Abstract
Axial chirality of biaryls can generate varied bioactivities. Gossypol is a binaphthyl compound made by cotton plants. Of its two axially chiral isomers, (−)-gossypol is the bioactive form in mammals and has antispermatogenic activity, and its accumulation in cotton seeds poses health concerns. Here we identified two extracellular dirigent proteins (DIRs) from Gossypium hirsutum, GhDIR5 and GhDIR6, which impart the hemigossypol oxidative coupling into (−)- and (+)-gossypol, respectively. To reduce cotton seed toxicity, we disrupted GhDIR5 by genome editing, which eliminated (−)-gossypol but had no effects on other phytoalexins, including (+)-gossypol, that provide pest resistance. Reciprocal mutagenesis identified three residues responsible for enantioselectivity. The (−)-gossypol-forming DIRs emerged later than their enantiocomplementary counterparts, from tandem gene duplications that occurred shortly after the cotton genus diverged. Our study offers insight into how plants control enantiomeric ratios and how to selectively modify the chemical spectra of cotton plants and thereby improve crop quality.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All relevant data supporting the findings of this study are available within the paper and its Supplementary Information. Public RNA-seq data can be obtained from the Sequence Read Archive repository (https://www.ncbi.nlm.nih.gov/sra) under accession nos. PRJNA248163, PRJNA265955 and PRJNA493958. Sequence data and gene IDs can be found in the Cottongen database (https://www.cottongen.org/). The template of DIR used for molecular modelling was downloaded from the PDB (https://www.rcsb.org/) with PDB ID 6OOD. Moreover, the datasets generated and/or analysed during the current study are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
Code availability
All code used in this study is available from the corresponding authors upon reasonable request.
References
Wencel-Delord, J., Panossian, A., Leroux, F. R. & Colobert, F. Recent advances and new concepts for the synthesis of axially stereoenriched biaryls. Chem. Soc. Rev. 44, 3418–3430 (2015).
Blackmond, D. G. The origin of biological homochirality. Cold Spring Harb. Perspect. Biol. 2, a002147 (2010).
Pizzolato, S. F. et al. Central-to-helical-to-axial-to-central transfer of chirality with a photoresponsive catalyst. J. Am. Chem. Soc. 140, 17278–17289 (2018).
Toenjes, S. T. & Gustafson, J. L. Atropisomerism in medicinal chemistry: challenges and opportunities. Future Med. Chem. 10, 409–422 (2018).
Evans, D. A. et al. Approaches to the synthesis of the vancomycin antibiotics: synthesis of orienticin C (bis-dechlorovancomycin) aglycon. J. Am. Chem. Soc. 119, 3419–3420 (1997).
McMahon, J. B. et al. Michellamine B, a novel plant alkaloid, inhibits human immunodeficiency virus-induced cell killing by at least two distinct mechanisms. Antimicrob. Agents Chemother. 39, 484–488 (1995).
Gesell, A. et al. CYP719B1 is salutaridine synthase, the C–C phenol-coupling enzyme of morphine biosynthesis in opium poppy. J. Biol. Chem. 284, 24432–24442 (2009).
Gil Girol, C. et al. Regio- and stereoselective oxidative phenol coupling in Aspergillus niger. Angew. Chem. Int. Ed. Engl. 51, 9788–9791 (2012).
Prag, A. et al. Regio- and stereoselective intermolecular oxidative phenol coupling in Streptomyces. J. Am. Chem. Soc. 136, 6195–6198 (2014).
Fang, W. et al. Naphthol radical couplings determine structural features and enantiomeric excess of dalesconols in Daldinia eschscholzii. Nat. Commun. 3, 1039 (2012).
Davin, L. B. et al. Stereoselective bimolecular phenoxy radical coupling by an auxiliary (dirigent) protein without an active center. Science 275, 362–366 (1997).
Effenberger, I. et al. Dirigent proteins from cotton (Gossypium sp.) for the atropselective synthesis of gossypol. Angew. Chem. Int. Ed. Engl. 54, 14660–14663 (2015).
Yu, Y. W. Probing into the mechanism of action, metabolism and toxicity of gossypol by studying its (+)- and (−)-stereoisomers. J. Ethnopharmacol. 20, 65–78 (1987).
Liu, Y. X., Wang, L. L., Zhao, L. & Zhang, Y. G. Structure, properties of gossypol and its derivatives—from physiological activities to drug discovery and drug design. Nat. Prod. Rep. 39, 1282–1304 (2022).
Liu, H. et al. Double-layered hyaluronic acid/stearic acid-modified polyethyleneimine nanoparticles encapsulating (−)-gossypol: a nanocarrier for chiral anticancer drugs. J. Mater. Chem. B 2, 5238–5248 (2014).
Matlin, S. A. et al. (−)-Gossypol: an active male antifertility agent. Contraception 31, 141–149 (1985).
Wang, N. G., Zhou, L. F., Guan, M. H. & Lei, H. P. Effect of (−)- and (+)-gossypol on fertility in male rats. J. Ethnopharmacol. 20, 21–24 (1987).
Benz, C. C. et al. Biochemical correlates of the antitumor and antimitochondrial properties of gossypol enantiomers. Mol. Pharmacol. 37, 840–847 (1990).
Gershenzon, J. & Dudareva, N. The function of terpene natural products in the natural world. Nat. Chem. Biol. 3, 408–414 (2007).
Nour-Eldin, H. H. et al. Reduction of antinutritional glucosinolates in Brassica oilseeds by mutation of genes encoding transporters. Nat. Biotechnol. 35, 377–382 (2017).
Bjornsdotter, E. et al. VC1 catalyses a key step in the biosynthesis of vicine in faba bean. Nat. Plants 7, 923–931 (2021).
Li, J. et al. Biofortified tomatoes provide a new route to vitamin D sufficiency. Nat. Plants 8, 611–616 (2022).
Sunilkumar, G., Campbell, L. M., Puckhaber, L., Stipanovic, R. D. & Rathore, K. S. Engineering cottonseed for use in human nutrition by tissue-specific reduction of toxic gossypol. Proc. Natl Acad. Sci. USA 103, 18054–18059 (2006).
Waltz, E. First edible cottonseed go-ahead. Nat. Biotechnol. 36, 1126 (2018).
Townsend, B. J. & Llewellyn, D. J. Reduced terpene levels in cottonseed add food to fiber. Trends Biotechnol. 25, 239–241 (2007).
Stipanovic, R. D., Puckhaber, L. S., Bell, A. A., Percival, A. E. & Jacobs, J. Occurrence of (+)- and (−)-gossypol in wild species of cotton and in Gossypium hirsutum var. marie-galante (Watt) Hutchinson. J. Agric. Food Chem. 53, 6266–6271 (2005).
Jaroszewski, J. W., Strom-Hansen, T., Hansen, S. H., Thastrup, O. & Kofod, H. On the botanical distribution of chiral forms of gossypol. Planta Med. 58, 454–458 (1992).
Tian, X. et al. Characterization of gossypol biosynthetic pathway. Proc. Natl Acad. Sci. USA 115, E5410–E5418 (2018).
Huang, J. Q. et al. Aromatization of natural products by a specialized detoxification enzyme. Nat. Chem. Biol. 16, 250–256 (2020).
Ma, D. et al. Genetic basis for glandular trichome formation in cotton. Nat. Commun. 7, 10456 (2016).
Wang, J. Y. et al. VdNEP, an elicitor from Verticillium dahliae, induces cotton plant wilting. Appl. Environ. Microbiol. 70, 4989–4995 (2004).
Uchida, K., Akashi, T. & Aoki, T. The missing link in leguminous pterocarpan biosynthesis is a dirigent domain-containing protein with isoflavanol dehydratase activity. Plant Cell Physiol. 58, 398–408 (2017).
Fryxell, P. A. A redefinition of tribe Gossypieae. Bot. Gaz. 129, 296–308 (1968).
Park, S. H., Scheffler, J., Scheffler, B., Cantrell, C. L. & Pauli, C. S. Chemical defense responses of upland cotton, Gossypium hirsutum L. to physical wounding. Plant Direct 3, e00141 (2019).
Bezemer, T. M., Wagenaar, I. R., van Dam, N. M., van der Putten, W. H. & Wackers, F. L. Above- and below-ground terpenoid aldehyde induction in cotton, Gossypium herbaceum, following root and leaf injury. J. Chem. Ecol. 30, 53–67 (2004).
Opitz, S., Kunert, G. & Gershenzon, J. Increased terpenoid accumulation in cotton (Gossypium hirsutum) foliage is a general wound response. J. Chem. Ecol. 34, 508–522 (2008).
Hedin, P. A., Parrott, W. L. & Jenkins, J. N. Effects of cotton plant allelochemicals and nutrients on behavior and development of tobacco budworm. J. Chem. Ecol. 17, 1107–1121 (1991).
Lusas, E. W. & Jividen, G. M. Glandless cottonseed—a review of the 1st 25 years of processing and utilization research. J. Am. Oil Chem. Soc. 64, 839–854 (1987).
Rathore, K. S. et al. Ultra-low gossypol cottonseed: selective gene silencing opens up a vast resource of plant-based protein to improve human nutrition. CRC Crit. Rev. Plant Sci. 39, 1–29 (2020).
Pösö, H., Wichmann, K., Jänne, J. & Luukkainen, T. Gossypol, a powerful inhibitor of human spermatozoal metabolism. Lancet 1, 885–886 (1980).
de Peyster, A. & Wang, Y. Y. Gossypol—proposed contraceptive for men passes the Ames test. N. Engl. J. Med. 301, 275–276 (1979).
Kitada, S. et al. Bcl-2 antagonist apogossypol (NSC736630) displays single-agent activity in Bcl-2–transgenic mice and has superior efficacy with less toxicity compared with gossypol (NSC19048). Blood 111, 3211–3219 (2008).
Balakrishnan, K., Burger, J. A., Wierda, W. G. & Gandhi, V. AT-101 induces apoptosis in CLL B cells and overcomes stromal cell-mediated Mcl-1 induction and drug resistance. Blood 113, 149–153 (2009).
Haasler, L. et al. The BH3 mimetic (+/−) gossypol induces ROS-independent apoptosis and mitochondrial dysfunction in human A375 melanoma cells in vitro. Arch. Toxicol. 95, 1349–1365 (2021).
Huang, G., Huang, J. Q., Chen, X. Y. & Zhu, Y. X. Recent advances and future perspectives in cotton research. Annu. Rev. Plant Biol. 72, 437–462 (2021).
O’Leary, B. M., Rico, A., McCraw, S., Fones, H. N. & Preston, G. M. The infiltration–centrifugation technique for extraction of apoplastic fluid from plant leaves using Phaseolus vulgaris as an example. J. Vis. Exp. 94, e52113 (2014).
Kim, S. S. & Sattely, E. S. Dirigent proteins guide asymmetric heterocoupling for the synthesis of complex natural product analogues. J. Am. Chem. Soc. 143, 5011–5021 (2021).
Meng, Q. et al. Pterocarpan synthase (PTS) structures suggest a common quinone methide-stabilizing function in dirigent proteins and proteins with dirigent-like domains. J. Biol. Chem. 295, 11584–11601 (2020).
Gasper, R. et al. Dirigent protein mode of action revealed by the crystal structure of AtDIR6. Plant Physiol. 172, 2165–2175 (2016).
Kim, K. W. et al. Trimeric structure of (+)-pinoresinol-forming dirigent protein at 1.95 A resolution with three isolated active sites. J. Biol. Chem. 290, 1308–1318 (2015).
Ziegler, J. & Facchini, P. J. Alkaloid biosynthesis: metabolism and trafficking. Annu. Rev. Plant Biol. 59, 735–769 (2008).
Yonekura-Sakakibara, K. et al. Seed-coat protective neolignans are produced by the dirigent protein AtDP1 and the laccase AtLAC5 in Arabidopsis. Plant Cell 33, 129–152 (2021).
Hosmani, P. S. et al. Dirigent domain-containing protein is part of the machinery required for formation of the lignin-based Casparian strip in the root. Proc. Natl Acad. Sci. USA 110, 14498–14503 (2013).
Aghova, T. et al. Fossils know it best: using a new set of fossil calibrations to improve the temporal phylogenetic framework of murid rodents (Rodentia: Muridae). Mol. Phylogenet. Evol. 128, 98–111 (2018).
Gao, X., Britt, R. C.Jr, Shan, L. & He, P. Agrobacterium-mediated virus-induced gene silencing assay in cotton. J. Vis. Exp. 20, 2938 (2011).
Shan, C. M. et al. Control of cotton fibre elongation by a homeodomain transcription factor GhHOX3. Nat. Commun. 5, 5519 (2014).
Ma, X. et al. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol. Plant 8, 1274–1284 (2015).
Burton, R. A. et al. Cellulose synthase-like CslF genes mediate the synthesis of cell wall (1,3;1,4)-beta-d-glucans. Science 311, 1940–1942 (2006).
Jefferson, R. A., Kavanagh, T. A. & Bevan, M. W. GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 6, 3901–3907 (1987).
Liu, M. L., Cheng, Y. M. & Jia, M. C. LM23 is essential for spermatogenesis in Rattus norvegicus. Front. Biosci. (Elite Ed.) 2, 187–194 (2010).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46, W296–W303 (2018).
Morris, G. M. et al. AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J. Comput. Chem. 30, 2785–2791 (2009).
Cousins, K. R. ChemDraw Ultra 9.0. CambridgeSoft, 100 CambridgePark Drive, Cambridge, MA 02140. www.cambridgesoft.com. See Web site for pricing options. J. Am. Chem. Soc. 127, 4115–4116 (2005).
Kumar, S., Tamura, K. & Nei, M. MEGA: Molecular evolutionary genetics analysis software for microcomputers. Comput. Appl. Biosci. 10, 189–191 (1994).
Seifert, E. OriginPro 9.1: Scientific data analysis and graphing software—software review. J. Chem. Inf. Model. 54, 1552 (2014).
Motulsky, H. J. Analyzing Data with GraphPad Prism, 1999, GraphPad Software Inc., San Diego CA, www.graphpad.com
Wendel, J. F. & Grover, C. E. Taxonomy and evolution of the cotton genus, Gossypium. Cotton 57, 25–44 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Pickel, B. et al. An enantiocomplementary dirigent protein for the enantioselective laccase-catalyzed oxidative coupling of phenols. Angew. Chem. Int. Ed. Engl. 49, 202–204 (2010).
Shi, H. et al. Overexpression of cotton (Gossypium hirsutum) dirigent1 gene enhances lignification that blocks the spread of Verticillium dahliae. Acta Biochim. Biophys. Sin. (Shanghai) 44, 555–564 (2012).
Effenberger, I., Harport, M., Pfannstiel, J., Klaiber, I. & Schaller, A. Expression in Pichia pastoris and characterization of two novel dirigent proteins for atropselective formation of gossypol. Appl. Microbiol. Biotechnol. 101, 2021–2032 (2017).
Teufel, F. et al. SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat. Biotechnol. 40, 1023–1025 (2022).
Acknowledgements
We thank W. Hu, X. Xu, S. Wang and X. Liao for help with the HPLC–MS analyses and S. Zhan, X. Wang, Y. Li, C. Li, B. Yang, P. Chen and L. Chen for their kind and generous help. The research was supported by grants from the National Key R&D Program of China (no. 2022YFF1001400) to J.-Q.H., the Chinese Academy of Sciences (no. XDB27020207) to X.-Y.C., the National Natural Science Foundation of China (nos. 31788103 and 31690092) to X.-Y.C., the Foundation of Youth Innovation Promotion Association of the Chinese Academy of Sciences to J.-Q.H., the Young Elite Scientists Sponsorship Program by CAST (no. 2019QNRC001) to J.-Q.H., the Hainan Yazhou Bay Seed Laboratory (no. B21HJ0223) to X.-Y.C. and the Yunnan Revitalization Talent Support Program ‘Top Team’ Project to X.-Y.C. We also acknowledge the support of the SANOFI Scholarship Program (to J.-Q.H.). Leaves of diploid or allotetraploid Gossypium species and Gossypioides kirkii were acquired from the Sanya National Research Station of Wild Cotton Germplasm, China.
Author information
Authors and Affiliations
Contributions
J.-Q.H., J.-L.L. and X.-Y.C. conceived the idea and designed the study. C.M., X.F., B.X., N.-J.L. and L.-J.W. discussed the results and provided advice. J.-L.L. and J.-Q.H. isolated the DIRs and characterized their function. J.-Q.H., J.-L.L. and L.-J.W. generated the transgenic plants. J.-L.L., W.-K.W. and F.-Y.C. completed the LC–MS analysis. J.-Q.H., J.-L.L., X.-X.G. and J.-F.H. evaluated the resistance and toxicity of the transgenic plants. J.-L.L., J.-Q.H. and Z.-W.C. carried out the bioinformatic analysis. J.-X.L. performed the protein modelling. J.-Q.H., X.-Y.C., J.-L.L. and C.M. wrote, reviewed and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Reuben Peters, Xiaoquan QI and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
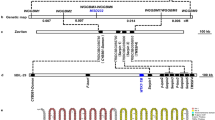
Extended Data Fig. 1 Transcriptome analysis to identify the (−)-gossypol biosynthetic gene in upland cotton.
a, Scatter plot, constructed from 35 RNA-seq samples, illustrates the correlation values of the downregulated genes in glandless cotton (edgeR, log2[FC] < = −8, log2[FC]: log2-transformed fold change) with CYP71BE79. Differential RNA expression of genes was analyzed in R software using edgeR68. Point sizes indicate the FDR-adjusted P-value of differential expression. FDR-adjusted P values were calculated using Benjamini-Hochberg correction of the obtained P values. Different colors distinguish between protein families. Correlation analysis was measured by Pearson’s correlation coefficient with ‘Cor’ function from R software. b, Sequence alignment of Gh_A04G0066 (GhDIR5) and Gh_A05G2499 (GhDIR6) conducted by BioEdit software 7.2.5. c, Global heatmap showing the abundance of gossypol in different organs or in ovules at different stages. d, Global heatmap showing the expressions of CYP71BE79, Gh_A04G0066 (GhDIR5) and Gh_A05G2499 (GhDIR6) in different organs or in ovules at different stages. For c and d: DPA, days post anthesis. ROT: root; STE, stem; LEA, leaf; PET, petal; PIS, pistil; CAL, calycle. e, Relative expressions of Gh_A04G0066 (GhDIR5) and Gh_A05G2499 (GhDIR6) in leaf and ovule (35 DPA) of the wild-type (gland, G) and gossypol-deficient (glandless, GL) cultivars, in cotyledons two days post-elicitor treatment, and in GoPGF-silencing leaves. GoPGF is the regulator of pigment gland formation. FPKM (fragments per kilobase million) values obtained from the gland cotton leaves or ovules and the elicitor-treated cotyledons were set to 1 and used to estimate relative values of the paired samples. Data shown are means ± s. d. (n = 3 biological replicates).
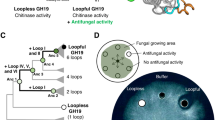
Extended Data Fig. 2 Phylogenetic analysis of the DIR family proteins from upland cotton (n = 121), rice (n = 49), and Arabidopsis thaliana (n = 26).
Proteins whose function has been described in the literature are indicated. [AtDIR6 and AtDIR5, (−)-pinoresinol-forming DIRs from A. thaliana69, in the DIR-a subfamily; AtDP1/AtDIR12, a neolignan biosynthesis DIR from A. thaliana52, in the DIR-a subfamily; AtDIR10 (ESB1), a Casparian band lignin-forming DIR from A. thaliana53, in the DIR-e subfamily; GhDIR1, a DIR involved in cotton lignification which could limit the spread of fungal pathogen Verticillium dahlia70, in the DIR-b/d subfamily]. GhDIR5, GhDIR6 and the previously reported GhDIR412 and GhDIR371 (another Gossypium hirsutum DIR for (+)-2 formation) are classified to the DIR-b/d subfamily, in which GhDIR6 belongs to a branch separated earlier.
Extended Data Fig. 3 GhDIR5 and GhDIR6 were computationally predicted to contain N-terminal signal peptide.
a,b, Signal peptides were predicted using the SignalP-6.0 (https://services.healthtech.dtu.dk/service.php?SignalP)72.
Extended Data Fig. 4 Subcellular localization of GhDIR5 and GhDIR6.
a, Confocal microscopy (Leica TSC SP8 STED 3X) of pGhDIR5::GhDIR5-EGFP and pGhDIR6::GhDIR6-EGFP transgenic cotton leaves. Scale bars, 30 μm. b, GUS staining of pGhDIR5::GUS and pGhDIR6::GUS transgenic cotton leaves. Scale bars, 50 μm. c, Immunoelectron microscopy with monoclonal antibodies against GhDIR5 and gold-labelled secondary antibodies, showing the localization of GhDIR5 in pigmented glands. The magnification of the box is shown in the panel below. Arrows indicate the immunogold particles in cotton pigmented glands. Scale bars of original and amplified panels are 10 μm and 500 nm, respectively. For controls the primary antibodies were replaced with the non-immune serum. All experiments were repeated independently at least two times with similar results, representative images are shown.
Extended Data Fig. 5 Sequence similarity network analysis of DIRs.
Protein sequence similarity network (SSN) of DIRs from eight Gossypium species and from Gossypioides kirkii. Each node within the resulting SSN contains sequences with >93% amino acid identity, rendering different DIRs into different clusters. Each color denotes a plant species. Protein sequences were downloaded from CottonGen (https://www.cottongen.org/). The SSN was constructed by EFI-EST webtool (http.//efi.igb.illinois.edu/efi-est/) and visualized with Cytoscape 3.9.1.
Extended Data Fig. 6 GhDIR5 and GhDIR6 expression and activity assays.
a, Western blot of purified GhDIR5 and GhDIR6 proteins extracted from agroinfiltrated leaves of N. benthamiana and affinity purified with Ni sepharose column. Mouse anti-His antibody (TA-02, ORIGENE) was used at a dilution of 1:5000 using goat-anti-mouse IgG horseradish peroxidase conjugate (1:10000, ZB2305, ORIGENE) as secondary antibody. Molecular weight of selected bands was indicated in kDa. The experiments were repeated independently three times with similar results. b, Enantiomeric ratio of (±)-2 resulted from oxidative coupling of 1, the reaction was incubated with commercialized RvLac and GhDIR5 or GhDIR6, as indicated. Reaction without DIR was used as control. (Data are means ± s. d., n = 3 independent experiments). c, Production of total 2 increased with the amounts of GhDIR5 (0.5–5 μg in 100 μL), RvLac was used as the oxidative enzyme. 2 formed in the absence of GhDIR5 was set to 1 (means ± s. d., n = 2 or 3 independent experiments).
Extended Data Fig. 7 VIGS analysis of GhDIR5.
a–e, Relative contents of hemigossypolone and heliocides in the virus-mediated GhDIR5-silencing leaves of G. hirsutum. Value of the empty tobacco rattle virus (TRV) vector control (TRV:00) was set to 1 (means ± s. d., n = 5 independent experiments), hereinafter inclusive. f, g, Relative expression levels of GhDIR5 (f) and GhDIR4 (g) in the TRV:GhDIR5 leaves, compared to the EV control (means ± s. d., n = 3 independent experiments).
Extended Data Fig. 8 Identification of CRISPR/Cas9–edited GhDIR5 knockout lines.
a, Western blot detection of leaf proteins after SDS-PAGE. GhDIR5 antibody (Abmart) was used at a dilution of 1:1000, using goat-anti-mouse IgG horseradish peroxidase conjugate (1:10000, ZB2305, ORIGENE) as secondary antibody. The experiments were repeated independently three times with similar results. b-c, CRISPR/Cas9-introduced GhDIR5 mutation didn’t led to off-target mutations at the GhDIR4 (b) and GhDIR6 (c) loci, as determined by Sanger sequencing after PCR amplification.
Extended Data Fig. 9 Content of phytoalexins in CR-Ghdir5 lines.
a-d, Relative contents of (±)-2 in WT and CR-Ghdir5 cotton plant organs. e-n, Relative contents of hemigossypolone and heliocides in WT and CR-Ghdir5 cotton leaf (e-i) and sepal (j-n). Content value of the WT cotton was set to 1 (means ± s. d., n = 3 independent experiments).
Extended Data Fig. 10 Representative HE-stained semi-thin sections of testes of the control and the cottonseed extracts-treated mice.
The seed extracts contained 1 mM racemic 2 (WT) or (+)-2 (CR-Ghdir5) from the WT and CR-Ghdir5 seeds, respectively. The mock group was treated with DMSO. Scale bars are indicated in the right. The experiments were repeated independently three times with similar results.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6, Tables 1 and 2, and source data for Supplementary Fig. 5.
Supplementary Data
Statistical source data.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed western blots.
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lin, JL., Fang, X., Li, JX. et al. Dirigent gene editing of gossypol enantiomers for toxicity-depleted cotton seeds. Nat. Plants 9, 605–615 (2023). https://doi.org/10.1038/s41477-023-01376-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-023-01376-2
This article is cited by
-
Targeted genome editing for cotton improvement: prospects and challenges
The Nucleus (2024)
-
The resilient cotton plant: uncovering the effects of stresses on secondary metabolomics and its underlying molecular mechanisms
Functional & Integrative Genomics (2023)