Abstract
Cutting edge deep learning techniques allow for image segmentation with great speed and accuracy. However, application to problems in materials science is often difficult since these complex models may have difficultly learning meaningful image features that would enable extension to new datasets. In situ electron microscopy provides a clear platform for utilizing automated image analysis. In this work, we consider the case of studying coarsening dynamics in supported nanoparticles, which is important for understanding, for example, the degradation of industrial catalysts. By systematically studying dataset preparation, neural network architecture, and accuracy evaluation, we describe important considerations in applying deep learning to physical applications, where generalizable and convincing models are required. With a focus on unique challenges that arise in high-resolution images, we propose methods for optimizing performance of image segmentation using convolutional neural networks, critically examining the application of complex deep learning models in favor of motivating intentional process design.
Similar content being viewed by others
Introduction
In situ and operando experimental techniques, where dynamic process can be observed with high temporal and spatial resolution, have allowed scientists to observe chemical reactions, interfacial phenomena, and mass transport processes to give not only a better understanding of the physics of materials phenomena, but also a view into how materials react under the conditions in which they are designed to perform1,2. As the use of in situ techniques continues to expand, and technology to enable these experiments continues to develop, we are faced with the fact that more data can be produced than can be feasibly analyzed by traditional methods3,4. This is particularly true for in situ electron microscopy experiments, where high-resolution images are captured at very high frame rates. In practice, hundreds of images can be captured per second. However, many experimental analyses consider less than one frame per second, or even one frame for every several minutes5. Methods for fast and efficient processing of high-resolution imaging data will allow for not only full utilization of existing and developing technologies, but also for producing results with more statistical insight based on the sheer volume of data being analyzed.
Simultaneously, the field of computer vision provides well understood tools for image processing, edge detection, and blob localization that are helpful for moving from raw image data to quantifiable material properties. These techniques are easy to apply in many common computer programming languages and libraries. However, more recent research highlights the processing speed and accuracy of results obtained through the use of machine learning6,7. Previously, a combination of traditional image processing and advanced statistical analysis has be shown to successfully segment medical images8,9. Deep learning—generally using multilayer neural network models—expands on other machine learning techniques by using complex connections between learned parameters, and the addition of nonlinear activation functions, to achieve the ability to approximate nearly any type of function10. With regards to image segmentation and classification, the use of convolutional neural networks (CNNs), in which high-dimensional learned kernels are applied across grouped image pixels, is widespread. CNNs provide the benefit that their learned features are translationally equivariant, meaning that image features can be recognized regardless of their position in the image. This makes such models useful for processing images with multiple similar features, and robust against variation in position or imaging conditions11. Additionally, the feature richness of high-dimensional convolutional filters and the large number of connections between hidden layers in a neural network allows for the learning of features that, conventionally, are too complex to represent, and that make intuitive interpretation difficult. Much of the literature studying CNNs focuses on high-accuracy segmentation/classification of large, complex, multiclass image datasets or upon improving data quality through super-resolution inference, rather than quantitative analysis of high-resolution images12. Though conventionally used to specify atomic-resolution imaging in the field of electron microscopy, in this work we use the term high-resolution to refer to the pixel resolution of the microscope camera. While additional memory requirements alone make processing of high-resolution images difficult, the scale of features and possible level of precision also changes as a function of image resolution. Most importantly, for the simple case of particle edge detection, the boundary between classes in a high-resolution image may spread across several pixels, making segmentation difficult even by hand. Generally, literature studies of CNNs for image classification are used for many-class classification with coarse—if any—object localization, while in the field of electron microscopy fewer individual object classes exist in a single image, yet precise positioning is required.
Though, it seems, the tools for rapid segmentation of high-resolution imaging data exist, several points of concern regarding the use of deep learning must be acknowledged. First, while the ease of implementation using common programming tools enables extension of methods to new applications by nonexperts, the complexity and still-developing fundamental understanding of deep learning can lead to misinterpretation of results and poor reproducibility13,14. Moreover, models can be prone to overfitting—memorizing the data rather than learning important features from limited training examples—which can go unnoticed without careful error analysis15,16. Overfitting occurs when a model has enough parameters that an unrealistically complex function can be fit to match every point in a dataset. Thus, a model which accurately labels data by overfitting will likely fail when shown new data, since its complex function does not describe the true variation in the data. Therefore, an overfitted model isn’t useful for future work. Finally, the high dimensionality of data at intermediate layers of a neural network combined with the compound connections between hidden layers makes representation, and therefore understanding, of learned features impossible without including more assumptions into the analysis. These challenges—specifically representation and visualization of CNN models—are areas of active research17,18.
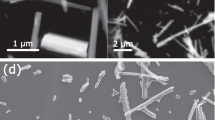
We focus on semantic segmentation of environmental transmission electron microscopy (ETEM) images of supported gold nanoparticles19,20,21,22. Ensembles of supported nanoparticles are important for industrial catalysis, deriving their exceptional catalytic activity from surface energy resulting from the high amount of under-coordinated surface atoms relative to the particle’s bulk volume. On a thermodynamic basis, the high surface energy that allows for effective catalysis also provides a driving force for nanoparticle sintering through a variety of mechanisms23,24. Theory exists to describe the mean-field process of Ostwald ripening and basics of nanoparticle coalescence, yet local effects and interparticle interactions cause deviations from our theoretical understanding. Obtaining precise sizes and locations of nanoparticles as a function of space and time is imperative to describing nanostructural evolution, and developing a physical understanding of the processes leading to catalyst degradation by particle growth. Thus, our high-contrast images of supported gold nanoparticles provide a simple, yet important, case study for developing efficient methods of image segmentation so that individual particle-scale changes can be studied.
Building on previous work on image segmentation, automated analysis, and merging deep learning within the field of materials science, we study a variety of CNN architectures to define the most important aspects for the practical application of deep learning to our task. We discuss how image resolution affects segmentation accuracy, and the role of regularization and preprocessing in controlling model variance. Further, we investigate how image features are learned, so that model architectures can be better designed depending on the task at hand. By using a simpler approach to semantic segmentation, in contrast to poorly understood and highly complex techniques, we intend to show that conventional tools can be utilized to construct models that are both accurate and extensible.
Results
High-resolution image segmentation
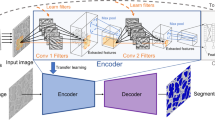
Particularly in the field of medical imaging, studies regarding similar image segmentation tasks have been published25,26. In these cases, an encoder–decoder, or ‘hourglass’-type CNN architecture was found to be well suited to segmentation tasks, where spatial positions of features are key. With this approach, successively deeper convolutional/max-pooling layer pairs (added to decrease spatial resolution while simultaneously increasing feature richness) are combined with up-sampling convolutional layers that aim to rescale the image back to a higher resolution, while decreasing the feature dimension of the image source21,27,28. In many cases, however, these tasks are used to identify whether a specific feature or object is present or absent, not to measure the size of such features with any level of precision. Correspondingly, our tests show that this network structure successfully identifies nanoparticle pixels in our images with 512 × 512 resolution, yet consistently misses the centers of the largest particles (Supplementary Fig. 1).
To improve the segmentation performance, we moved to a more complex architecture inspired by the UNet29. This model, rather than increasing kernel size with the goal of expanding the receptive field, uses skip connections to tie activations in the encoding stage to feature maps in the decoding stage, in order to improve feature localization. Skip connections work by concatenating encoded and decoded images of the same resolution followed by a single convolutional layer and activation function to relate unique aspects of both images (see visual representation in Supplementary Fig. 2). This improves upon the similar hourglass architecture by maintaining local environments from the original image to map features to the output. Results using the UNet-type architecture on our image set show that the model is able to consistently recognize both large and small particles, and that it is robust against varied imaging conditions and datasets (Fig. 1 shows results on images from experiments not represented in the training set).
Segmentation results for UNet-type architecture on 512 × 512 resolution images. a Raw output from the model overlaid on the raw image; notice the sharp activation cutoff at the particle edges. b Threshold applied to image to show final segmentation result. Yellow arrows indicate small particles that were successfully recognized. Scale bar represents 50 nm.
Using our earlier approach, we trained the same UNet on higher resolution images (1024 × 1024 pixels), however, as seen in Fig. 2, this network was not able to accurately label pixels at nanoparticle edges, showing instead a blur of uncertainty at the edges. Moreover, we noticed that training the same model on the same data more than once would produce different results: while in some cases training produced image segmentation with wide edge variation, other training instances gave segmentation results with nearly perfectly identified particles, with little to no variation at particle edges. These results likely signal overfitting of the dataset, with the model “memorizing” the noise rather than actual features, as raw activation maps (Fig. 3) show that in fact no features of particles are learned by the model and instead only noise patterns in the background areas are recognized. This model, therefore, produces a very accurate particle measurement on the training dataset, but would not generalize to data from other experiments or with particles of different sizes (i.e., the same dataset with a different magnification). This is further highlighted by the instability of the model with respect to the length of training time.
Rather than solely increasing the width and depth of the model to improve performance and stability (we used a four-step UNet-type architecture for 1024 × 1024 images, as depicted in Supplementary Fig. 2), the greatest improvement in model performance comes through understanding where the model fails when increasing image resolution. Fifteen unique UNet models were tested with architectural modifications inspired by the errors observed in our tests. These modifications, and the motivation for each, are described in Table 1. The effect of learning rate on model performance was also investigated empirically in order to determine how to best sample the loss landscape, but in this regard, we found that a learning rate of 0.0001 is practical and effective for all deep models on our dataset.
Results from all fifteen models are shown in Supplementary Fig. 3. Our initial gauge of performance is qualitatively based on the ability to detect particles of varying size, sensitivity to noise and illumination variation in the raw image, and the sharpness of the activation cutoff at particle edges. Based on these criteria, best performance is seen in models with batch normalization only and batch normalization combined with extra convolutional layers (Fig. 4, Norm and TwoConv_Norm, respectively). From this, it appears that batch normalization is the most important factor for learning particle features from 1024 × 1024 images. Visual inspection of Fig. 4 also shows that, in general, blurred images detect edges further toward the interior of the nanoparticle, and models with an additional convolutional layer (and no blurring) are virtually indistinguishable from those with a single up-sampling convolution. More importantly, only models without blur are able to consistently and accurately label small, low-contrast particles.
Aside from applying batch normalization, we find that the only way to achieve significant segmentation improvement on high-resolution images is to increase the size of the convolutional kernel, here from 3 × 3 pixels to 7 × 7 (Supplementary Fig. 4). However, this greatly increases the number of trainable parameters and training time for the model.
To briefly summarize the practical implications of our findings, continual batch normalization through successive convolutional layers has a significant positive effect on the performance. For our dataset, increasing network depth does not appear to increase the performance of the CNN. A slow learning rate produces the best results and most stable models, while preprocessing training images with Gaussian blur seems to increase the risk of overfitting.
Evaluating detection accuracy
Variation in the color scale at particle edges, as seen Fig. 2, led us to believe that our particle measurement would vary greatly as a function of the chosen softmax-activation threshold. Intensity line profiles, as shown in Fig. 5, are helpful in illustrating this edge variation for two models compared to the intensity of the raw image. These plots check how two different models perform in comparison with the edge contrast in the raw image. As the intensity approaches 1, both models show a slope toward the particle center showing the extent of uncertainty in classification at the particle-support interface. Supplemental Figure 5 collects precision and recall scores for the batch-normalized CNN as a function of threshold value, and the amount of Gaussian blur applied compared to a set of 50 validation set labels. Here, high precision means that the model produces few false positives (pixels labeled as particle that actually correspond to background), while recall measures the proportion of particle pixels that were successfully identified by the model (see individual plots in Supplemental Fig. 4). Based on these results, we could expect that the normalized models with no applied blur, and blur (σ = 1) are stable with respect to precision and recall at a particle activation threshold values <0.7. The model trained on blurred images with σ = 2, shows similar performance over a smaller range of stable thresholds. For our case of binary classification of an unbalanced dataset, where recognizing particles pixels is more important than recognizing background, recall is likely the most important measure for determining a threshold for use in practice. While we see convergence with maximum precision for the model without blur around a threshold of 0.7, we realize that our empirically selected value of 0.4 gives better recall with essentially the same precision as compared to thresholding at 0.7.
Learning features with a simpler model
Training stability and model overfitting pose large risk for image segmentation CNNs that are to be used and continually developed on varied datasets. While performance often increases with the addition of tunable model parameters, achieving training convergence and interpretation of the model’s output become increasingly difficult. With this in mind, we developed a significantly pared down CNN, with a single convolutional layer consisting of a single learnable filter followed by softmax activation on our training data that produced the segmentation shown in Fig. 6b. The benefit of such an architecture is that, since the dimensionality of the kernel is the same as that of the image, we can easily visualize the learned weights (Fig. 6a). Previous work confirms that edges and other spatially evident image features are generally learned in the early convolutional layers of a CNN30. Repeating the same method with another kernel size, this time 7 × 7 pixels rather than the initial 9 × 9, produces a similar filter, showing that the results are not an artifact of the feature scale. Such a single-layer model with logistic activation can be compared, in practice, to a sparse convolutional autoencoder, or even the application of a linear support vector machine for logistic regression31.
While this model is useful for illustrating the power of simpler machine learning methods, minimal changes are needed to extend this idea to a model that provides usable, practical segmentation. Using one convolutional layer, now with 32 filters, followed by a second, 1 × 1 convolutional layer to combine the features into a segmented image, we test a shallow but wide CNN architecture. Again, aside from the convolutional layer used to combine the extracted features, filters from this shallow network can be visualized to see what features are being learned from the data. The F1 score of this simpler model (Fig. 7a, blue line) is comparable to the performance of the most accurate deep network described above (batch normalization with no applied blur—red line). These results illustrate that a model with significantly fewer parameters and quicker training time can still produce a usable segmentation. Indeed, as shown in Fig. 7b, the edges detected by the simpler CNN are in many cases closer to the actual particle edge than those of the deep model; in this light, the decrease F1 score in Fig. 7a is likely due to the high rate of false positives in the simple model. In practice any false positive clusters are significantly smaller than true nanoparticles, so filtering by size to further increase accuracy is possible. Our results suggest that shallow, wide CNNs have enough expressive power to segment high-resolution image data32.
a Mean F1 score for UNet (only modified by adding batch normalization) and simple one-layer CNN architectures as a function of Softmax threshold cutoff. Red and blue curves, and image contours represent results from the UNet and simplified architecture, respectively. Error bands in a represent the range within a standard deviation of the mean F1 across the validation set. b Visual comparison of nanoparticle detection, using the Otsu threshold, for the simplified model (blue) and the best performing model (red).
Discussion
Our initial experiments revealed the importance of a segmentation model developing an understanding of a pixel’s broader environment, rather than simply identifying features based on intensity or distance to an edge. The fact that the simple, hourglass-style CNNs cannot identify the interior of particle as such, can be attributed to an inability of the CNN to learn similar features with different size scales; we suspect that, in an edge-detecting model, the lack of variation in the interior of a particle appears similar to the in the background leading to improper classification. This clearly indicates the importance of semantic understanding, in which the local environment is considered in detail. Indeed, increasing the receptive field (kernel size) of the network to incorporate more local information improves detection accuracy, yet this approach drastically increases the number of learnable parameters in the CNN and the training time required for convergence. This is reinforced in seeing the improved performance of the UNet compared to the hourglass CNN. Max-pooling after each convolutional layer effectively increases the receptive field of the next convolutional layer; concatenating encoding and decoding activations serves as a comparison of the same features over a variety of length scales.
While segmentation of 512 × 512 pixel images is possible and seemingly accurate, higher measurement precision can be achieved by utilizing higher resolution cameras available on most modern electron microscopes. For an image with a fixed side length, increasing pixel resolution decreases the relative size of each pixel. Decreasing the pixel size increases the possible measurement precision, and therefore, high-resolution images are needed to provide both accurate, and consistent particle measurements. Along these lines, the error introduced by mislabeling a single pixel decreases as pixel density (image resolution) increases. It’s important to note that though the accuracy of manual particle measurements from images with different resolutions likely changes very little (assuming accuracy is mainly dependent on the care taken by the person making measurements), changes in resolution, particularly around particle edges, can greatly influence automated labeling performance since edge contrast decreases as interfaces are spread across multiple pixels. Thus, a unique challenge for high-resolution image segmentation is developing a model that is able to recognize interface pixels, which appear fundamentally different from the interior of a nanoparticle, as contributing to the particle and not the background. To account for increased complexity of the features in higher resolution images, we expanded our network architecture both in depth and width with the idea that a larger number of parameters would increase the expressive power of the model. In fact, this deeper and wider model (seen in Fig. 2) showed little increase in performance compared to the one for low-resolution images. A more effective approach would match the strengths of the segmentation models to the features of the data. For our case of relatively simple images, increasing the complexity of the model alone does not achieve this goal.
Our findings show that regularization, in this case by batch normalization, is vital to accurate labeling of an image. When training from scratch, i.e., without pretrained weights, it has been shown that the loss function is smoother and model convergence is better when using batch normalization, which may have a significant effect on higher resolution images due to the combinations of strong noise and lack of visually discriminative features on the scale of the receptive field33. Properly pairing regularization, in attempt to maintain the distribution of intensity values in the image, with an activation function suited to allowing such a distribution is essential. As such, the dying rectified linear unit (ReLU) problem, where CNN outputs with a negative value are pushed to zero, removing a significant portion of the actual distribution of the data, causes loss of information and difficult convergence10,34. Our use of ReLU activation functions essentially produces output values in the range (0,∞), which presents a risk of activation divergence, and can be mitigated by normalization in successive convolutional layers before the final softmax activation. Leaky ReLU allows activations on the range (-∞,∞), and the small activation for negative pixel values combined with batch normalization works to avoid increasing variance with the number of convolutional layers. In practice, we find that using Leak ReLU activation solves the problem seen in Fig. 3, where no activation is seen for the particle class.
These results suggest that, for a common segmentation task, regularization is more effective than the depth or complexity of a CNN. This is easily justified, considering that the proper classification of boundary pixels, spread across several pixels in high-resolution images, requires the sematic information stored in the total local intensity distribution that is lost as the variance of the intensity histogram increases.
As shown in Fig. 7, the choice of an activation threshold for identifying nanoparticles can greatly influence the labeling error. The steep slope of the softmax-activation function used in the final CNN layer works to force activation values toward 0 or 1—in an ideal case the number of pixels with activation values between these values would be minimal. Our experience shows that the Otsu threshold, which separates the intensity histogram such that the intraclass variance is minimized, is a practical choice for segmenting our data35. This makes sense, since, qualitatively, CNN output shows a large peak close to 0 activation representing the background with nearly all pixels with higher activation values corresponding to particles. However, it can be shown mathematically that the calculated Otsu threshold may mislabel the class with a wider intensity distribution36. Therefore, thresholding datasets with a lower signal-to-noise ratio would likely be more difficult. In these cases, it is imperative that a large dataset—which is representative of the data in question—is used for training, as choosing low threshold values, even when they produce usable results, makes it difficult to recognize overfitting.
An effective machine learning model requires a balance between the number of learnable parameters, the complexity of a model, and the amount of training data available in order to prevent overfitting and ensure deep learning efficiency32,37. In an efficient model, a vast majority of the weights are used, and vital to the output. In practice though, deep networks generally have some amount of redundant or trivial weights38. In addition to efficiency, several issues have come to light regarding the use of deep learning for physical tasks that require an interpretable and explainable model, as this often leads to better reproducibility and results that generalize well18,37. Even for computer vision tasks, where feature recognition doesn’t necessarily give physical insight, an interpretable model is valuable so that sources of error can be understood when applied to datasets consisting of thousands of images, each of which cannot feasibly be checked for accuracy. Our main goal in employing a single-layer neural network was to provide a method for visualizing learned kernels that show the most important features of an image for binary classification. The visualization of our trained kernel (Fig. 6a) can be interpreted in two ways. First, we can conceive that the algorithm is learning vertical and horizontal lines (dark lines), potentially similar to basic Gabor filters for edge detection—though it is missing the characteristic oscillatory component—combined with some amount of radially symmetric blur (light gray). Alternatively, we can envision that the horizontal/vertical lines could be an artifact of the electron camera or data augmentation method meaning that the learned filter represents an intensity spread similar to a Laplacian of Gaussian (LoG) filter, which is used to detect blobs by highlighting image intensity contours. As a simple test of our supposition, Supplementary Fig. 6 shows that a sum of a horizonal Gabor filter, vertical Gabor filter, and Gaussian filter qualitatively produces a pattern similar to our learned kernel.
As mentioned, increasing the width of a shallow network (in this case from 1 to 32 filters) is enough to make a simple model more usable. Though 32 filters (visualized in Supplementary Fig. 7) may be too many filters to easily compare for visually extracting useful information, it is possible to see a general trend: filters are learning faint curved edges. Moreover, taking the mean of all 32 filters (Supplementary Fig. 8) shows a similar pattern as Fig. 6a with slight rotation. Further analysis of the set of 32 filters would require regularization of the entire set of weights to allow for more direct comparison; however, it is possible to imagine a case where, with a properly tuned receptive field in the convolutional layer, more subtle image features than hard lines could be revealed through visualizing a learned kernel. Based on these results, we expect that designing a shallower neural network that retains the local semantics learned in an encoder–decoder or UNet architecture would make a generalizable model for particle segmentation more realistic.
In summary, we have systematically tested several design aspects of CNNs with the goal of evaluating deep learning as tool for segmentation high-resolution ETEM images. With proper dataset preparation and continual regularization, standard CNN architectures can easily be adapted to our application. While overfitting, class imbalance, and data availability are overarching challenges for the use of machine learning in materials science, we find that knowledge of data features and hypothesis-focused model design can still produce accurate and precise results. Moreover, we demonstrate that meaningful features can be learned in a single convolutional layer, allowing us to move closer to a balance between state-of-the-art deep learning methods and physically interpretable results. We evaluate the accuracy of several deep and shallow CNN models and find evidence that, for a relatively simple segmentation task, important image features are learned in the initial convolutional layers. While we apply common accuracy measures to evaluate our models, we note that other specially designed metrics may help to define exactly where mistakes are made, and thereby which features a model is unable to represent. Whether or not these simplified models reach the accuracy required for quantification of segmented images, a learned indication of important low-level image features can help guide the design of an efficient, parallelizable pipeline for conventional image processing.
We present a method for simultaneously segmenting images and visualizing the features most important for a low-level description of the system. While we don’t derive any physical insight from the learned features of our images, this approach could potentially be extended, for example, to a multiclass classification task where learned kernels could elucidate subtle pixel-scale differences between feature classes. For our needs, the interpretability of this basic model helps us to design a segmentation process where measurement accuracy is limited by the resolution of our instrumentation, not by our ability to identify and localize features. Simple segmentation tasks may not fully utilize a deep CNN’s ability to recognize very rich, inconspicuous features, but the breadth of literature and open-source tools from the computer science community are available for use in other fields and must be applied in order to determine their limitations. In this regard, we hope to provide a clear description of how architectural features can be tweaked for best performance for the specific challenge of segmenting high-resolution ETEM images.
In all, while computer science research trends toward complicated, yet highly accurate deep learning models, we suggest a data-driven approach, in which deep learning is used to motivate and enhance the application of more straightforward data processing techniques, as a means for producing results that can be clearly interpreted, easily quantified, and reproducible on generalized datasets. In practice, the wide availability of technical literature, programming tools, and step-by-step tutorials simultaneously makes machine learning accessible to a wide audience, while obscuring the fact that application to specific datasets requires an understanding of unique, meaningful data features, and of how models can be harnessed to give usable and meaningful analyses. While common in the field of computer vision, in practice many of the techniques we discuss are added to a machine learning model as a black box, with little understanding of their direct effects on model performance. Framing deep learning challenges in the light of real physical systems, we propose means both for thoughtful model design, and for an application of machine learning where the learned features can be visualized and understood by the user. In this way, analysis of data from high-throughput in situ experiments can become feasible.
Methods
Sample preparation
An ~1 nm Au film was deposited by electron beam assisted deposition in Kurt J. Lesker PVD 75 vacuum deposition system to form nanoparticles with an approximate diameter of 5 nm. The film was directly deposited onto DENSsolutions Wildfire series chips with SiN support suitable for in situ TEM heating experiments.
TEM imaging
Samples were imaged in an FEI Titan 80–300 S/TEM ETEM operated at 300 kV. Film evolution was studied in vacuum (TEM column base pressure 2 × 10−7 Torr) at 950 °C. High frame rate image capture utilized a Gatan K2-IS direct electron detector camera at 400 frames per second. Selected images (Fig. 1, Supplementary Fig. 1) were acquired on a JEOL F200 S/TEM operated at 200 kV, with images collected on a Gatan OneView camera.
Automated training set generation
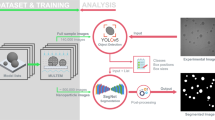
Raw ETEM images are processed using a series of Gaussian filters, Sobel filters, morphological opening and closing, and thresholding algorithms to produce pseudo-labeled training images (see provided code for reproducing specifics). All operations are features of the SciKit Image python package39. As a note, we specify that our dataset is pseudo labeled, because we take automatically labeled images as ground truth, while traditionally labeled data is produced manually by experts in the field. Parameters for each of these processing steps, such as the width of the Gaussian filter, are chosen empirically, and the same parameters are applied to all images in the dataset. Depending on the resolution of the image, and the amount of contrast between the nanoparticles and background in the dataset (which determines the number of required processing steps), automated image processing takes between 10 and 30 s per image. Segmentation by this method is faster than manual labeling for particle measurement and localization, which would take hours per image. Training set accuracy is evaluated by overlaying labels on raw images and visually inspecting the difference, as there is no way to quantitatively check the ground truth. Examples of processing steps and training data are shown in Supplemental Fig. 9.
A set of training data was made up of 2400 full ETEM images (1792 × 1920 pixels), collected during a single experiment, downsized via interpolation to a resolution of 512 × 512 pixels. Additionally, a second training set with 1024 × 1024 pixel resolution, made by cropping appropriately sized sections from a full 1792 × 1920 image, was created to study the impact of increasing pixel resolution on image segmentation performance. In practice, it is important to consider artifacts introduced by resizing images; stretching or compressing images through interpolation/extrapolation may change local signal patterns. Cropping sections of images maintains the scale of features in as-collected images, meaning that a model could potentially be trained on many small images (requiring less GPU memory), and then directly evaluated on full images since convolution neural networks do not require specific input/output sizes once training is complete. Augmentation of the dataset was carried out using affine transformations and image rotation, as successive images captured in a short time are not entirely unique/independent.
Programming and training machine learning models
All programming was done in Python, with machine learning aspects using the PyTorch framework40. The final dataset consisted of 2400 1024 × 1024 pixel images, which was randomly split into training (70%, or 1680 images) and validation (30%, or 720 images) sets. In order to avoid inherent bias due to strong correlation between training and test sets in randomly split consecutive images, a third validation set, collected at a different time but under the same conditions, should be included; we neglect to use this extra dataset, as we only work to show trends in performance as a function of CNN architecture.
In many cases a balanced dataset, where sample sizes of positive and negative examples are roughly equivalent, is required to avoid systematic error and bias while training a CNN. In the images considered here, particle pixels correspond to ~15% of any given image. Though this is quite unbalanced, we find that the general sparsity of features, and the fact that clear edges are the most important factor in identification of nanoparticles in these images, reduce the negative impact of any imbalance.
All CNNs used ReLU activation after each convolutional layer (except where noted later), the Adam optimizer, and cross-entropy loss functions41,42. Since cross-entropy loss in PyTorch includes a final softmax activation, a softmax layer was applied to model outputs for inference. All models were trained for 25 epochs on our System76 Thelio Major workstation using four Nvidia GeForce RTX 2080Ti GPUs, with each model taking 1–2 h to train. We note that longer training periods may be required; we used this time frame to make experimentation with network architecture, data preprocessing, and hyper-parameter tuning more feasible in-house. We gauge that models were stable in this training time by tracking loss as a function of epoch number and seeing general convergence. The binary segmentation map that classifies individual pixels as particle or background was obtained by thresholding predicted softmax output for each pixel.
To obtain quantitative data on the particles themselves, both the training set and CNN segmentation output were processed by a connected components algorithm to produce a labeled image that groups pixels into particle regions from which properties such as size and position can be extracted. This labeling, performed on a binary image, generally takes only one second or less per image
Our base UNet-type architecture for segmenting 512 × 512 images consisted of three convolutional layers with max-pooling or up-sampling (where applicable) on both downscaling and upscaling sides29,43. The base model for 1024 × 1024 images adds an additional level of convolutional layers to each side of the model. Adding convolutional layers, as described later to increase segmentation accuracy, refers to adding a successive convolutional layer after each down-/up-sampling level of a UNet-type architecture. Supplementary Fig. 2 shows a representation of the CNN architecture used here.
Data availability
Contact the corresponding author with requests to view raw data. Sample image sets and all python code used are publicly available in the GitHub repository for this project (link provided below).
Code availability
Python code for training image generation, UNet training, and evaluation of results are available at https://github.com/jhorwath/CNN_for_TEM_Segmentation.
References
Zheng, H., Meng, Y. S. & Zhu, Y. Frontiers of in situ electron microscopy. MRS Bull. 40, 12–18 (2015).
Tao, F. & Salmeron, M. In situ studies of chemistry and structure of materials in reactive environments. Science 331, 171–174 (2011).
Taheri, M. L. et al. Current status and future directions for in situ transmission electron microscopy. Ultramicroscopy 170, 86–95 (2016).
Hill, J. et al. Materials science with large-scale data and informatics: unlocking new opportunities. MRS Bull. 41, 399–409 (2016).
Simonsen, S. B. et al. Direct observations of oxygen-induced platinum nanoparticle ripening studied by in situ TEM. J. Am. Chem. Soc. 132, 7968–7975 (2010).
Badea, M. et al. The use of deep learning in image segmentation, classification, and detection. arxiv 1605.09612 (2016).
Chen, X. W. & Lin, X. Big data deep learning: challenges and perspectives. IEEE Access 2, 514–525 (2014).
Dheeba, J. & Tamil Selvi, S. Classification of malignant and benign microcalcification using SVM classifier. 2011 Int. Conf. Emerg. Trends Electr. Comput. Technol. ICETECT 2011, 686–690 (2011).
Bengio, Y., Lee, D.-H., Bornschein, J., Mesnard, T. & Lin, Z. Towards biologically plausible deep learning. arxiv 1502.04156 (2015).
Goodfellow, I., Bengio, Y. & Courville, A. Deep Learning (MIT Press, 2016).
Moen, E. et al. Deep learning for cellular image analysis. Nat. Methods 16, 1233–1246 (2019).
Yang, W., Zhang, X., Tian, Y., Wang, W. & Xue, J.-H. Deep learning for single image super-resolution: a brief review. IEEE Trans. Multimedia 21, 1–17 (2018).
Zhang, C., Bengio, S., Hardt, M., Recht, B. & Vinyals, O. Understanding deep learning requires rethinking generalization. arxiv 1611.03530 (2017).
Wang, Z. Deep learning for Image segmentation-a short survey. arxiv 1904.08483 (2019).
Dietterich, T. Overftting Overlifting and undercomputing in machine learning. ACM Comput. Surv. 27, 326–327 (1995).
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Selvaraju, R. R. et al. Grad-CAM: visual explanations from deep networks via gradient-based localization. Proc. IEEE Int. Conf. Comput. Vis. 2017, 618–626 (2017).
Umehara, M. et al. Analyzing machine learning models to accelerate generation of fundamental materials insights. npj Comput. Mater 5, 1–9 (2019).
Madsen, J. et al. A deep learning approach to identify local structures in atomic-resolution transmission electron microscopy images. Adv. Theory. Simul. 1, 1–12 (2018).
Schneider, N. M., Park, J. H., Norton, M. M., Ross, F. M. & Bau, H. H. Automated analysis of evolving interfaces during in situ electron microscopy. Adv. Struct. Chem.Imaging 2, 1–11 (2017).
Ziatdinov, M. et al. Deep learning of atomically resolved scanning transmission electron microscopy images: chemical identification and tracking local transformations. ACS Nano 11, 12742–12752 (2017).
Zakharov, D. N. et al. Towards Real time quantitative analysis of supported nanoparticle ensemble evolution investigated by environmental TEM. Microsc. Microanal. 24, 540–541 (2018).
Hansen, T. W., Delariva, A. T., Challa, S. R. & Datye, A. K. Sintering of catalytic nanoparticles: particle migration or ostwald ripening? Acc. Chem. Res. 46, 1720–1730 (2013).
Ostwald, W. Uber die vermeintliche Isomerie des roten und gelben Quecksilberoxyds und die Oberflachenspannung fester Korper. Zeritschrift fur Phys. Chem. 34, 495 (1900).
Shen, D., Wu, G. & Suk, H.-I. Deep learning in medical image analysis. Annu. Rev. Biomed. Eng. 19, 221–248 (2017).
Wilson, R. S. et al. Automated single particle detection and tracking for large microscopy datasets. R. Soc. Open Sci. 3, 160225 (2016).
Badrinarayanan, V., Kendall, A. & Cipolla, R. SegNet: a deep convolutional encoder-decoder architecture for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 39, 2481–2495 (2017).
Long, J., Shelhamer, E. & Darrell, T. Fully convolutional networks for semantic segmentation. IEEE Conf. Comput. Vis. Pattern Recognit. 3431–3440 (2015).
Ronneberger, O., Fischer, P. & Brox, T. U-net: convolutional networks for biomedical image segmentation. Lect. Notes Comput. Sci. 9351, 234–241 (2015).
Yosinski, J., Clune, J., Bengio, Y. & Lipson, H. How transferable are features in deep neural networks?. Adv. Neural. Inf. Process Syst 27, 3320–3328 (2014).
Baudat, G. & Anouar, F. Kernel-based methods and function approximation. Proc. Int. Jt. Conf. Neural. Netw. 2, 1244–1249 (2001).
Lu, Z. et al. The expressive power of neural networks: a view form the width. Adv. Neural. Inf. Proc. Sys. 30 (2017).
Santurkar, S., Tsipras, D., Ilyas, A. & Madry, A. How does batch normalization help optimization?. Adv. Neural. Inf. Process. Syst. 31, 2488–2498 (2018).
Maas, A. L., Hannun, A. Y. & Ng, A. Y. Rectifier nonlinearities improve neural network acoustic models. Icml ’13 28, 6 (2013).
Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man. Cybern. SMC-9, 62–66 (1979).
Xu, X., Xu, S., Jin, L. & Song, E. Characteristic analysis of Otsu threshold and its applications. Pattern Recognit. Lett. 32, 956–961 (2011).
Kabkab, M., Hand, E. & Chellappa, R. On the size of convolutional neural networks and generalization performance. In 2016 23rd International Conference on Pattern Recognition (ICPR) 3572–3577 (IEEE, 2016).
Han, S. et al. Learning both weights and connections for efficient neural network. Adv. Neural. Inf. Proc. Sys. 2015, 1135–1143 (2015).
van der Walt, S. et al. scikit-image: image processing in Python. PeerJ 2, e453 (2014).
Paszke, A. et al. Automatic differentiation in PyTorch. NIPS 2017 (2017).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. ICLR 2015, 1–15 (2015).
Nair, V. & Hinton, G. Rectified linear units improve restricted Boltzmann machines. Int. Conf. on Mach. Learn. (2010).
Scherer, D., Müller, A. & Behnke, S. Evaluation of pooling operations in convolutional architectures for object recocnition. Art. Neural. Net-ICANN 6354, 92–101 (2010).
Acknowledgements
J.P.H and E.A.S acknowledge support through the National Science Foundation, Division of Materials Research, Metals and Metallic Nanostructures Program under Grant 1809398. This research used resources of the Center for Functional Nanomaterials, which is a U.S. DOE Office of Science Facility, at Brookhaven National Laboratory under Contract No. DE-SC0012704, which also provided support to D.N.Z. The data acquisition were initially supported under Laboratory Directed Research and Development funding at Brookhaven National Laboratory. The authors thank Yuwei Lin and Shinjae Yoo from Brookhaven National Laboratory for their insights and comments on the manuscript.
Author information
Authors and Affiliations
Contributions
E.A.S, D.N.Z., and R.M. conceived of the ideas for data analysis and experimentation. D.N.Z. collected TEM images with minor contributions from J.P.H., and computational experiments and model design were performed by J.P.H. with guidance from R.M. All authors contributed to preparing the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Horwath, J.P., Zakharov, D.N., Mégret, R. et al. Understanding important features of deep learning models for segmentation of high-resolution transmission electron microscopy images. npj Comput Mater 6, 108 (2020). https://doi.org/10.1038/s41524-020-00363-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-020-00363-x
This article is cited by
-
nNPipe: a neural network pipeline for automated analysis of morphologically diverse catalyst systems
npj Computational Materials (2023)
-
Mapping microstructure to shock-induced temperature fields using deep learning
npj Computational Materials (2023)
-
Time-resolved transmission electron microscopy for nanoscale chemical dynamics
Nature Reviews Chemistry (2023)
-
Bridging microscopy with molecular dynamics and quantum simulations: an atomAI based pipeline
npj Computational Materials (2022)
-
A reusable neural network pipeline for unidirectional fiber segmentation
Scientific Data (2022)