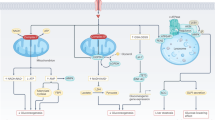
Abstract
Iron is essential for both microorganisms and their hosts. Although effects of dietary iron on gut microbiota have been described, the effect of systemic iron administration has yet to be explored. Here, we show that dietary iron, intravenous iron administration, and chronic transfusion in mice increase the availability of iron in the gut. These iron interventions have consistent and reproducible effects on the murine gut microbiota; specifically, relative abundance of the Parabacteroides and Lactobacillus genera negatively correlate with increased iron stores, whereas members of the Clostridia class positively correlate with iron stores regardless of the route of iron administration. Iron levels also affected microbial metabolites, in general, and indoles, in particular, circulating in host plasma and in stool pellets. Taken together, these results suggest that by shifting the balance of the microbiota, clinical interventions that affect iron status have the potential to alter biologically relevant microbial metabolites in the host.
Similar content being viewed by others
Introduction
Iron is a critically important nutrient, required for the viability of virtually all organisms.1 Iron deficiency is the most common form of malnutrition worldwide, increasing the risks of disability and death in >2 billion people.2 Efforts to prevent and correct iron deficiency by iron supplementation, particularly in children in low-income countries with increased exposure to pathogenic bacteria and parasites, have resulted in increased incidence of infection and death.3,4,5,6 One leading hypothesis to explain how iron supplementation adversely affects outcomes is through modification of the microbiota, promoting the growth of pathogenic bacteria, and hampering the survival of protective microbes.7 Indeed, dietary iron fortification of infants in low-income countries does alter the gut microbiota, favoring enteric pathogens, increasing intestinal inflammation and diarrhea, and reducing the efficacy of antibiotic treatment against potential enteropathogens.8,9,10 However, it is not known whether changing systemic iron levels by blood transfusion has similar effects.
Red blood cell transfusion is a prominent, but often inadvertent method of iron delivery. In patients undergoing hematopoietic stem cell transplantations (HSCT), transfusion-induced iron overload is a risk factor for poorer outcomes.11,12,13 In particular, transfusional iron overload is associated with an increased incidence of infection in the transplant setting.14,15,16 Furthermore, owing to patient blood management efforts to reduce adverse effects of red blood cell transfusions, intravenous iron is increasingly administered to anemic patients with iron deficiency.17 Nonetheless, it is unknown whether these interventions influence the gut microbiota similarly to dietary iron manipulations.
In this study, we generated a mouse model of transfusional iron overload to investigate the impact of transfusions on iron-associated alterations to commensal flora in the gut. Furthermore, we compared these results with those obtained in murine models of iron manipulations using dietary and intravenous approaches. Finally, given the iron-mediated changes observed in the gut microbiota, we examined the potential effects on microbial metabolic pathways and altered microbial metabolites circulating in the host. Taken together, these findings represent an important step toward better understanding the role of iron replacement therapies and transfusional iron overload on the gut microbiota and as a possible cause of dysbiosis leading to altered health outcomes.
Results
Intravenous iron and chronic transfusions increase fecal iron in mice fed either iron-deficient or iron-replete diet
Mice fed an iron-deficient diet became anemic, which was corrected by either intravenous iron or transfusion (Fig. 1a). In mice-fed iron-replete or iron-supplemented diets, chronic transfusion resulted in mild polycythemia. Splenic and hepatic iron content increased with increasing dietary iron, and with intravenous iron or chronic transfusion; however, liver iron increased more with intravenous iron as compared with transfusion (Fig. 1b, c). Serum ferritin levels paralleled liver iron levels (Fig. 1d). Chronic transfusion and intravenous iron increased hepcidin levels in mice fed iron-deficient or iron-replete diets, but did not significantly affect hepcidin levels in mice fed an iron-supplemented diet (Fig. 1e). Mean transferrin saturation was also below 50% for all groups except those on an iron-supplemented diet transfused or infused with iron (Fig. 1f); suggesting that significant levels of labile plasma iron were not produced in these former groups.18 The iron content in the diet was reflected in the fecal iron concentration. Furthermore, in parallel with hepcidin levels, chronic transfusion or intravenous iron increased fecal iron concentrations in mice fed iron-deficient or iron-replete diets, but not in mice fed an iron-supplemented diet (Fig. 1g). Furthermore, the various treatments did not induce liver toxicity as evidenced by no difference in alanine and aspartate aminotransferase (ALT and AST, respectively) levels (Fig. 1h, i), body weight (Fig. 1j), and microscopic histologic findings in the liver (Supplementary Fig. 1). Finally, the treatments only had a minor effect on expression levels of pro- and anti-inflammatory molecules in the liver and duodenum (Supplementary Fig. 2).
Iron status. a–g Iron parameters as labeled in each panel, were measured for each mouse on an iron-deficient (Def; n = 15), iron-replete (Rep; n = 15), or iron-supplemented (Sup; n = 20) diet, with intravenous iron infusion (IV; n = 10 per group) or weekly blood transfusions (Trans; n = 10 per group). Results shown are combined from four separate and complementary experiments. h–i ALT and AST activity in the liver was measured in the same cohorts of mice for one representative replicate (n = 5 per group); j Body weight was recorded in three of the complementary experiments. Dot plots graphs report mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 by one-way ANOVA with Tukey’s multiple comparison test
Dietary and systemic iron administration affect the microbiota composition
The dietary iron content affected microbial alpha- and beta diversity, as measured by Shannon index and weighted UniFrac, respectively (Fig. 2a–e, Supplementary Table 1). When stratifying by dietary iron content, systemic iron administration by intravenous infusion or chronic transfusion affected both alpha- and beta diversity; however, this effect became less significant as the dietary iron content increased. Furthermore, alpha diversity was positively correlated with liver iron levels regardless of the diet and how iron was delivered systemically (Fig. 2f). Finally, 16 S copy number analysis of cecal content revealed only minor differences in bacterial load among the groups (Supplementary Fig. 3). Taken together, these results suggest that systemic iron administration, by either transfusion or intravenous iron, modifies the murine gut flora.
Iron status modulates microbiota diversity. Weighted UniFrac Principal Coordinate Analysis (PCoA) plot of a representative experiment of cohorts of mice a fed an iron-deficient (Def), iron-replete (Rep) or iron-supplemented (Sup) diet; b fed an iron-deficient diet and either infused with intravenous iron (Def+IV iron) or chronically transfused (Def+Trans); c fed an iron-replete diet and either infused with intravenous iron (Rep+IV iron) or chronically transfused (Rep+Trans); and d fed an iron-supplemented diet and either infused with intravenous iron (Sup+IV iron) or chronically transfused (Sup+Trans). e Shannon alpha diversity index of the mice represented in a–d and f Spearman correlation dot plot comparing logarithm of Shannon alpha diversity index and liver iron for each mouse in a representative experiment; n = 5 mice per group; *p < 0.05, **p < 0.01
Iron levels negatively correlate with bacteria belonging to the Lactobacillus and Parabacteroides genera and positively correlate with bacteria belonging to the Clostridia class
To investigate whether iron levels per se affect specific bacteria, we performed a Spearman’s correlation analysis between iron measures (e.g., fecal, liver and spleen iron, serum ferritin and hepcidin) and the relative abundance of bacteria OTUs aggregated at different taxonomic rank. Only bacterial taxa detected in all four complementary experiments were included in this analysis (Supplementary Data 1). Lactobacillus and Parabacteroides genera exhibited the strongest negative association with iron status (Fig. 3, Supplementary Data 2). Conversely, only bacteria belonging to the Clostridia class positively correlated with iron status.
Correlation of bacterial taxa with iron status. Heatmap of Spearman’s correlation between iron parameters and bacterial taxonomic levels of all mice combined from four experiments. Red indicates a negative and green a positive correlation (color intensity reflective of Spearman’s r). a p_ = phylum, b c_ = class, c o_ = order, d f_ = family, e g_ = genus, f s_ = species. *p < 0.0001 for the Spearman’s correlation
Iron status modifies the gut microbiota independently of the administration route
To investigate whether administration of iron by intravenous infusion or chronic transfusion differentially affects the microbiota, we performed Linear Discriminant Analysis (LDA) effect size (LEfSe)19 on the same bacterial data set used for the Spearman’s correlation analysis. The gut microbiota composition of cohorts of mice receiving intravenous iron or chronic transfusion were compared with cohorts of untreated mice for each respective diet. Differentially abundant bacterial taxa at different taxonomic rank with a p value of 0.01 and LDA score higher than 2.0 were identified. Twenty-four and 20 bacteria at different taxonomic rank were modified by intravenous iron infusion and chronic transfusion, respectively (Fig. 4). Among these, 17 were common to both iron treatments.
LEfSe analysis of bacterial taxa modulated by either intravenous iron infusion or chronic transfusion. Table and corresponding Venn diagrams of the total number of bacteria at different taxonomic rank increased (+) or decreased (−) by intravenous iron infusion (IV Iron; blue), chronic transfusions (yellow), or both (green) in the three different diet groups, as labeled. p_ = phylum, c_ = class, o_ = order, f_ = family, g_ = genus; *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001 of Least Discriminant Analysis effect size (LDA) score
To confirm the LEfSe results and to account for the variability between experiments, a mixed model analysis was performed, considering diet and parenteral treatments as fixed effects and each experiment as a random effect. Using mice fed an iron-replete diet as the control, 18 bacterial taxa emerged as being modified by diet and/or parenteral iron treatment with a p value of <0.0001 (Fig. 5, Table 1). Among these, 17 bacterial taxa were affected by intravenous iron administration or chronic transfusion in a similar fashion to the dietary iron supplementation. Taken together, these results suggest that the delivered iron plays the dominant role in modifying the microbiota, irrespective of the route of administration.
Relative abundance of all taxonomic ranks. Relative abundance of all taxa from four combined experiments in cohorts of mice fed an iron-deficient (Def), iron-replete (Rep) or iron-supplemented diet (Sup), and from cohorts of mice infused with intravenous iron (IV) or chronically transfused (Trans). Each bar represents the mean of the relative abundance of the corresponding taxa, across four complementary experiments (n = 10–20 mice per group); * denotes bacteria modulated by diet and/or treatments (IV iron or chronic transfusion) with p < 0.0001 from mixed model analysis
Combining the three different approaches to identify bacterial taxa modified by iron status (i.e., Spearman’s correlation, Fig. 3; LEfSe analysis, Fig. 4; mixed effects model, Fig. 5, Table 1), Proteobacteria phylum, Clostridia class in phylum Firmicutes, and its dependent bacterial taxa Clostridiales order, Clostridiaceae and Peptococcaceae families and Dehalobacterium genus, and Parabacteroides genus in phylum Bacteroidetes, were most consistently associated with iron levels and iron treatment (Fig. 6).
Combination of all three approaches used to analyze the effects of iron on the microbiome. Table and Venn diagram indicating the distribution of the bacteria at different taxonomic rank modified by intravenous iron and chronic transfusion that were identified by Spearman’s correlation (25), LEfSe (17) and mixed model analysis (17) from the four complementary experiments. Taxa highlighted in dark gold were identified by all three approaches. p_ = phylum, c_ = class, o_ = order, f_ = family, g_ = genus, s_ = species
Iron status affects the predicted functional profile of the gut microbiota
To predict whether iron levels affect the functional profile of the gut microbiota, PICRUSt analysis was performed to predict potential pathways affected by iron availability in the gut. Based on fecal iron content, we compared the cohort of mice fed an iron-deficient diet with cohorts fed an iron-replete diet, or an iron-deficient diet with intravenous iron administration, or an iron-deficient diet with chronic transfusion. At Kyoto Encyclopedia of Genes and Genomes (KEGG) level 3, and as compared with mice fed an iron-deficient diet, 100 pathways were predicted to be significantly different in mice fed an iron-replete diet. Furthermore, 101 and 54 pathways were predicted in the cohorts of mice infused with intravenous iron or chronically transfused, respectively. Among these, 51 pathways were shared by all three cohorts of mice with increased iron level as compared with the iron-deficient mice (Supplementary Fig. 4a, b). PCA analysis confirmed the overlap and similarity of the predicted pathways at KEGG level 3 and demonstrated that the mice in the cohort receiving only the iron-deficient diet cluster separately from the three groups with increased fecal iron (Supplementary Fig. 4c). Thus, the PICRUSt analysis predicted that iron status may functionally affect microbial metabolism and prompted an exploration regarding whether microbial metabolites are modified by iron status.
Untargeted metabolomics analysis reveals that metabolites of tryptophan degradation are negatively associated with increasing iron availability
To investigate metabolic changes associated with iron status, untargeted metabolomics analysis was performed on cecal feces, stool pellets, and plasma. Spearman’s correlation between detected metabolites and iron parameters revealed that increasing iron levels correlate with levels of multiple metabolites (Supplementary Fig. 5, Supplementary Data 3–5). Of the microbially produced metabolites examined, indoles, which are derived from tryptophan degradation with protective effects on the gut mucosa,20,21 exhibited the strongest negative correlation with fecal and plasma iron levels (Fig. 7).
Iron modulation of “Indole and Tryptophan” pathway metabolites. a Spearman’s correlation of metabolites belonging to the “Indole and Tryptophan” pathway in cecal feces, stool pellets, and plasma as correlated to standard iron parameters. b Correlation between Log10 of quantified indole Area Under the Curve (AUC) and Log10 of liver iron in stool pellets and plasma, as indicated. *p < 0.0001 for the Spearman’s correlation
Discussion
Iron is a vital element for biological functions in both eukaryotic and prokaryotic cells. Increased iron availability affects bacterial virulence22,23 and changes the balance of commensal bacteria in the gut, promoting growth of more pathogenic Enterobacteriaceae at the expense of protective bacteria that are less dependent on iron (e.g., Lactobacilli).7,24,25 However, the effects of iron delivered systemically, such as by intravenous iron therapy or red blood cell transfusion, have not been fully characterized.26 In the current study, although dietary iron had the greatest impact on fecal iron levels, both intravenous iron infusion and chronic transfusion affected fecal iron levels when murine diets were not iron supplemented. Fecal iron was associated with alpha and beta diversity changes in the composition of the murine gut microbiota; furthermore, intravenous iron infusion and chronic red blood cell transfusion had similar effects. Taken together, these findings suggest that the delivery of iron, irrespective of the route of administration, has strong effects on the murine gut microbiota.
Owing to the variability of the baseline microbiota in repeat experiments,27,28,29 we identified bacterial taxa that were similarly modulated by iron across four complementary experiments, analyzing a total of 110 mice, and integrating three different analytical approaches (i.e., LEfSe, mixed model analysis, and Spearman’s correlation); this was done to minimize the effects of non-controllable variables on the microbiota. Using this strategy, bacterial taxa belonging to Clostridia class (Clostridiales order, Clostridiaceae and Peptococcaceae families, and Dehalobacterium genus) were found to proliferate with increasing levels of iron; in contrast, Parabacteroides genus in the Bacteriodetes phylum and Proteobacteria phylum decreased in abundance with increasing iron availability. Finally, there was a significant negative correlation between bacteria in the Lactobacillaceae family and iron status using two of the three analytical approaches (i.e., Spearman’s correlation and mixed model analysis). Clostridiaceae, Lactobacillaceae, and Parabacteroides were previously shown to be modified by oral iron;7,8,9,30,31 the current results extend these findings to systemic iron administration. In contrast, very little is known about the relationship between iron and members of the Peptococcaceae family, Gram-positive obligate anaerobic bacteria, which are positively associated with production of trimethylamine N-oxide (TMAO),32 a metabolite that increases thrombotic risk by enhancing platelet activation.33 Thus, in this study, we demonstrate that this bacterial taxa has a strong positive association with iron availability.
Although we did not observe major differences in systemic and duodenal inflammation (Supplementary Fig. 2) in these wild-type and healthy mice, the protective role of Lactobacillaceae and Parabacteroides in the gut has been shown in a mouse model of dextran sulfate sodium-induced colitis, where the presence of Lactobacillaceae and Parabacteroides have been associated with increased intestinal barrier function and reduced intestinal inflammation, respectively.34,35 Furthermore, in both murine and human studies, Lactobacillus and Parabacteroides are beneficial in the setting of HSCT and acute graft-versus-host disease (GVHD), the latter a dangerous adverse outcome of HSCT.36,37,38 Thus, our findings that Lactobacillus and Parabacteroides are reduced by chronic transfusion are particularly relevant in the context of HSCT, considering that these patients often become iron overloaded as a result of their transfusion requirements and that this iron overload portends worse outcomes.11,12 Finally, patients with chronic kidney disease are often treated with intravenous iron and microbiome studies in these patients suggest dysbiosis with a particular reduction in Lactobacillus species.39,40 Whether iron is responsible for these findings in patients with chronic kidney disease has yet to be investigated.
The gut microbiota affects the host by producing various metabolites. In this study, we show that these iron-mediated microbiota changes are also associated with changes in metabolites produced by microbiota, particularly indoles, which originate from the microbial metabolism of tryptophan,41,42 in both stool pellets and plasma. Using untargeted metabolomics profiling, we demonstrate that increased iron levels are associated with decreased indole metabolites. Indoles protect the gut mucosa, by increasing tight-junction protein expression, reducing epithelial inflammation,20 and conferring resistance to dextran sodium sulfate-induced colitis in mice.21 Furthermore, acting via Type I interferon, indoles reduce GVHD pathology and mortality in mice43 and low levels of urinary 3-indoxyl-sulfate are associated with higher transplant-related mortality and lower overall survival in allogeneic HSCT patients.44 Whether these iron-mediated alterations to the commensal microbiota and to microbiota-derived metabolites, such as indoles, induce some of the clinical effects associated with iron overload, remains unknown and requires further investigation.
Although iron metabolism in laboratory mice is generally similar to that of humans,45 there are notable differences. For example, mice do not absorb heme iron as well as humans.46 Furthermore, mice have greater iron losses relative to iron stores,45 and have a more active iron excretion pathway as compared with humans.47,48,49 Thus, a limitation of this study is that the increases in fecal iron content observed with chronic transfusion or intravenous iron infusion in this murine model may not fully translate to humans. However, we hypothesize that increasing iron status in humans may lead to an increased iron concentration in the gut lumen through the action of hepcidin, the iron-regulatory hormone,1 which would prevent dietary iron absorption, resulting in increasing iron availability to gut microbiota through sloughing of iron-loaded enterocytes into the gastrointestinal lumen at the end of their lifespan. Furthermore, limited bile excretion pathways may exist in iron-overloaded humans.47,50,51 To determine how well these murine findings translate to the human setting, we are currently conducting a human study (ClinicalTrials.gov # NCT02990988) in which stool samples are collected from iron-deficient volunteers before and after repletion by intravenous iron infusion. Finally, another limitation of our study is that other forms of iron administration were not tested; however, chronic transfusion, oral iron supplementation, and intravenous iron represent the main routes of changing iron status in patients.
In conclusion, we observe that intravenous iron infusions and chronic transfusion, in a mouse model, modified fecal iron levels, and the gut microbiota in similar ways to those observed with dietary iron supplementation. These changes affected multiple predicted functional pathways in gut microbiota and, most importantly, metabolite levels, such as indoles, in feces and plasma. Although the human clinical implications of these findings in a murine model are yet to be determined, the fact that indoles are known to affect clinical outcomes and, in our setting, were modulated by iron, strengthens the importance of better understanding iron-mediated host-microbiota interactions.
Methods
Mice
Weanling female wild-type C57BL/6 mice were purchased from Charles River Laboratories (Stone Ridge, NY), housed in a pathogen-free facility, and used at 3 weeks of age. Cohorts of mice were fed ad libitum either an iron-deficient (TD.110592, Envigo, 0–5 ppm Fe), iron-replete (TD. 110593, Envigo, 45 ppm Fe), or iron-supplemented (TD.110594, Envigo, 220 ppm Fe) diet for 6 weeks. Two additional cohorts of mice were generated for each of these dietary groups by intravenous infusion of iron dextran (0.33 mg weekly × 6; Henry Schein Animal Health, Dublin, OH; 2 mg iron total over 6 weeks) or transfusing red blood cells (0.33 mL at 60% hematocrit weekly × 6 using fresh blood collected from wild-type C57BL/6 mice on a regular chow diet; mouse equivalent to 10 human units of blood transfused or 2 mg total iron administered). Following 6 weeks on defined diets, with or without additional iron treatment, all cohorts were euthanized and tissues, blood, and feces from rectum and cecum were collected for analysis. Between 10 and 20 mice per group total, in four complementary experiments were analyzed. For all experiments, only mice on the same diet and provided with the same treatment were cohoused (n = 5 per cage), so that coprophagic behavior would not influence the results. All animal studies were approved by the Columbia University Institutional Animal Care and Use Committee.
16 S sequencing and copy number, microbiota, and PICRUSt analysis
Cecal fecal samples were collected and DNA extracted using QIAamp power fecal DNA kit (Qiagen). 16 S rDNA V4 or V3–V4 variable regions were amplified by PCR and sequenced using the Illumina MiSeq platform. For the first three experiments, 16 S rDNA V4 sequencing was performed at MR DNA (www.mrdnalab.com, Shallowater, TX, USA) and operational taxonomic units (OTUs) were defined by clustering at 97% similarity. Final OTUs were taxonomically classified using BLASTn against a curated database derived from Greengenes, RDPII and NCBI (www.ncbi.nlm.gov, www.greengenes.lbl.gov, http://rdb.cme.msu.edu). For the fourth experiment, 16 S rDNA V3–V4 sequencing was performed at the Microbiota Core facility at the Columbia University Irving Medical Center (CUIMC). QIIME52 was used to cluster OTUs at 97% sequence similarity and aligned against the Greengenes database with noise filtering. A minimum cut off of 10,000 counts for a sample to be analyzed was used for all experiments. The relative proportion of multiple OTU sequences within each sample, mapping to the designated taxonomic classification, was used to perform the mixed model analysis and the pairwise linear discriminant analysis effect size (LEfSe),19 excluding the bacterial taxa not detected in all four experiments. 16 S rDNA data were also used for metagenome functional prediction with PICRUSt on the online Galaxy platform (http://huttenhower.sph.harvard.edu/galaxy/).53 Sequences were mapped to the Greengenes ver 13.5 database and normalized to check for differences in 16 S rDNA copy number between OTUs. STAMP54 software was used to analyze KEGG Level 2 and 3 pathway profiles. Bacterial load was analyzed by qPCR of 16 S rDNA copies using RT2 SYBR green ROX qPCR Mastermix (Qiagen) on an AriaMx Real-Time PCR system as described by Staffas et al.55
Metabolomics
Plasma and weighted cecal feces and stool pellets were collected after 6 weeks of dietary manipulation and/or iron treatment. Blood was obtained via cardiac puncture into heparinized syringes and plasma separated after centrifugation at 2500 × g for 10 min at 4 °C. All samples were stored at − 80 °C until further processing. Metabolomics analyses of indole metabolites were performed as previously described.56 In brief, stable isotope standards tryptophan (15N2 - NLM-800–0.25) were purchased from Cambridge Isotopes and samples were thawed on ice, then 20 µL (fluid) or 10 mg (tissue) was extracted with 480 µL or 1 mL respectively, of ice cold extraction buffer (5:3:2 MeOH:MeCN:H2O) containing 0.1 μM each of the heavy standards. Extraction was performed by vigorous agitation at 4 °C for 30 min followed by centrifugation at 12,000 rpm, 4 °C for 10 min A 100 μL aliquot of supernatant was transferred to a glass vial, dried under N2 flow, and resuspended in an equal volume of water containing 0.1% (v/v) formic acid. Aqueous extracts were analyzed by ultra-high pressure liquid chromatography-mass spectrometry (UHPLC-MS) on a Thermo Vanquish UHPLC (San Jose, CA) coupled to a Thermo Q Exactive mass spectrometer (Bremen, Germany) via positive electrospray ionization. Solvents were water (phase A) and acetonitrile (phase B) supplemented with formic acid (0.1%) and flow rate was 0.25 mL/min. Metabolites were separated using a Kinetex C18 (Phenomenex, Torrance, CA) column (2.1 × 150 mm, 1.7 um) with a 6 min gradient of 0–2 min 2% B; 2–2.5 min increase to 25% B; 2.5–4 min hold at 25% B; 4–4.01 min decrease to 2% B; 4.01–6 min hold at 2% B. The Q Exactive mass spectrometer was operated in full scan mode over the range of 65–950 m/z. Samples were randomized and a quality control sample was injected every 10 runs. Data analysis was performed using Maven (Princeton University) following file conversion by MassMatrix (Case Western Reserve University). Other metabolites were analyzed through the same platform, with methods and workflows extensively described in prior publications.57
Iron-related measurements
All iron-related measurements were obtained at the time of euthanasia. Non-heme iron of spleen, liver, and stool pellets were determined using a wet ashing procedure.58 In brief, the wet weight of samples obtained at necropsy was quantified and portions of liver, spleen, or stool pellets, were placed in 2 ml micro-tubes. Following desiccation at 65 °C for 24 h, 1 ml of acid digestion mixture (3 M hydrochloric acid, 10% trichloracetic acid) was added and samples were heated at 65 °C for an additional 24 h. The acidified sample (50 μl) was then incubated for 30 min with 200 μl of chromogen (1.6 M bathophenanthroline, 2 M sodium acetate, 11.5 M thioglycolic acid). Absorbance at 535 nm of samples and iron standards was measured in duplicate and mean values used for calculating total iron. Iron transferrin saturation in plasma was calculated using Iron/TIBC Reagent Set (Pointe Scientific, Canton, MI). Plasma ferritin, plasma hepcidin, and liver ALT levels in liver were measured by ELISA following the manufacturer’s instructions (Kamiya Biomedical, Seattle, WA; Intrinsic LifeSciences, La Jolla, CA; and Biomatik, Wilmington, DE, respectively). AST was measured in liver using the AST activity assay kit (Sigma Aldrich, St. Louis, MO).
Histology
At necropsy, liver was removed, fixed overnight with 10% neutral-buffered formalin and embedded in paraffin. Sections were stained with hematoxylin and eosin and images captured using a 10 × Olympus objective on an Olympus BX51 microscope and an Infinity HD digital camera (B&B Microscopes Ltd.)
Quantitative reverse transcriptase PCR
Liver and duodenal tissue (20–50 mg) were collected at necropsy in RNAlater RNA Stabilization Solution (Invitrogen). After 24 h of incubation at +4 °C, tissues were homogenized in RLT buffer for RNA extraction using RNeasy Plus mini Kit (Qiagen) followed by processing with RT2 First strand kit (Qiagen) for cDNA synthesis. Quantitative PCR for Mcp1 (Fw 5′-CCTGGATCGGAACCAAATGAGATCAG-3′; Rev 5′-AGTGCTTGAGGTGGTTGTGGAA-3′), Lcn2 (Fw 5′-CCACCACGGACTACAACCAG-3′; Rev 5′-AGCTCCTTGGTTCTTCCATACA-3′), Saa1/2 (Fw 5′-GCGAGCCTACACTGACATG A-3′; Rev 5′-GGCAGTCCAGGAGGTCTGTA-3′) and Il10 (Fw 5′-CGGGAAGACAATAACTG-3′; Rev 5′-ATTTCCGATAAGGCTTGG-3′) genes was performed using RT2 SYBR® green ROX qPCR Mastermix (Qiagen) on an AriaMx Real-Time PCR system and normalized to the housekeeping gene Gapdh (Fw 5′-ATGACTCCACTCACGGCAAAT TC -3′; Rev 5′-ACACCAG TAGACTCCACGACATAC-3′).
Statistical analysis
Significance among means was calculated using one-way ANOVA with the Tukey Multiple Comparison Test or Kruskal–Wallis test with Dunn’s Multiple Comparison Test, as appropriate. A p value of <0.05 was considered significant. Statistical analyses were performed using Prism 7 (GraphPad Software, Inc., La Jolla, CA) and means ± standard deviations (SD) are illustrated in the figures. PermANOVA (Permutational Multivariate Analysis of Variance Using Distance Matrices) test of weighted and unweighted UniFrac distance was calculated with R (version 3.4.2) using the vegan package (adonis function). For LEfSe analysis, a p value of 0.01 for the Kruskal–Wallis test and LDA score higher than 2.0 were used. SAS studio was used for mixed model analysis and Spearman’s correlation. A conservative p value of <0.0001 was considered significant in these analyses to limit false discovery given a Bonferroni correction for examining 400 bacterial taxa. The heatmap and dot plot graphs of Spearman’s correlation were generated using ggplot2 and ggpubr packages in R software, respectively. Venn diagrams describing the overlap between the detected bacterial taxa and functional profiles were computed with Biovenn59 and manually edited with Inkscape 0.92 (https://inkscape.org/).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The authors declare that all relevant data supporting the findings of the study are available in this article and its Supplementary Information file, or from the corresponding author upon request.
References
Andrews, N. C. Forging a field: the golden age of iron biology. Blood 112, 219–230 (2008).
Camaschella, C. Iron-deficiency anemia. N. Engl. J. Med. 372, 1832–1843 (2015).
Barry, D. M. & Reeve, A. W. Increased incidence of gram-negative neonatal sepsis with intramuscula iron administration. Pediatrics 60, 908–912 (1977).
Brewster, U. C., Coca, S. G., Reilly, R. F. & Perazella, M. A. Effect of intravenous iron on haemodialysis catheter microbial colonization and blood-borne infection. Nephrology 10, 124–128 (2005).
Sazawal, S. et al. Effect of routine prophylactic supplementation with iron and folic acid on admission to hospital and mortality in preschool children in a high malaria transmission setting: community based, randomised, placebo-controlled trial. Lancet 367, 133–143 (2006).
Drakesmith, H. & Prentice, A. M. Hepcidin and the iron-infection axis. Science 338, 768–772 (2012).
Zimmermann, M. B. et al. The effects of iron fortification on the gut microbiota in African children: a randomized controlled trial in Côte d’Ivoire. Am. J. Clin. Nutr. 92, 1406–1415 (2010).
Jaeggi, T. et al. Iron fortification adversely affects the gut microbiome, increases pathogen abundance and induces intestinal inflammation in Kenyan infants. Gut 64, 731–742 (2015).
Paganini, D., Uyoga, M. A. & Zimmermann, M. B. Iron fortification of foods for infants and children in low-income countries: effects on the gut microbiome, gut inflammation, and diarrhea. Nutrients 8, 494 (2016).
Paganini, D. et al. Iron-containing micronutrient powders modify the effect of oral antibiotics on the infant gut microbiome and increase post-antibiotic diarrhoea risk: a controlled study in Kenya. Gut 68, 645–653 (2018).
Pullarkat, V. et al. Iron overload adversely affects outcome of allogeneic hematopoietic cell transplantation. Bone Marrow Transplant. 42, 799–805 (2008).
Meyer, S. C. et al. Prognostic impact of posttransplantation iron overload after allogeneic stem cell transplantation. Biol. Blood Marrow Transplant. 19, 440–444 (2013).
Armand, P. et al. Does iron overload really matter in stem cell transplantation? Am. J. Hematol. 87, 569–572 (2012).
Fernández-Ruiz, M. et al. Association between baseline serum hepcidin levels and infection in kidney transplant recipients: potential role for iron overload. Transpl. Infect. Dis. 20, e12807 (2018).
Kanda, J. et al. Pretransplant serum ferritin and C-reactive protein as predictive factors for early bacterial infection after allogeneic hematopoietic cell transplantation. Bone Marrow Transplant. 46, 208–216 (2011).
Miceli, M. H. et al. Iron overload is a major risk factor for severe infection after autologous stem cell transplantation: a study of 367 myeloma patients. Bone Marrow Transplant. 37, 857–864 (2006).
Desai, N., Schofield, N. & Richards, T. Perioperative patient blood management to improve outcomes. Anesth. Analg. 127, 1211–1220 (2018).
de Swart, L. et al. Second international round robin for the quantification of serum non-transferrin-bound iron and labile plasma iron in patients with iron-overload disorders. Haematologica 101, 38–45 (2016).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Alexeev, E. E. et al. Microbiota-derived indole metabolites promote human and murine intestinal homeostasis through regulation of iinterleukin-10 receptor. Am. J. Pathol. 188, 1183–1194 (2018).
Shimada, Y. et al. Commensal bacteria-dependent indole production enhances epithelial barrier function in the colon. PLoS ONE 8, e80604 (2013).
Sheldon, J. R., Laakso, H. A. & Heinrichs, D. E. Iron acquisition strategies of bacterial pathogens. Virulence Mech. Bact. Pathog. 4, 43–85 (2016).
Skaar, E. P. The battle for iron between bacterial pathogens and their vertebrate hosts. PLoS Pathog. 6, e1000949 (2010).
Archibald, F. Lactobacillus plantarum, an organism not requiring iron. FEMS Microbiol. Lett. 19, 29–32 (1983).
Pandey, A., Bringel, F. & Meyer, J.-M. Iron requirement and search for siderophores in lactic acid bacteria. Appl. Microbiol. Biotechnol. 40, 735–739 (1994).
Lee, T. et al. Oral versus intravenous iron replacement therapy distinctly alters the gut microbiota and metabolome in patients with IBD. Gut 66, 863–871 (2017).
Ericsson, A. C. et al. The influence of caging, bedding, and diet on the composition of the microbiota in different regions of the mouse gut. Sci. Rep. 8, 4065 (2018).
Franklin, C. L. & Ericsson, A. C. Microbiota and reproducibility of rodent models. Lab. Anim. 46, 114–122 (2017).
Wesolowska-Andersen, A. et al. Choice of bacterial DNA extraction method from fecal material influences community structure as evaluated by metagenomic analysis. Microbiome 2, 19 (2014).
Kortman, G. A. M. et al. Low dietary iron intake restrains the intestinal inflammatory response and pathology of enteric infection by food-borne bacterial pathogens. Eur. J. Immunol. 45, 2553–2567 (2015).
Dostal, A. et al. Effects of iron supplementation on dominant bacterial groups in the gut, faecal SCFA and gut inflammation: a randomised, placebo-controlled intervention trial in South African children. Br. J. Nutr. 112, 547–556 (2014).
Org, E. et al. Relationships between gut microbiota, plasma metabolites, and metabolic syndrome traits in the METSIM cohort. Genome Biol. 18, 70 (2017).
Zhu, W. et al. Gut microbial metabolite TMAO enhances platelet hyperreactivity and thrombosis risk. Cell 165, 111–124 (2016).
Anderson, R. C., Cookson, A. L., McNabb, W. C., Kelly, W. J. & Roy, N. C. Lactobacillus plantarum DSM 2648 is a potential probiotic that enhances intestinal barrier function. FEMS Microbiol. Lett. 309, no–no (2010).
Kverka, M. et al. Oral administration of Parabacteroides distasonis antigens attenuates experimental murine colitis through modulation of immunity and microbiota composition. Clin. Exp. Immunol. 163, 250–259 (2011).
Ladas, E. J. et al. The safety and feasibility of probiotics in children and adolescents undergoing hematopoietic cell transplantation. Bone Marrow Transplant. 51, 262–266 (2016).
Jenq, R. R. et al. Regulation of intestinal inflammation by microbiota following allogeneic bone marrow transplantation. J. Exp. Med. 209, 903–911 (2012).
Liu, C. et al. Associations between acute gastrointestinal GvHD and the baseline gut microbiota of allogeneic hematopoietic stem cell transplant recipients and donors. Bone Marrow Transplant. 52, 1643–1650 (2017).
Vaziri, N. D. et al. Chronic kidney disease alters intestinal microbial flora. Kidney Int. 83, 308–315 (2013).
Sampaio-Maia, B., Simões-Silva, L., Pestana, M., Araujo, R. & Soares-Silva, I. J. The role of the gut microbiome on chronic kidney disease. Adv. Appl. Microbiol. 96, 65–94 (2016).
Zelante, T. et al. Tryptophan catabolites from microbiota engage aryl hydrocarbon receptor and balance mucosal reactivity via interleukin-22. Immunity 39, 372–385 (2013).
Cervantes-Barragan, L. et al. Lactobacillus reuteri induces gut intraepithelial CD4+CD8αα+ T cells. Science 357, 806–810 (2017).
Swimm, A. et al. Indoles derived from intestinal microbiota act via type I interferon signaling to limit graft-versus-host disease. Blood 132, 2506–2519 (2018).
Weber, D. et al. Low urinary indoxyl sulfate levels early after transplantation reflect a disrupted microbiome and are associated with poor outcome. Blood 126, 1723–1728 (2015).
Coffey, R. & Ganz, T. Iron homeostasis: san anthropocentric perspective. J. Biol. Chem. 292, 12727–12734 (2017).
Fillebeen, C. et al. Mice are poor heme absorbers and do not require intestinal Hmox1 for dietary heme iron assimilation. Hematologica 100, e334–e337 (2015).
Green, R. et al. Body iron excretion in man: sa collaborative study. Am. J. Med. 45, 336–353 (1968).
Musumeci, M. et al. Iron excretion in iron dextran-overloaded mice. Blood Transfus. 12, 485–490 (2014).
Hunt, J. R., Zito, C. A. & Johnson, L. K. Body iron excretion by healthy men and women. Am. J. Clin. Nutr. 89, 1792–1798 (2009).
Brissot, P. et al. Iron overload and the biliary route. in Advances in Experimental Medicine and Biology (eds. Hershko, C., Konijn, A. M. & Aisen, P.) 356, 277–283 (Springer, Boston, MA, 1994).
Hultcrantz, R., Angelin, B., Björn-Rasmussen, E., Ewerth, S. & Einarsson, K. Biliary excretion of iron and ferritin in idiopathic hemochromatosis. Gastroenterology 96, 1539–1545 (1989).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Langille, M. G. I. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31, 814–821 (2013).
Parks, D. H. & Beiko, R. G. Identifying biologically relevant differences between metagenomic communities. Bioinformatics 26, 715–721 (2010).
Staffas, A. et al. Nutritional support from the intestinal microbiota improves hematopoietic reconstitution after bone marrow transplantation in mice. Cell Host Microbe 1–11 (2018). https://doi.org/10.1016/j.chom.2018.03.002.
Greene, L. I. et al. A role for tryptophan-2,3-dioxygenase in cd8 T-cell suppression and evidence of tryptophan catabolism in breast cancer patient plasma. Mol. Cancer Res. 17, 131–139 (2019).
Nemkov, T., Hansen, K. C. & D’Alessandro, A. A three-minute method for high-throughput quantitative metabolomics and quantitative tracing experiments of central carbon and nitrogen pathways. Rapid Commun. Mass Spectrom. 31, 663–673 (2017).
Overmoyer, B. A., McLaren, C. E. & Brittenham, G. M. Uniformity of liver density and nonheme (storage) iron distribution. Arch. Pathol. Lab. Med. 111, 549–554 (1987).
Hulsen, T., de Vlieg, J. & Alkema, W. BioVenn – a web application for the comparison and visualization of biological lists using area-proportional Venn diagrams. BMC Genomicsa 9, 488 (2008).
Acknowledgements
This work was supported in part by a grant from the National Heart, Lung, and Blood Institute, National Institutes of Health (R01 HL121275 and R21 HL145319 to EAH). We express our gratitude to Dr. Jonathan U. Peled (Memorial Sloan Kettering Cancer Center) for kindly providing the Blautia 16 S rDNA gene sequence plasmid for the 16 S copy number analysis and all the members of the Laboratory of Transfusion Biology (Columbia University Irving Medical Center) for their support and encouragement.
Author information
Authors and Affiliations
Contributions
F.L.C. and E.A.H. designed the study. F.L.C., B.S.W. and A.R. performed the experiments and collected data. A.D. carried out metabolomics analysis. F.L.C., M.K.A., D.F., A.C.U. and E.A.H. analyzed data. F.L.C. and EAH wrote the manuscript and all authors edited drafts and reviewed the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
Unrelated to the contents of the manuscript, A.D. declares to be the founder of Omix Technologies, Inc. and a consultant for Hemanext, Inc. The other authors declare no competing financial interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
La Carpia, F., Wojczyk, B.S., Annavajhala, M.K. et al. Transfusional iron overload and intravenous iron infusions modify the mouse gut microbiota similarly to dietary iron. npj Biofilms Microbiomes 5, 26 (2019). https://doi.org/10.1038/s41522-019-0097-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41522-019-0097-2
This article is cited by
-
Role of iron in host-microbiota interaction and its effects on intestinal mucosal growth and immune plasticity in a piglet model
Science China Life Sciences (2023)
-
Gut microbiota bridges dietary nutrients and host immunity
Science China Life Sciences (2023)
-
Plasma non-transferrin-bound iron uptake by the small intestine leads to intestinal injury and intestinal flora dysbiosis in an iron overload mouse model and Caco-2 cells
Science China Life Sciences (2023)
-
Transferrin predicts trimethylamine-N-oxide levels and is a potential biomarker of cardiovascular disease
BMC Cardiovascular Disorders (2022)
-
Supplementation with Sucrosomial® iron leads to favourable changes in the intestinal microbiome when compared to ferrous sulfate in mice
BioMetals (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.