Abstract
Appropriate root system architecture (RSA) can improve maize yields in densely planted fields, but little is known about its genetic basis in maize. Here we performed root phenotyping of 14,301 field-grown plants from an association mapping panel to study the genetic architecture of maize RSA. A genome-wide association study identified 81 high-confidence RSA-associated candidate genes and revealed that 28 (24.3%) of known root-related genes were selected during maize domestication and improvement. We found that modern maize breeding has selected for a steeply angled root system. Favourable alleles related to steep root system angle have continuously accumulated over the course of modern breeding, and our data pinpoint the root-related genes that have been selected in different breeding eras. We confirm that two auxin-related genes, ZmRSA3.1 and ZmRSA3.2, contribute to the regulation of root angle and depth in maize. Our genome-wide identification of RSA-associated genes provides new strategies and genetic resources for breeding maize suitable for high-density planting.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Data supporting the findings of this work are available within the paper and its Supplementary Information. The genotype set, population structure and kinship data can be downloaded from the Maizego website (http://www.maizego.org/Resources.html). All root phenotype data for the 380 inbred maize lines are included in Supplementary Table 25. The RNA-sequencing reads used to construct the co-expression network and the root transcriptome sequencing reads were deposited in the NCBI Sequence Read Archive (https://www.ncbi.nlm.nih.gov/) under accession codes PRJNA694491 and PRJNA693427, respectively. Source data are provided with this paper.
Code availability
All scripts for GWAS, co-expression network analysis, selective sweep detection for domestication and improvement, and obtaining aligned sequences of high-priority candidate genes and known root-related genes (https://doi.org/10.5281/zenodo.7112683) are available on Zenodo.
References
Tilman, D., Balzer, C., Hill, J. & Befort, B. L. Global food demand and the sustainable intensification of agriculture. Proc. Natl Acad. Sci. USA 108, 20260–20264 (2011).
Duvick, D. N. The contribution of breeding to yield advances in maize (Zea mays L.). Adv. Agron. 86, 83–145 (2005).
Tian, J. et al. Teosinte ligule allele narrows plant architecture and enhances high-density maize yields. Science 365, 658–664 (2019).
Wang, B. et al. Genome-wide selection and genetic improvement during modern maize breeding. Nat. Genet. 52, 565–571 (2020).
Hochholdinger, F. Untapping root system architecture for crop improvement. J. Exp. Bot. 67, 4431–4433 (2016).
Lynch, J. P. Root phenotypes for improved nutrient capture: an underexploited opportunity for global agriculture. N. Phytol. 223, 548–564 (2019).
Lynch, J. P. Steep, cheap and deep: an ideotype to optimize water and N acquisition by maize root systems. Ann. Bot. 112, 347–357 (2013).
Mi, G., Chen, F., Yuan, L. & Zhang, F. Ideotype root system architecture for maize to achieve high yield and resource use efficiency in intensive cropping systems. Adv. Agron. 139, 73–97 (2016).
Thorup-Kristensen, K. et al. Digging deeper for agricultural resources, the value of deep rooting. Trends Plant Sci. 25, 406–417 (2020).
Shao, H. et al. Genotypic difference in the plasticity of root system architecture of field-grown maize in response to plant density. Plant Soil 439, 201–217 (2019).
Gamuyao, R. et al. The protein kinase Pstol1 from traditional rice confers tolerance of phosphorus deficiency. Nature 488, 535–539 (2012).
Uga, Y. et al. Control of root system architecture by DEEPER ROOTING 1 increases rice yield under drought conditions. Nat. Genet. 45, 1097–1102 (2013).
Kirschner, G. K. et al. ENHANCED GRAVITROPISM 2 encodes a STERILE ALPHA MOTIF-containing protein that controls root growth angle in barley and wheat. Proc. Natl Acad. Sci. USA 118, e2101526118 (2021).
Bray, A. L. & Topp, C. N. The quantitative genetic control of root architecture in maize. Plant Cell Physiol. 59, 1919–1930 (2018).
Hochholdinger, F., Yu, P. & Marcon, C. Genetic control of root system development in maize. Trends Plant Sci. 23, 79–88 (2018).
Zhang, X. M. et al. Genetic variation in ZmTIP1 contributes to root hair elongation and drought tolerance in maize. Plant Biotechnol. J. 18, 1271–1283 (2020).
Schneider, H. M. et al. Root angle in maize influences nitrogen capture and is regulated by calcineurin B-like protein (CBL)-interacting serine/threonine-protein kinase 15 (ZmCIPK15). Plant Cell Environ. 45, 837–853 (2022).
Schneider, H. M. et al. Genetic control of root architectural plasticity in maize. J. Exp. Bot. 71, 3185–3197 (2020).
Zheng, Z. et al. Shared genetic control of root system architecture between Zea mays and Sorghum bicolor. Plant Physiol. 182, 977–991 (2020).
Chen, Z. et al. Plasticity of root anatomy during domestication of a maize-teosinte derived population. J. Exp. Bot. 73, 139–153 (2022).
Burton, A. L., Brown, K. M. & Lynch, J. P. Phenotypic diversity of root anatomical and architectural traits in Zea species. Crop Sci. 53, 1042–1055 (2013).
Gaudin, A. C. M., McClymont, S. A., Soliman, S. S. M. & Raizada, M. N. The effect of altered dosage of a mutant allele of Teosinte branched 1 (tb1-ref) on the root system of modern maize. BMC Genet. 15, 23 (2014).
Perkins, A. C. & Lynch, J. P. Increased seminal root number associated with domestication improves nitrogen and phosphorus acquisition in maize seedlings. Ann. Bot. 128, 453–468 (2021).
Wang, H. et al. Natural variation and domestication selection of ZmCKX5 with root morphological traits at the seedling stage in maize. Plants 10, 1 (2021).
Gaudin, A. C. M., McClymont, S. A. & Raizada, M. N. The nitrogen adaptation strategy of the wild teosinte ancestor of modern maize, Zea mays subsp. parviglumis. Crop Sci. 51, 2780–2795 (2011).
Gao, K., Chen, F., Yuan, L., Zhang, F. & Mi, G. A comprehensive analysis of root morphological changes and nitrogen allocation in maize in response to low nitrogen stress. Plant Cell Environ. 38, 740–750 (2015).
Mano, Y. et al. Variation for root aerenchyma formation in flooded and non-flooded maize and teosinte seedlings. Plant Soil 281, 269–279 (2006).
Ning, P., Li, S., Li, X. & Li, C. New maize hybrids had larger and deeper post-silking root than old ones. Field Crop Res. 166, 66–71 (2014).
Chen, X. et al. Changes in root size and distribution in relation to nitrogen accumulation during maize breeding in China. Plant Soil 374, 121–130 (2014).
Mano, Y. & Omori, F. Flooding tolerance in interspecific introgression lines containing chromosome segments from teosinte (Zea nicaraguensis) in maize (Zea mays subsp. mays). Ann. Bot. 112, 1125–1139 (2013).
Wang, H., Studer, A. J., Zhao, Q., Meeley, R. & Doebley, J. F. Evidence that the origin of naked kernels during maize domestication was caused by a single amino acid substitution in tga1. Genetics 200, 965–974 (2015).
Li, Z. et al. Enhancing auxin accumulation in maize root tips improves root growth and dwarfs plant height. Plant Biotechnol. J. 16, 86–99 (2018).
Zhao, Y. et al. The interaction between rice ERF3 and WOX11 promotes crown root development by regulating gene expression involved in cytokinin signaling. Plant Cell 27, 2469–2483 (2015).
Asim, M., Ullah, Z., Oluwaseun, A., Wang, Q. & Liu, H. B. Signalling overlaps between nitrate and auxin in regulation of the root system architecture: insights from the Arabidopsis thaliana. Int. J. Mol. Sci. 21, 2880 (2020).
Woll, K. et al. Isolation, characterization, and pericycle-specific transcriptome analyses of the novel maize lateral and seminal root initiation mutant rum1. Plant Physiol. 139, 1255–1267 (2005).
von Behrens, I. et al. Rootless with undetectable meristem 1 encodes a monocot-specific AUX/IAA protein that controls embryonic seminal and post-embryonic lateral root initiation in maize. Plant J. 66, 341–353 (2011).
Tang, J. et al. Genetic dissection of plant height by molecular markers using a population of recombinant inbred lines in maize. Euphytica 155, 117–124 (2007).
Li, H. et al. Genome-wide association study dissects the genetic architecture of oil biosynthesis in maize kernels. Nat. Genet. 45, 43–50 (2013).
Wen, T. J., Hochholdinger, F., Sauer, M., Bruce, W. & Schnable, P. S. The roothairless1 gene of maize encodes a homolog of sec3, which is involved in polar exocytosis. Plant Physiol. 138, 1637–1643 (2005).
Huang, P. et al. Sparse panicle1 is required for inflorescence development in Setaria viridis and maize. Nat. Plants 3, 17054 (2017).
Rademacher, E. H. et al. Different auxin response machineries control distinct cell fates in the early plant embryo. Dev. Cell 22, 211–222 (2012).
Rinaldi, M. A., Liu, J., Enders, T. A., Bartel, B. & Strader, L. C. A gain-of-function mutation in IAA16 confers reduced responses to auxin and abscisic acid and impedes plant growth and fertility. Plant Mol. Biol. 79, 359–373 (2012).
Huang, J. et al. Formin homology 1 (OsFH1) regulates root-hair elongation in rice (Oryza sativa). Planta 237, 1227–1239 (2013).
Kitomi, Y., Inahashi, H., Takehisa, H., Sato, Y. & Inukai, Y. OsIAA13-mediated auxin signaling is involved in lateral root initiation in rice. Plant Sci. 190, 116–122 (2012).
Chen, H., Patterson, N. & Reich, D. Population differentiation as a test for selective sweeps. Genome Res. 20, 393–402 (2010).
Hufford, M. B. et al. Comparative population genomics of maize domestication and improvement. Nat. Genet. 44, 808–811 (2012).
Chen, Q. et al. The genetic architecture of the maize progenitor, teosinte, and how it was altered during maize domestication. PLoS Genet. 16, e1008791 (2020).
Wang, Y., Deng, D., Bian, Y., Lv, Y. & Xie, Q. Genome-wide analysis of primary auxin-responsive Aux/IAA gene family in maize (Zea mays. L.). Mol. Biol. Rep. 37, 3991–4001 (2010).
Rosero, A. et al. Arabidopsis FH1 formin affects cotyledon pavement cell shape by modulating cytoskeleton dynamics. Plant Cell Physiol. 57, 488–504 (2016).
Shi, J. et al. Ectopic expression of ARGOS8 reveals a role for ethylene in root-lodging resistance in maize. Plant J. 97, 378–390 (2019).
Evans, M. M. & Poethig, R. S. Gibberellins promote vegetative phase change and reproductive maturity in maize. Plant Physiol. 108, 475–487 (1995).
Trachsel, S., Kaeppler, S. M., Brown, K. M. & Lynch, J. P. Maize root growth angles become steeper under low N conditions. Field Crop Res 140, 18–31 (2013).
Kell, D. B. Breeding crop plants with deep roots: their role in sustainable carbon, nutrient and water sequestration. Ann. Bot. 108, 407–418 (2011).
Hund, A., Reime, R. & Messmer, R. A consensus map of QTLs controlling the root length of maize. Plant Soil 344, 143–158 (2011).
Schaefer, R. J. et al. Integrating coexpression networks with GWAS to prioritize causal genes in maize. Plant Cell 30, 2922–2942 (2018).
York, L. M., Galindo-Castaneda, T., Schussler, J. R. & Lynch, J. P. Evolution of US maize (Zea mays L.) root architectural and anatomical phenes over the past 100 years corresponds to increased tolerance of nitrogen stress. J. Exp. Bot. 66, 2347–2358 (2015).
Chen, F. et al. Breeding for high-yield and nitrogen use efficiency in maize: lessons from comparison between Chinese and US cultivars. Adv. Agron. 166, 251–275 (2021).
Xu, C. et al. Cooperative action of the paralogous maize lateral organ boundaries (LOB) domain proteins RTCS and RTCL in shoot-borne root formation. N. Phytol. 207, 1123–1133 (2015).
Suzuki, M., Sato, Y., Wu, S., Kang, B. H. & McCarty, D. R. Conserved functions of the MATE transporter BIG EMBRYO1 in regulation of lateral organ size and initiation rate. Plant Cell 27, 2288–2300 (2015).
Zhang, Y. et al. LATERAL ROOT PRIMORDIA 1 of maize acts as a transcriptional activator in auxin signalling downstream of the Aux/IAA gene rootless with undetectable meristem 1. J. Exp. Bot. 66, 3855–3863 (2015).
Taramino, G. et al. The maize (Zea mays L.) RTCS gene encodes a LOB domain protein that is a key regulator of embryonic seminal and post-embryonic shoot-borne root initiation. Plant J. 50, 649–659 (2007).
Zhang, M. et al. Auxin efflux carrier ZmPGP1 mediates root growth inhibition under aluminum stress. Plant Physiol. 177, 819–832 (2018).
Benjamins, R. & Scheres, B. Auxin: the looping star in plant development. Annu Rev. Plant Biol. 59, 443–465 (2008).
Ikeda, Y. et al. Local auxin biosynthesis modulates gradient-directed planar polarity in Arabidopsis. Nat. Cell Biol. 11, 731–738 (2009).
Lanza, M. et al. Role of actin cytoskeleton in brassinosteroid signaling and in its integration with the auxin response in plants. Dev. Cell 22, 1275–1285 (2012).
Banno, H. & Chua, N. H. Characterization of the arabidopsis formin-like protein AFH1 and its interacting protein. Plant Cell Physiol. 41, 617–626 (2000).
Martiniere, A., Gayral, P., Hawes, C. & Runions, J. Building bridges: formin1 of Arabidopsis forms a connection between the cell wall and the actin cytoskeleton. Plant J. 66, 354–365 (2011).
Li, G. et al. Rice actin-binding protein RMD is a key link in the auxin–actin regulatory loop that controls cell growth. Proc. Natl Acad. Sci. USA 111, 10377–10382 (2014).
Ogura, T. et al. Root system depth in Arabidopsis Is shaped by EXOCYST70A3 via the dynamic modulation of auxin transport. Cell 178, 400–412 (2019).
Yang, P. et al. Light modulates the gravitropic responses through organ-specific PIFs and HY5 regulation of LAZY4 expression in Arabidopsis. Proc. Natl Acad. Sci. USA 117, 18840–18848 (2020).
Yang, X. et al. Characterization of a global germplasm collection and its potential utilization for analysis of complex quantitative traits in maize. Mol. Breed. 28, 511–526 (2011).
Trachsel, S., Kaeppler, S. M., Brown, K. M. & Lynch, J. P. Shovelomics: high throughput phenotyping of maize (Zea mays L.) root architecture in the field. Plant Soil 341, 75–87 (2010).
Das, A. et al. Digital imaging of root traits (DIRT): a high-throughput computing and collaboration platform for field-based root phenomics. Plant Methods 11, 51 (2015).
Colombi, T. et al. Next generation shovelomics: set up a tent and REST. Plant Soil 388, 1–20 (2015).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48 (2015).
Fu, J. et al. RNA sequencing reveals the complex regulatory network in the maize kernel. Nat. Commun. 4, 2832 (2013).
Unterseer, S. et al. A powerful tool for genome analysis in maize: development and evaluation of the high density 600 k SNP genotyping array. BMC Genomics 15, 823 (2014).
Ganal, M. W. et al. A large maize (Zea mays L.) SNP genotyping array: development and germplasm genotyping, and genetic mapping to compare with the B73 reference genome. PLoS ONE 6, e28334 (2011).
Elshire, R. J. et al. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. PLoS ONE 6, e19379 (2011).
Liu, H. J. et al. MODEM: multi-omics data envelopment and mining in maize. Database 2016, baw117 (2016).
Yu, J. et al. A unified mixed-model method for association mapping that accounts for multiple levels of relatedness. Nat. Genet. 38, 203–208 (2006).
Zhang, Z. et al. Mixed linear model approach adapted for genome-wide association studies. Nat. Genet. 42, 355–360 (2010).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet 81, 559–575 (2007).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Maere, S., Heymans, K. & Kuiper, M. BiNGO: a Cytoscape plugin to assess overrepresentation of Gene Ontology categories in biological networks. Bioinformatics 21, 3448–3449 (2005).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Gui, S. et al. ZEAMAP, a comprehensive database adapted to the maize multi-omics era. iScience 23, 101241 (2020).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Mikheenko, A., Prjibelski, A., Saveliev, V., Antipov, D. & Gurevich, A. Versatile genome assembly evaluation with QUAST-LG. Bioinformatics 34, i142–i150 (2018).
Nakamura, T., Yamada, K. D., Tomii, K. & Katoh, K. Parallelization of MAFFT for large-scale multiple sequence alignments. Bioinformatics 34, 2490–2492 (2018).
Ishida, Y., Hiei, Y. & Komari, T. Agrobacterium-mediated transformation of maize. Nat. Protoc. 2, 1614–1621 (2007).
Wang, B. et al. Tryptophan-independent auxin biosynthesis contributes to early embryogenesis in Arabidopsis. Proc. Natl Acad. Sci. USA 112, 4821–4826 (2015).
Tan, H. et al. A crucial role of GA-regulated flavonol biosynthesis in root growth of Arabidopsis. Mol. Plant 12, 521–537 (2019).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
This study was financially supported by the National Key Research and Development Program of China (grant nos. 2021YFF1000500 (Q.P.), 2021YFD1200700 (F.C.) and 2016YFD0100700 (L.Y.)), the National Natural Science Foundation of China (grant nos. 31972485 (F.C.) and 31971948 (Q.P.)), the Hainan Natural Science Foundation Innovation Research Team Project (grant no. 321CXTD443 (F.C.)), the Hainan Provincial Science and Technology Plan Sanya Yazhou Bay Science and Technology City Joint Project (grant no. 320LH011 (Q.P.)) and the China Postdoctoral Science Foundation (grant no. 2021M693431 (W.R.)). The transgenic maize seeds were produced by the Center for Crop Functional Genomics and Molecular Breeding of China Agricultural University.
Author information
Authors and Affiliations
Contributions
Q.P., L.Y. and F.C. conceived and designed the research. W.R., L.Z., J. Liang, L.W., P.L., Z.L., X.L., Z. Zhang and J. Li performed phenotypic measurements. W.R. and Q.P. performed the data analyses. L.C. performed plasmid construction and genetic transformation. W.R., K.H. and Z. Zhao characterized the transgenic overexpression lines. J.Y. provided the maize inbred lines and genotype set. W.R. and Q.P. wrote the manuscript. F.A., G.M., J.Y., F.Z., F.C., L.Y. and Q.P. revised the manuscript. All authors contributed to the final version of the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Yusaku Uga, Ana Letycia Basso Garcia, Ana Caño-Delgado and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
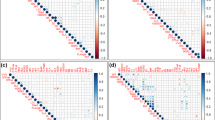
Extended Data Fig. 1 Pearson correlations among eight root traits.
The red lines represent positive correlations, and the green lines represent negative correlations. The line width represents the strength of the correlation. Yellow lines indicate that the correlation coefficient was close to zero.
Extended Data Fig. 2 Principal component analysis of eight root traits.
The red ellipse indicates the area-related traits; the yellow ellipse indicates the width-related traits; and the green ellipse indicates the angle-related traits.
Extended Data Fig. 3 Cluster analysis of 380 maize inbred lines based on root traits.
(a) Cluster analysis of 380 inbred lines based on ROA, RMEW, and AREA. (b) Representative inbred lines from the three clusters (groups 1–3). (c) Comparison of eight root traits among groups 1–3. (d) Comparison of eight root traits among four subpopulations. (e) Proportion of lines from each of four subgroups in the three cluster groups. The four subgroups (Mixed, SS, NSS, and TST) are based on genetic relationships among the different inbred lines. Mixed, mixed group; SS, stiff stalk group; NSS, non-stiff stalk group; TST, tropical and subtropical group.
Supplementary information
Supplementary Information
Supplementary Note and Figs. 1–19.
Supplementary Table 28
Supplementary Tables 1–28.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ren, W., Zhao, L., Liang, J. et al. Genome-wide dissection of changes in maize root system architecture during modern breeding. Nat. Plants 8, 1408–1422 (2022). https://doi.org/10.1038/s41477-022-01274-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-022-01274-z
This article is cited by
-
Natural variation in ZmNAC087 contributes to total root length regulation in maize seedlings under salt stress
BMC Plant Biology (2023)
-
Hybrid performance evaluation and genome-wide association analysis of root system architecture in a maize association population
Theoretical and Applied Genetics (2023)
-
Laser ablation tomography monitors lateral root development in maize: a pictorial-based study case
Brazilian Journal of Botany (2023)
-
Genetic architecture of ear traits based on association mapping and co-expression networks in maize inbred lines and hybrids
Molecular Breeding (2023)
-
The genetic architecture of prolificacy in maize revealed by association mapping and bulk segregant analysis
Theoretical and Applied Genetics (2023)