Abstract
Recent progress in material data mining has been driven by high-capacity models trained on large datasets. However, collecting experimental data (real data) has been extremely costly owing to the amount of human effort and expertise required. Here, we develop a novel transfer learning strategy to address problems of small or insufficient data. This strategy realizes the fusion of real and simulated data and the augmentation of training data in a data mining procedure. For a specific task of grain instance image segmentation, this strategy aims to generate synthetic data by fusing the images obtained from simulating the physical mechanism of grain formation and the “image style” information in real images. The results show that the model trained with the acquired synthetic data and only 35% of the real data can already achieve competitive segmentation performance of a model trained on all of the real data. Because the time required to perform grain simulation and to generate synthetic data are almost negligible as compared to the effort for obtaining real data, our proposed strategy is able to exploit the strong prediction power of deep learning without significantly increasing the experimental burden of training data preparation.
Similar content being viewed by others
Introduction
There has been considerable interest over the last few years in accelerating the process of materials design and discovery1. In the past decade, a new interdisciplinary research field called materials informatics, which combines data and information science with materials science, has led to an increasing number of successful materials discoveries2,3,4,5,6,7. Machine learning methods have played a key role in many of these studies. In general, under the premise of ensuring data quality, the larger the training dataset is, the more accurate the trained machine learning model becomes. This is especially the case for deep neural networks, which have been shown to have exceptional predictive performance when trained with a large amount of data. As a result, some accelerated methods for data accumulation, such as high-throughput computation and high-throughput experiments, have been developed to build large databases. However, in many materials studies, especially for new materials, we still face the dilemma of lacking high-quality data to construct reliable machine learning models8. The major obstacle comes from the typically time-consuming or technically difficult process of collecting experimental data (real data). Although computational data may have a lower cost than experimental data, there is still a significant gap between the two types of data for many applications in materials science.
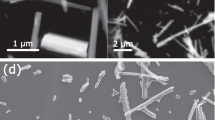
Material microstructure data are an important type of material data to build the intrinsic relationship between composition, structure, process, and properties that is fundamental to material design. Therefore, the quantitative analysis of microstructures is essential for the control of the material properties and performances of metals or alloys in various industrial applications9,10. Microscopic images are often used to understand the important structures of a material related to certain properties of interest. One of the key steps in the design process is microscopic image processing using computational algorithms and tools to efficiently extract useful information from the images11,12. For example, image segmentation13, which is a task that generates pixel-wise labels of a raw image, is commonly used to extract significant information from microscopic images in the field of material structure characterization14. For example, in polycrystalline iron (Fig. 1), which is a fundamental and popular subject in practical materials research, the objective of image processing algorithms is to detect and depart each grain from raw microscopic images to obtain accurate descriptions of the microstructure, such as geometric and topological characteristics.
Recent progress in segmentation of material microscopic image15,16,17 has been driven by high-capacity models trained on large datasets. Unfortunately, the generalization performance of these models has been hindered by the lack of a large training data due to the time-consuming labeling of material microscopic images. Generating data of images with pixel-wise semantic labels is known to be very challenging due to the amount of human effort and expertise required to trace the object boundaries accurately18. Therefore, high quality large training datasets are often difficult to obtain for a new task of material design.
In this work, we develop a novel transfer learning strategy to address problems of small or insufficient data in material data mining. This strategy realizes the fusion of real data and simulated/calculated data based on transfer learning theory and the augmentation of training data in a data mining procedure so that a classification or regression model with better performance can be established based on only a small set of real data. We explore the use of a 3D simulated model to create large-scale pixel-accurate data to train image segmentation models. For example, as shown in Fig. 1c, a Monte Carlo Potts model can represent the polycrystalline microstructure of materials because it possesses geometrical and topological microstructure characteristics of grains similar to those of the real grain structure19,20. It is easy to obtain large data with pixel-level labels from a simulated 3D model using computational methods (Fig. 1d, e). However, the acquired contents in the simulated data are unrealistic, due to some theoretical approximations and simplifications in the modeling process, which causes a challenge to simply apply simulated data to a real microscopic image processing system. To fill in the missing information in the simulated data, we use an image-to-image conversion technique to transfer simulated images into synthetic images that incorporates features from real images. This image-to-image conversion can be simply described as an image style transfer model that takes in a simulated image as input and outputs a realistic image21,22. We exploit the transformation power of image style transfer models that input and output images shared the same underlying global structure even if the appearances are very different. Specifically, we pick generative adversarial networks (GANs) as the underlying generative model based on prior studies that demonstrated its outstanding performance on the task of image style transfer23,24,25,26,27.
In general, our presented strategy can create synthetic data by fusing the images obtained from simulating the physical mechanism of grain formation and the “image style” information in real images, as outlined in Fig. 2. The acquired synthetic images are more realistic than the simulated images as they captures features in real images through the transfer learning process. Based on quantitative analyses using the polycrystalline data, we show that the acquired synthetic data can be used as a partial replacement to the real data when training a segmentation model. In particular, we demonstrate that the model trained with synthetic data and only 35% of the real data can achieve competitive performance to the model trained on all of the real data. Because the time required to perform grain simulation and to generate synthetic images are almost negligible as compared with the effort for obtaining real data, our proposed strategy is able to exploit the strong prediction power of deep learning without significantly increasing the experimental burden of training data preparation. Besides, we believe that this strategy can be easily applied to other data mining tasks, structured data analysis for example, or even those outside of materials science.
The presented strategy can create synthetic data by fusing the images obtained from simulating the physical mechanism of grain formation and the “image style” information in real images. Then the acquired synthetic data can be used as data augmentation for the training procedure of the machine learning model.
Results
Datasets
We define a data point as a pair of an image and its corresponding label. Real dataset denotes real images taken from experiment with manually annotated labels; simulated dataset refers to Monte Carlo Potts simulated images with labels directly extracted from the simulations; synthetic dataset denotes synthetic images generated from our image style transfer model with corresponding simulated labels. Segmentation models trained with different data sets are used to provide predicted labels for a separated test images obtain from experiment. Throughout the paper, we consider the manually annotated labels as ground true (or true labels) because these are the best references we can get and we have confirmed the consistency of results between different experts.
Real dataset
The real dataset contains a total of 136 serial section optical images of polycrystalline iron with a resolution of 2800 × 1600 pixels, of which the ground truth has 2 semantic classes (grain and grain boundary) and is manually labeled by material scientists. The dataset is randomly split into 100 training and 36 test data. The original images were pre-processed into small images (patches) with a size of 400 × 400 pixels in order to reduce the computational burden. Finally, the training set consists of 2800 patches with resolution of 400 × 400 pixels, cropped from the original 100 training images, while the test set consists of 1008 patches cropped from the 36 test images. The test data is separated at the beginning of our experiment and is not used for any of the model training, including the image style transfer models and segmentation models.
Simulated dataset
We established a large 3D simulated model of the polycrystalline materials using the Monte Carlo Potts model. Then, 2D images were obtained by slicing a simulated 3D image in the normal direction (Fig. 1d). Finally, we retained only the boundary pixels of each grain to obtain the labels for the simulated 2D images (Fig. 1e). The final simulated dataset contains a total of 28,800 images, which was one order of magnitude greater than the real dataset. During the simulation process, we ensured that the geometric and topological information of the simulated data was statistically consistent with that of the real image. However, the simulated data contains only grain boundary information with no defects or noises. For the real images, the range of pixel values of the grain boundary obtained by optical microscopy was [0, 255], and the specific pixel value was affected by grain appearance, the light intensity of the microscope and noise introduced during the sample preparation. For the simulated 2D images, the range of pixel values was [0, N], with N denoting the total number of grains in the 3D simulated model, which was controlled by the grain-growth simulation model. The pixel values are basically the identification numbers for the grains. Simulated data cannot be directly used in machine learning-based algorithms because there are differences in the nature of real and simulated data.
Synthetic dataset
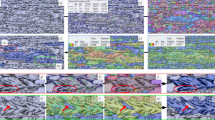
We trained our image style transfer model using real dataset. Then, we applied our model to transform all the labels images of the simulated 2D images (simulated label) into synthetic images. As shown in Fig. 3, there are four columns of images: from left to right are real image, simulated image, simulated label and synthetic image. The synthetic image has both label information and an “image style” similar to those of the real image. It can be used as data augmentation for the real data in data mining or machine learning tasks.
We compare the time consumption of the two image production methods, i.e., experiment and synthetization, in Table 1. Due to the complex experimental procedures, including sample preparation, polishing, etching, and photographing, the real image takes the longest time, ~1200 s per image. It should be noted that we do not consider the increase in time cost caused by failed experiments. In reality, the experimental process is likely to require even longer time.
The preparation of synthetic dataset includes three steps: the design and construction of simulation model, the training of style transfer model, and the generation of synthetic image. By virtue of a high-speed computer system, the construction of simulation model cost only 1% of the experimental time, ~12 s per simulated data. And the training of style transfer model cost ~23 h, which translated to ~3 s per synthetic image. Note that the training time of style transfer model depends on the volume of training data. In practice, a smaller training set could be enough to obtain a reasonably good model, meaning that a shorter time would be needed. Finally, when we have a trained style transfer model, the generation of one synthetic image takes ~0.1 s during the inference of the generative adversarial network. As a result, the time cost of setting up a synthetic dataset is approximated 1% of the time cost of getting experimental data.
Evaluation models and metrics
Model setting
There are two deep learning-based models in our experiment: image style transfer model and image segmentation model.
For image style transfer model, we used pix2pix23 to transform simulated label to synthetic image and the detail structure can be found in latter section. During the training stage, we set the batch size equal to 8 and optimized the model by Adam with an initial learning rate of 2 × 10−4 in 200 epochs.
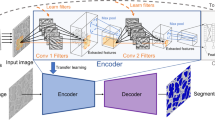
For image segmentation model, U-net28 is the most commonly used supervised learning method in the field of materials and medical image processing29. Therefore, we used U-net as our baseline to compare with data augmentation algorithms. U-net is an encoder–decoder network; the input goes through a series of convolution-pooling-normalization group layers until it reaches the bottleneck layer, where the underlying information is shared with the output. U-net joins the layer-skip connection to transfer the features extracted from the down-sampling layers directly to the upper sampling layers, thus making the pixel location of the network more accurate. During the training stage, we jointly trained the model on real and synthetic dataset using batch gradient descent with mini-batches of 8 images, including 4 real and 4 synthetic images, which is the same process used in the work18. It took 28,000 iterations to converge with a learning rate of 1 × 10−4. Although the training pattern is important for network training, this paper does not discuss this issue because it is beyond the scope of our topic. All models were trained with the same pattern to ensure fairness.
Metrics
Grain segmentation is an instance segmentation (or dense segmentation) task. A successful algorithm has to detect and depart each grain in an image. For example, there are ~57 grains in each 400 × 400 real image. Based on the segmentation results, researchers can extract size and shape distribution of grains from the image to build the relationship between microstructure and macroscopic material performance. In practice, different types of noises are introduced to a sample during the preparation step, which seriously influences the grain segmentation algorithms. We should choose an effective metrics to evaluate the performance of algorithms in this task. Discussion of different image segmentation metrics is included in Supplementary Note 1 with Supplementary Fig. 1. We decided to use two effective metrics to evaluate our algorithm: mean average precision (MAP)30,31 and adjusted rand index (ARI)32,33,34.
MAP is a classical measure in image segmentation and object detection tasks. In this paper, we evaluate it at different intersection over union (IoU) thresholds. The IoU of a proposed set of object pixels and a set of true object pixels is calculated as:
The metric sweeps over a range of IoU thresholds at each point, calculating an average precision value. The threshold values range from 0.5 to 0.95 with a step size of 0.05: (0.5, 0.55, 0.6, 0.65, 0.7, 0.75, 0.8, 0.85, 0.9, 0.95). In other words, at a threshold of 0.5, a predicted object is considered a “hit” if its IoU with a ground truth object is >0.5. Generally, it can be considered that the segment is correct when its IoU is >0.5. The other higher value is aimed at ensuring the correct results.
At each threshold value t, a precision value is calculated based on the number of true positives (TP), false negatives (FN), and false positives (FP) resulting from comparing the predicted object to all ground truth objects. The average precision of a single image is then calculated as the mean of the above precision values at each IoU threshold.
Finally, the MAP score returned by the metric is the mean taken over the individual average precisions of each image in the test set.
ARI is the corrected-for-chance version of the rand index (RI), which is a measure of the similarity between two data clustering methods32,33,34. From a mathematical standpoint, ARI or RI is related to accuracy. In addition, image segmentation can be considered a clustering task that splits all pixels in an image into n partitions or segments.
Given a set S of n elements (pixels) and two groupings or partitions of these elements, namely, X = {X1, X2, …, Xr} (a partition of S into r subsets) and Y = {Y1, Y2, …, XS} (a partition of S into s subsets), the overlap between X and Y can be summarized in a contingency table [nij] (see Table 2), where each entry nij denotes the number of objects in common between Xi and Yj: \(n_{ij} = |X_i \cap Y_j|\). For the image segmentation task, X and Y can be treated as ground truth and predicted result, respectively.
The ARI is defined as follows:
where nij, ai, bj are values from the contingency table and \(\left( {\begin{array}{*{20}{c}} n \\ 2 \end{array}} \right)\) is calculated as n(n − 1)/2.
The MAP is a strict and effective metric because it is the mean score of each threshold, so the score is always smaller than the ARI.
For all models, the higher the metrics, the better the model is. For fair comparison, all the models were evaluated on the same real test set initially separated out from the training process.
Our implementation of this algorithm was derived from the publicly available Python35, Pytorch framework36, and OpenCV toolbox37. The image style transfer model and U-net’s training and testing were performed on Nvidia DGX Station using 4 NVIDIA Tesla V100 GPU with 128 GB memory.
Image segmentation by the proposed augmentation method
First, we compared the performance of three conventional image processing algorithms; a threshold-based method called OTSU38, a boundary-based method called Canny39 and a morphology-based method called Watershed40. Second, we chose K-means41 as a representative for unsupervised learning method. Third, we chose U-net as a representative for deep learning method (Real100% in Table 3). As shown in Table 3, due to the high capability of feature extraction, the U-net achieves the best performance compared with unsupervised learning method and traditional image processing algorithms. And some visualization results can be found in Supplementary Note 2 with Supplementary Fig. 2.
Finally, we explored the use of a simulated dataset as data augmentation to replace the real dataset for training a neural network. As shown in Table 3, using the same real test set initially separated out from the training, we compared the performance of the model trained on the whole training set of real data, named the Real100% dataset, and the performance on the whole simulated dataset, named the Simulated100% dataset. The subscript denotes the percentage of the specific dataset used in the training set. We found that simply using the Simulated100% dataset achieved poor performance when compared with the Real100% dataset. In addition, the performance was degraded when we mixed these two datasets as a training set. We assume that there are two possible reasons. One possibility is that we did not have sufficient simulated data to achieve proper model training. However, additional experiments (Supplementary Table 1) shown in Supplementary Note 3 indicated that models trained with simulated data performed very well on the simulated test data, so we ruled out this possibility. The other reason could be that the simulated data were unrealistic, i.e., containing only grain boundary information without any defects that may appear in real images. This problem can be addressed by the image style transfer model.
We used the image style transfer model to create synthetic image by fusing the pixel-level label of simulated label and the “image style” information of real image. We believe that this processing will make the simulated data more realistic and could be used as data augmentation for real data in material data mining.
We evaluated the performance of the image style transfer model using MAP and ARI. There are two stages in our approach; image style transfer and image segmentation, both of which need to be trained. The main motivation of our study is to reduce the demand of real data for training the segmentation network. Therefore, we started our experiment from using only 5% of the whole real datasets and increased the amount by 5% until we reached the same performance as the Real100% case. First, we used the selected amount of real data to train an image style transfer model. Then, we used the trained image style transfer model to convert all images of simulated label to a set of synthetic images. The simulated labels and synthetic images form the synthetic dataset, denoted as \({\mathrm{Synthetic}}_{100{\mathrm{\% }}}^{5{\mathrm{\% }}}\) if we used 5% real data. The superscript in \({\mathrm{Synthetic}}_{100{\mathrm{\% }}}^{5{\mathrm{\% }}}\) denotes the proportion of real dataset used to train the style transfer model and the subscript of Real5% or \({\mathrm{Synthetic}}_{100{\mathrm{\% }}}^{5{\mathrm{\% }}}\) denotes the proportion of real or synthetic dataset, respectively, used to train the segmentation network.
We show the performance of model trained with synthetic dataset in Table 4. We compared the performance with models trained on real dataset and a mixture of both. We found that the case of using mixed dataset performed better than the case of using only real dataset. If we have only 5% of the real dataset, the performance will increase by ~8% on MAP and 10% on ARI after using the synthetic dataset as data augmentation. This finding suggests that using synthetic dataset as data augmentation will bring significant performance improvement, especially when there is only a small amount of real data.
As shown in Table 4, with the increase in the amount of real data, the performance of the models improved for both cases (using only the real data or using the mixed data for training). When we used 35% of the real data, the performance (MAP is 0.586 and ARI is 0.875) of the case of mixed training (Real35% + \({\mathrm{Synthetic}}_{100{\mathrm{\% }}}^{35{\mathrm{\% }}}\)) achieved competitive performance to the case of using the whole real data (Real100%) in both metrics. This finding proves that our method can significantly reduce the amount of real data in image segmentation (around 65% in this case), which means reducing the pressure of obtaining and labeling real images from costly experiment. In order to demonstrate the statistical significance of resulted values, we have conducted extra tests to estimate the mean and standard deviation of the performance of our best model (see Supplementary Note 4 with Supplementary Table 2).
The visualization of image segmentation is shown in Fig. 4. From left to right are real image, manually annotated label, the results of model training with Real35% and the results of model training with mixed data (\({\mathrm{Real}}_{35{\mathrm{\% }}} + {\mathrm{Synthetic}}_{100{\mathrm{\% }}}^{35{\mathrm{\% }}}\)). The orange rectangles and their magnified results (shown in upper left of the figure) denote the areas in which the model trained with small real data cannot close the grain boundary, but the model trained with data augmentation can segment those areas correctly. The experimental results demonstrate that the proposed data augmentation method improves the performance in terms of quantitative assessment and result visualization.
In addition, we compare the performance of our data augmentation strategy with traditional augmentation methods (such as rotation and flipping) and pre-training method (Ternausnet42) in Supplementary Notes 5 and 6 with Supplementary Tables 3, 4, respectively. We confirmed that our proposed augmentation method provides extra improvement to the segmentation task on top of the traditional data augmentation methods. Furthermore, the typical pre-training method (e.g., Ternausnet) did not achieve the same level of improvement as our proposed method did.
Discussion
Modern machine learning methods, such as deep learning, have outstanding prediction performance when the model is trained with a sufficiently large amount of data. However, for most applications in materials science, there is a chronic lack of experimental data for any given task, i.e., the small data dilemma. In the present work, we developed a novel transfer learning strategy to address problems of small or insufficient data in material data mining. This strategy realizes the fusion of experimental data (real data) and simulated/calculated data based on transfer learning theory and the augmentation of training data in a data mining procedure, so that classification or regression models with better performance can be established based on only a small set of real data. Then, we applied this strategy to the task of grain instance segmentation. First, we transferred the “image style” information of a real image to an image of simulated label to generate a synthetic image. Such a transfer learning technique extracted important features from the real image to modify the images from simulation, making them more relevant to the segmentation task of real images43. Second, by supplementing a machine learning model with synthetic dataset (synthetic image and simulated label), the model captured useful features from the synthetic dataset that are transferable to the real dataset and achieved a promising performance.
Computational simulation is a good way to acquire data efficiently. We assumed that the Monte Carlo Potts model could generate simulated data that mimic the pattern of grain growth under ideal conditions. In materials science, simulated data partially capture the actual physical mechanism. However, the Monte Carlo Potts model could not simulate the perturbation of data in practical experiments, mainly coming from the process of sample preparation in the experiment. As shown in Table 3, when there is a large difference between the simulated data and the real data, it is not enough to simply mix the simulated dataset into the real dataset, which may sometimes lead to a negative effect.
Recently, generative adversarial networks (GAN), a deep learning based generative model has been proposed to learn the data distribution and achieve promising performance in image generation. It can learn a loss that tries to classify whether the output image is real or fake while simultaneously training a generative model to minimize this loss44. We chose it as the primary architecture behind our image style transfer model. We exploit the generative power of GAN to capture the small perturbation of real data that commonly occurs in actual experiments. The image style transfer model, then, generates synthetic images by mixing in realistic features into the images from computational simulation.
By treating the synthetic dataset as data augmentation for a machine learning model, the model achieves a promising performance when trained on a mix of 35% real dataset and the whole synthetic dataset, which is competitive to the performance of the model trained on the whole real dataset. In addition, as the time required to generate the synthetic dataset is almost negligible when compared with the time required to generate real dataset, our method is able to achieve high-quality segmentation without a high cost of preparing training data. Hence, this strategy can be used to address the problems of small or insufficient data in material data mining.
This paper has demonstrated the viability of the combination of simulated and real experimental data, suggesting that simulated data (after performing image style transfer from the real data) could be useful in data mining or machine learning problems. We believed that the proposed strategy could be easily applied to other data mining tasks, structured data analysis, or even those outside materials science. Our next step will be to investigate a more efficient transfer learning technique for this segmentation task to fully suppress the need to use real experimental dataset for training.
Methods
Experimental
A commercial hot-rolled iron plate with a purity of >99.9 wt% was used in this work. The plate was forged into a round bar with a diameter of 30 mm and then fully recrystallized by annealing at 1153 K for 3 h. The samples for metallographic characterization were spark cut from the bar. One surface of the samples was mechanically polished for a fixed time, and metallographic photographs were then taken with an optical microscope after etching using a 4 vol% nital solution. The steps of the polish-etching photograph above were repeated to obtain serial section photographs. The average thickness interval between the two sections is ~1.8 μm.
Monte Carlo Potts simulated model
The Monte Carlo Potts model19,20 was used to simulate three-dimensional normal grain growth and the processed images were exported as simulated data to train segmentation models. We used a 400 × 400 × 400 cubic lattice with full periodic boundary conditions to represent the continuum microstructure. A positive integer, Si, termed an index number, was assigned to each site in the lattice sequentially. The index number of a site corresponded to the orientation of the grain that it belonged to. Sites with the same index were considered part of the same grain, and the grain boundary existed only between neighbors with different orientations. The Potts model served as the grain boundary energy function:
where E is the boundary energy, N is the system size (the total sites in the system), J is the positive constant that scales the boundary energy, NN is the number of nearest neighbors of site i, and δ is the Kronecker function with \(\delta _{S_iS_j} = 1\) if Si = Sj and 0 otherwise. Here, NN = 26, which means that the interactions between a given site and its 6 first-nearest neighbor sites, 12 second-nearest neighbor sites, and 8 third-nearest neighbor sites were considered in the calculation of grain boundary energy. The simulation of grain growth involved a reorientation attempt for each site. Thus, the net energy change associated with the reorientation trial can be expressed by:
where Ei1 and Ei2 are the boundary energies of site i before and after the reorientation trial. The possibility of this reorientation is given by:
where kB is the Boltzmann constant and T is the Monte Carlo temperature. The term kBT defines the thermal fluctuations within the simulated system. Time is scaled by the Monte Carlo step (MCS), which corresponds to as many orientation trials as there are lattice sites.
The probability of the trial reorientation for the target is determined by Eq. (3) with the constant \(\left( {\frac{{k_{\mathrm{B}}T}}{J}} \right) = 0.5\), which is large enough to reduce lattice pinning but small enough to avoid lattice break-up.
Figure 1c shows the Monte Carlo simulation structure. Figure 5 shows the distribution of grain area and the distribution of the number of grain edge. The pure iron structure and Monte Carlo simulation structure have features that are similar in terms of grain morphology, grain area distribution, and edge number distribution.
Image style transfer
We used the image style transfer model to generate simulated data by extracting the “image style” information from real images. Specifically, the conditional GAN (pix2pix)23,45 was adopted to carry out the image-to-image transformation. A GAN consists of two modules: a generator G and a discriminator D. Assume that x is the source image and y is the target image such that (x, y) is a pair of data, for example, two styles of an image. The generator G produces an output image G(x) given the original image x before style transfer as a condition, while the discriminator D distinguishes G(x) from the corresponding y. Both modules are optimized by adversarial training to make the “fake” G(x) closer to the “true” y.
The objective function of a conditional GAN can be expressed as:
where G tries to minimize this objective and D tries to maximize it.
A previous study21 indicated that the generator has to not only fool the discriminator but also be similar to the ground truth output in an L1 sense. The L1 loss is described as follows:
The final objective is as follows:
We performed image-to-image transfer to convert image of a simulated label to a synthetic image, as shown in Fig. 6. Our generator used the encoding–decoding network U-net28. The network structure bridges the input and output through a bottleneck layer, which helps to retain the underlying structural similarity in a lower dimensional space. The discriminator calculates the loss of local patches between the output and ground truth to represent the consistency of local details46. In the training stage, the generator is given an image of manually annotated label as input and is trained to produce a synthetic image, similar to the real image. The discriminator, on the other hand, is trained to distinguish the “real ones” versus the generated “synthetic ones”. During the training, the generator acquires the ability to transfer the input labels to synthetic images. Then, we use this generative model to perform inference on the images of simulated label, i.e., converting all images of simulated labels into synthetic images.
Data availability
The datasets and codes are available from the corresponding author on reasonable request. The third-party image style transfer codes that we used are available online45.
Change history
11 February 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41524-020-00415-2
References
Lookman, T., Alexander, F. J. & Rajan, K. Information Science for Materials Discovery and Design. (Springer Cham. Press, Switzerland, 2016).
Butler, K. T., Davies, D. W., Cartwright, H., Isayev, O. & Walsh, A. Machine learning for molecular and materials science. Nature 559, 547–555 (2018).
Raccuglia, P. et al. Machine-learning-assisted materials discovery using failed experiments. Nature 533, 73–76 (2016).
Luna, P. D., Wei, J., Bengio, Y., Aspuru-Guzik, A. & Sargent, E. Use machine learning to find energy materials. Nature 552, 23–27 (2017).
Sanchez-Lengeling, B. & Aspuru-Guzik, A. Inverse molecular design using machine learning: Generative models for matter engineering. Science 361, 360–365 (2018).
Wu, S. et al. Machine-learning-assisted discovery of polymers with high thermal conductivity using a molecular design algorithm. npj Comput. Mater. 5, 66 (2019).
Wang, C. S., Fu, H. D., Jiang, L., Xue, D. Z. & Xie, J. X. A property-oriented design strategy for high performance copper alloys via machine learning. npj Comput. Mater. 5, 87 (2019).
Wu, Y. J., Fang, L. & Xu, Y. B. Predicting interfacial thermal resistance by machine learning. npj Comput. Mater. 5, 56 (2019).
Dursun, T. & Soutis, C. Recent developments in advanced aircraft aluminium alloys. Mater. Des. 56, 862–871 (2014).
Hu, J., Shi, Y. N., Sauvage, X., Sha, G. & Lu, K. Grain boundary stability governs hardening and softening in extremely fine nanograined metals. Science 355, 1292–1296 (2017).
Sonka, M., Hlavac, V. & Boyle, R. Image Processing, Analysis, and Machine Vision. 4th edn (Cengage Learning. Press, Boston, 2014).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. Imagenet classification with deep convolutional neural networks. NeurIPS. 1097–1105 (2012).
Long, J., Shelhamer, E. & Darrell, T. Fully convolutional networks for semantic segmentation. CVPR. 3431–3440 (2015).
Lewis, A. C. & Howe, D. Future directions in 3D materials science: outlook from the first international conference on 3D materials science. JOM 66, 670–673 (2014).
Li, W., Field, K. G. & Morgan, D. Automated defect analysis in electron microscopic images. npj Comput Mater. 4, 35 (2018).
Azimi, S. M., Britz, D., Engstler, M., Fritz, M. & Mucklich, F. Advanced steel microstructural classification by deep learning methods. Sci. Rep. 8, 2128 (2018).
Ma, B. Y. et al. Deep learning-based image segmentation for al-la alloy microscopic images. Symmetry 10, 107 (2018).
Richter, S. R., Vineet, V., Roth, S. & Koltun, V. Playing for data: ground truth from computer games. ECCV. 102–118 (2016).
Anderson, M. P., Grest, G. S. & Srolovitz, D. J. Computer simulation of normal grain growth in three dimensions. Philos. Mag. B 59, 293–329 (1989).
Radhakrishnan, B. & Zacharia, T. Simulation of curvature-driven grain growth by using a modified Monte Carlo algorithm. Metall. Mater. Trans. A. 26, 167–180 (1995).
Gatys, L. A., Ecker, A. S. & Bethge, M. Image style transfer using convolutional neural networks. CVPR. 2414–2423 (2016).
Gatys, L. A., Ecker, A. S. & Bethge, M. A neural algorithm of artistic style. Preprint at https://arxiv.org/abs/1508.06576 (2015).
Isola, P., Zhu, J. Y., Zhou, T. & Efros, A. A. Image-to-image translation with conditional adversarial networks. CVPR. 1125–1134 (2017).
Zhu, J. Y., Park, T., Isola, P. & Efros, A. A. Unpaired image-to-image translation using cycle-consistent adversarial networks. ICCV. 2223–2232 (2017).
Shrivastava, A. et al. Learning from simulated and unsupervised images through adversarial training. CVPR. 2107–2116 (2017).
Choi, Y. et al. Stargan: unified generative adversarial networks for multi-domain image-to-image translation. CVPR. 8789–8797 (2018).
Zhu, J. Y. et al. Toward multimodal image-to-image translation. NeurIPS. 465–476 (2017).
Ronneberger, O., Fischer, P. & Brox, T. U-net: convolutional networks for biomedical image segmentation. MICCAI. 234–241 (2015).
Falk, T. et al. U-Net: deep learning for cell counting, detection, and morphometry. Nat. Methods 16, 67–70 (2019).
Kaggle data science bowl 2018. https://www.kaggle.com/c/data-science-bowl-2018/overview/evaluation (2019).
Lin, T. Y. et al. Microsoft COCO: common objects in context. ECCV. 740–755 (2014).
Rand, W. M. Objective criteria for the evaluation of clustering methods. J. Am. Stat. Assoc. 66, 846–850 (1971).
Hubert, L. & Arabie, P. Comparing partitions. J. Classif. 2, 193–218 (1985).
Vinh, N. X., Epps, J. & Bailey, J. Information theoretic measures for clusterings comparison: Variants, properties, normalization and correction for chance. JMLR 11, 2837–2854 (2010).
Python language reference. http://www.python.org (2019).
Pytorch. https://pytorch.org/ (2019).
Laganière, R. OpenCV 3 Computer Vision Application Programming Cookbook. 3rd edn (Packt Publishing Ltd. Press, Birmingham, 2017).
Otsu, N. A threshold selection method from gray-level histograms. IEEE T Syst. Man CY-S 9, 62–66 (1979).
Canny, J. A computational approach to edge detection. TPAMI 6, 679–698 (1986).
Roerdink, J. B. & Meijster, A. The watershed transform: definitions, algorithms and parallelization strategies. Fund Inform. 41, 187–228 (2000).
Jain, A. K. Data clustering: 50 years beyond K-means. ICPR 31, 651–666 (2010).
Iglovikov, V. & Shvets, A. Ternausnet: U-net with vgg11 encoder pre-trained on imagenet for image segmentation. Preprint at https://arxiv.org/abs/1801.05746 (2018).
Yamada, H. et al. Predicting materials properties with little data using shotgun transfer learning. ACS Central Sci. 5, 1717–1730 (2019).
Goodfellow, I. J. et al. Generative adversarial. Netw. NeurIPS 3, 2672–2680 (2014).
Pix2Pix in Pytorch. https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix (2019).
Li, C. & Wand, M. Precomputed real-time texture synthesis with markovian generative adversarial networks. ECCV. 702–716 (2016).
Acknowledgements
The authors acknowledge financial support from the National Key Research and Development Program of China (No. 2016YFB0700500), the National Science Foundation of China (No. 51574027, No. 61572075, No. 6170203, No. 61873299), the Finance science and technology project of Hainan province (No. ZDYF2019009), and the Fundamental Research Funds for the University of Science and Technology Beijing (No. FRF-BD-19-012A, No. FRF-TP-19-043A2). The computing work is partly supported by USTB MatCom of Beijing Advanced Innovation Center for Materials Genome Engineering. And this manuscript was edited for English language by Springer Nature. This work was supported by Beijing Top Discipline for Artificial Intelligent Science and Engineering, University of Science and Technology Beijing.
Author information
Authors and Affiliations
Contributions
B.M. conceived the idea, designed the experiment and wrote the paper; X.W. participated in the paper writing and conducted experiments with C.L.; X.B. organized the research project; H.H. participated in the experimental design and discussion with S.W.; H.W. and W.X. prepared the data for the experiments. M.G. prepared the computational environment of the experiment. Y.S., Q.S., M.M., A.O.A., and H.S. were involved in the analyses of data. Besides, Y.S. provided financial support.
Corresponding authors
Ethics declarations
Competing interests
B.M., X.W., X.B., H.H., H.W., and W.X. declare the following competing interests: one patent has been registered (201910243002.6).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ma, B., Wei, X., Liu, C. et al. Data augmentation in microscopic images for material data mining. npj Comput Mater 6, 125 (2020). https://doi.org/10.1038/s41524-020-00392-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-020-00392-6
This article is cited by
-
Advances in machine learning- and artificial intelligence-assisted material design of steels
International Journal of Minerals, Metallurgy and Materials (2023)
-
Prediction of mechanical properties for deep drawing steel by deep learning
International Journal of Minerals, Metallurgy and Materials (2023)
-
A reusable neural network pipeline for unidirectional fiber segmentation
Scientific Data (2022)
-
Visualization of judgment regions in convolutional neural networks for X-ray diffraction and scattering images of aliphatic polyesters
Polymer Journal (2021)