).
Snyder, J. C. et al. Orbital-free bond breaking via machine learning. J. Chem. Phys. 139, 224104 (2013).
Li, L. et al. Understanding machine-learned density functionals. Int. J. Quantum Chem. 116, 819–833 (2016).
Nagai, R., Akashi, R., Sasaki, S. & Tsuneyuki, S. Neural-network Kohn-Sham exchange-correlation potential and its out-of-training transferability. J. Chem. Phys. 148, 241737 (2018).
Cybenko, G. Approximation by superpositions of a sigmoidal function. Math. Control Signals Syst. 2, 303–314 (1989).
Hornik, K. Approximation capabilities of multilayer feedforward networks. Neural Netw. 4, 251–257 (1991).
Slater, J. C. A simplification of the hartree-fock method. Phys. Rev. 81, 385–390 (1951).
Vosko, S. H., Wilk, L. & Nusair, M. Accurate spin-dependent electron liquid correlation energies for local spin density calculations: a critical analysis. Can. J. Phys. 58, 1200–1211 (1980).
Becke, A. D. Density-functional exchange-energy approximation with correct asymptotic behavior. Phys. Rev. A 38, 3098–3100 (1988).
Lee, C., Yang, W. & Parr, R. G. Development of the colle-salvetti correlation-energy formula into a functional of the electron density. Phys. Rev. B 37, 785–789 (1988).
Perdew, J. P., Burke, K. & Ernzerhof, M. Generalized gradient approximation made simple. Phys. Rev. Lett. 77, 3865–3868 (1996).
Tao, J., Perdew, J. P., Staroverov, V. N. & Scuseria, G. E. Climbing the density functional ladder: Nonempirical meta–generalized gradient approximation designed for molecules and solids. Phys. Rev. Lett. 91, 146401 (2003).
Sun, J., Ruzsinszky, A. & Perdew, J. P. Strongly constrained and appropriately normed semilocal density functional. Phys. Rev. Lett. 115, 036402 (2015).
Zhao, Y. & Truhlar, D. G. A new local density functional for main-group thermochemistry, transition metal bonding, thermochemical kinetics, and noncovalent interactions. J. Chem. Phys. 125, 194101 (2006).
Rumelhart, D. E., Hinton, G. E. & Williams, R. J. Learning representations by back-propagating errors. Nature 323, 533–536 (1986).
Gunnarsson, O., Jonson, M. & Lundqvist, B. I. Descriptions of exchange and correlation effects in inhomogeneous electron systems. Phys. Rev. B 20, 3136–3164 (1979).
Curtiss, L. A., Raghavachari, K., Trucks, G. W. & Pople, J. A. Gaussian-2 theory for molecular energies of first-and second-row compounds. J. Chem. Phys. 94, 7221–7230 (1991).
Čížek, J. On the correlation problem in atomic and molecular systems. calculation of wavefunction components in ursell-type expansion using quantum-field theoretical methods. J. Chem. Phys. 45, 4256–4266 (1966).
Purvis, G. D. & Bartlett, R. J. A full coupled-cluster singles and doubles model: the inclusion of disconnected triples. J. Chem. Phys. 76, 1910–1918 (1982).
Curtiss, L. A., Raghavachari, K., Redfern, P. C. & Pople, J. A. Assessment of gaussian-2 and density functional theories for the computation of enthalpies of formation. J. Chem. Phys. 106, 1063–1079 (1997).
Lynch, B. J., Zhao, Y. & Truhlar, D. G. Effectiveness of diffuse basis functions for calculating relative energies by density functional theory. J. Phys. Chem. A 107, 1384–1388 (2003).
Zhao, Y., González-García, N. & Truhlar, D. G. Benchmark database of barrier heights for heavy atom transfer, nucleophilic substitution, association, and unimolecular reactions and its use to test theoretical methods. J. Phys. Chem. A 109, 2012–2018 (2005).
Ernzerhof, M. & Scuseria, G. E. Assessment of the perdew-burke-ernzerhof exchange-correlation functional. J. Chem. Phys. 110, 5029–5036 (1999).
Stephens, P. J., Devlin, F. J., Chabalowski, C. F. & Frisch, M. J. Ab initio calculation of vibrational absorption and circular dichroism spectra using density functional force fields. J. Phys. Chem. 98, 11623–11627 (1994).
Zhao, Y. & Truhlar, D. G. The m06 suite of density functionals for main group thermochemistry, thermochemical kinetics, noncovalent interactions, excited states, and transition elements: two new functionals and systematic testing of four m06-class functionals and 12 other functionals. Theor. Chem. Acc. 120, 215–241 (2008).
Perdew, J. P., Parr, R. G., Levy, M. & Balduz, J. L. Jr Density-functional theory for fractional particle number: derivative discontinuities of the energy. Phys. Rev. Lett. 49, 1691 (1982).
Tozer, D. J., Ingamells, V. E. & Handy, N. C. Exchange-correlation potentials. J. Chem. Phys. 105, 9200–9213 (1996).
McLachlan, A. & Ball, M. Time-dependent Hartree–Fock theory for molecules. Rev. Mod. Phys. 36, 844 (1964).
Furche, F. Molecular tests of the random phase approximation to the exchange-correlation energy functional. Phy. Rev. B 64, 195120 (2001).
Burke, K. Perspective on density functional theory. J. Chem. Phys. 136, 150901 (2012).
Oliver, G. L. & Perdew, J. P. Spin-density gradient expansion for the kinetic energy. Phys. Rev. A 20, 397–403 (1979).
Clevert, D. A., Unterthiner, T. & Hochreiter, S. Fast and accurate deep network learning by exponential linear units (elus). ICLR2016 (2015).
Sun, Q. et al. Pyscf: the python-based simulations of chemistry framework. Wiley Interdiscip. Rev. Comput. Mol. Sci. 8, e1340 (2017).
Lebedev, V. I. & Laikov, D. A quadrature formula for the sphere of the 131st algebraic order of accuracy. in Dokl. Math. vol. 59, 477–481 (Pleiades Publishing, Ltd., 1999).
Treutler, O. & Ahlrichs, R. Efficient molecular numerical integration schemes. J. Chem. Phys. 102, 346–354 (1995).
Becke, A. D. A multicenter numerical integration scheme for polyatomic molecules. J. Chem. Phys. 88, 2547–2553 (1988).
Paszke, A. et al. Automatic differentiation in pytorch. NeurIPS (2017).
Acknowledgements
R.N. thanks Takahito Nakajima and Yoshiyuki Yamamoto for their enlightening comments. Part of the calculations were performed at the Supercomputer Center at the Institute for Solid State Physics at the University of Tokyo.
Author information
Authors and Affiliations
Contributions
R.N. designed the method, implemented the codes, and performed the calculation. All authors contributed to developing the concept, analyzing the results, and writing the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nagai, R., Akashi, R. & Sugino, O. Completing density functional theory by machine learning hidden messages from molecules. npj Comput Mater 6, 43 (2020). https://doi.org/10.1038/s41524-020-0310-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-020-0310-0
This article is cited by
-
Advances of machine learning in materials science: Ideas and techniques
Frontiers of Physics (2024)
-
Linear Jacobi-Legendre expansion of the charge density for machine learning-accelerated electronic structure calculations
npj Computational Materials (2023)
-
Machine learning and density functional theory
Nature Reviews Physics (2022)
-
Computational design of magnetic molecules and their environment using quantum chemistry, machine learning and multiscale simulations
Nature Reviews Chemistry (2022)
-
Rapidly predicting Kohn–Sham total energy using data-centric AI
Scientific Reports (2022)