Abstract
When confronted with heat stress, plants depend on the timely activation of cellular defences to survive by perceiving the rising temperature. However, how plants sense heat at the whole-plant level has remained unanswered. Here we demonstrate that shoot apical nitric oxide (NO) bursting under heat stress as a signal triggers cellular heat responses at the whole-plant level on the basis of our studies mainly using live-imaging of transgenic plants harbouring pHsfA2::LUC, micrografting, NO accumulation mutants and liquid chromatography–tandem mass spectrometry analysis in Arabidopsis. Furthermore, we validate that S-nitrosylation of the trihelix transcription factor GT-1 by S-nitrosoglutathione promotes its binding to NO-responsive elements in the HsfA2 promoter and that loss of function of GT-1 disrupts the activation of HsfA2 and heat tolerance, revealing that GT-1 is the long-sought mediator linking signal perception to the activation of cellular heat responses. These findings uncover a heat-responsive mechanism that determines the timing and execution of cellular heat responses at the whole-plant level.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data generated or analysed during this study are included in the published article and the Supplementary Information. All Arabidopsis genes involved in this study can be found at TAIR (www.arabidopsis.org), with the following accession numbers: GT-1 (AT1G13450), HsfA2 (AT2G26150), Hsp101 (AT1G74310), Hsp70 (AT3G12580), Hsp25.3 (AT4G27670), Hsp18.1 (AT5G59720), Hsp17.7 (AT5G12030), APX2 (AT3G09640) and GolS1 (AT2G47180). Source data are provided with this paper.
References
Choudhury, F. K., Rivero, R. M., Blumwald, E. & Mittler, R. Reactive oxygen species, abiotic stress and stress combination. Plant J. 90, 856–867 (2017).
Fancy, N. N., Bahlmann, A. K. & Loake, G. J. Nitric oxide function in plant abiotic stress. Plant Cell Environ. 40, 462–472 (2017).
Domingos, P., Prado, A. M., Wong, A., Gehring, C. & Feijo, J. A. Nitric oxide: a multitasked signaling gas in plants. Mol. Plant 8, 506–520 (2015).
Besson-Bard, A., Pugin, A. & Wendehenne, D. New insights into nitric oxide signaling in plants. Annu. Rev. Plant Biol. 59, 21–39 (2008).
Yun, B.-W. et al. Nitric oxide and S-nitrosoglutathione function additively during plant immunity. N. Phytol. 211, 516–526 (2016).
Gaupels, F., Durner, J. & Kogel, K.-H. Production, amplification and systemic propagation of redox messengers in plants? The phloem can do it all! N. Phytol. 214, 554–560 (2017).
Mur, L. A. J. et al. Nitric oxide in plants: an assessment of the current state of knowledge. AoB Plants 5, pls052 (2013).
Gould, K. S., Lamotte, O., Klinguer, A., Pugin, A. & Wendehenne, D. Nitric oxide production in tobacco leaf cells: a generalized stress response? Plant Cell Environ. 26, 1851–1862 (2003).
Yu, M., Lamattina, L., Spoel, S. H. & Loake, G. J. Nitric oxide function in plant biology: a redox cue in deconvolution. N. Phytol. 202, 1142–1156 (2014).
Bouchard, J. N. & Yamasaki, H. Heat stress stimulates nitric oxide production in Symbiodinium microadriaticum: a possible linkage between nitric oxide and the coral bleaching phenomenon. Plant Cell Physiol. 49, 641–652 (2008).
Uchida, A., Jagendorf, A. T., Hibino, T., Takabe, T. & Takabe, T. Effects of hydrogen peroxide and nitric oxide on both salt and heat stress tolerance in rice. Plant Sci. 163, 515–523 (2002).
Song, L., Ding, W., Zhao, M., Sun, B. & Zhang, L. Nitric oxide protects against oxidative stress under heat stress in the calluses from two ecotypes of reed. Plant Sci. 171, 449–458 (2006).
Hasanuzzaman, M., Nahar, K., Alam, M. M., Roychowdhury, R. & Fujita, M. Physiological, biochemical, and molecular mechanisms of heat stress tolerance in plants. Int. J. Mol. Sci. 14, 9643–9684 (2013).
Xuan, Y., Zhou, S., Wang, L., Cheng, Y. & Zhao, L. Nitric oxide functions as a signal and acts upstream of AtCaM3 in thermotolerance in Arabidopsis seedlings. Plant Physiol. 153, 1895–1906 (2010).
Feechan, A. et al. A central role for S-nitrosothiols in plant disease resistance. Proc. Natl Acad. Sci. USA 102, 8054–8059 (2005).
Lee, U., Wie, C., Fernandez, B. O., Feelisch, M. & Vierling, E. Modulation of nitrosative stress by S-nitrosoglutathione reductase is critical for thermotolerance and plant growth in Arabidopsis. Plant Cell 20, 786–802 (2008).
Nishizawa, A. et al. Arabidopsis heat shock transcription factor A2 as a key regulator in response to several types of environmental stress. Plant J. 48, 535–547 (2006).
Charng, Y. Y. et al. A heat-inducible transcription factor, HsfA2, is required for extension of acquired thermotolerance in Arabidopsis. Plant Physiol. 143, 251–262 (2007).
Chen, S.-T., He, N.-Y., Chen, J.-H. & Guo, F.-Q. Identification of core subunits of photosystem II as action sites of HSP21, which is activated by the GUN5-mediated retrograde pathway in Arabidopsis. Plant J. 89, 1106–1118 (2017).
Yu, H.-D. et al. Downregulation of chloroplast RPS1 negatively modulates nuclear heat-responsive expression of HsfA2 and its target genes in Arabidopsis. PLoS Genet. 8, e1002669 (2012).
Wang, L. et al. Hydrogen peroxide acts upstream of nitric oxide in the heat shock pathway in Arabidopsis seedlings. Plant Physiol. 164, 2184–2196 (2014).
Begara-Morales, J. C. et al. Differential transcriptomic analysis by RNA-seq of GSNO-responsive genes between Arabidopsis roots and leaves. Plant Cell Physiol. 55, 1080–1095 (2014).
Kaplan-Levy, R. N., Brewer, P. B., Quon, T. & Smyth, D. R. The trihelix family of transcription factors—light, stress and development. Trends Plant Sci. 17, 163–171 (2012).
Kotak, S. et al. Complexity of the heat stress response in plants. Curr. Opin. Plant Biol. 10, 310–316 (2007).
Mittler, R., Finka, A. & Goloubinoff, P. How do plants feel the heat? Trends Biochem. Sci. 37, 118–125 (2012).
Ohama, N., Sato, H., Shinozaki, K. & Yamaguchi-Shinozaki, K. Transcriptional regulatory network of plant heat stress response. Trends Plant Sci. 22, 53–65 (2017).
Penfield, S. Temperature perception and signal transduction in plants. N. Phytol. 179, 615–628 (2008).
Saidi, Y., Finka, A. & Goloubinoff, P. Heat perception and signalling in plants: a tortuous path to thermotolerance. N. Phytol. 190, 556–565 (2011).
Vierling, E. The roles of heat-shock proteins in plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 42, 579–620 (1991).
Lam Dai, V., Gevaert, K. & De Smet, I. Feeling the heat: searching for plant thermosensors. Trends Plant Sci. 24, 210–219 (2019).
Zhu, J.-K. Abiotic stress signaling and responses in plants. Cell 167, 313–324 (2016).
Ruelland, E. & Zachowski, A. How plants sense temperature. Environ. Exp. Bot. 69, 225–232 (2010).
Hayes, S., Schachtschabel, J., Mishkind, M., Munnik, T. & Arisz, S. A. Hot topic: thermosensing in plants. Plant Cell Environ. 44, 2018–2033 (2021).
Finka, A., Cuendet, A. F. H., Maathuis, F. J. M., Saidi, Y. & Goloubinoff, P. Plasma membrane cyclic nucleotide gated calcium channels control land plant thermal sensing and acquired thermotolerance. Plant Cell 24, 3333–3348 (2012).
Kumar, S. V. & Wigge, P. A. H2A.Z-containing nucleosomes mediate the thermosensory response in Arabidopsis. Cell 140, 136–147 (2010).
Kumar, S. V. et al. Transcription factor PIF4 controls the thermosensory activation of flowering. Nature 484, 242–U127 (2012).
Jung, J.-H. et al. Phytochromes function as thermosensors in Arabidopsis. Science 354, 886–889 (2016).
Legris, M. et al. Phytochrome B integrates light and temperature signals in Arabidopsis. Science 354, 897–900 (2016).
Lamers, J., van der Meer, T. & Testerink, C. How plants sense and respond to stressful environments. Plant Physiol. 182, 1624–1635 (2020).
Wigge, P. A. Ambient temperature signalling in plants. Curr. Opin. Plant Biol. 16, 661–666 (2013).
Sanchez-Vicente, I., Lechon, T., Fernandez-Marcos, M., Sanz, L. & Lorenzo, O. Nitric oxide alters the pattern of auxin maxima and PIN-FORMED1 during shoot development. Front. Plant Sci. 12, 630792 (2021).
Airaki, M. et al. Detection and quantification of S-nitrosoglutathione (GSNO) in pepper (Capsicum annuum L.) plant organs by LC-ES/MS. Plant Cell Physiol. 52, 2006–2015 (2011).
Barroso, J. B. et al. Localization of S-nitrosoglutathione and expression of S-nitrosoglutathione reductase in pea plants under cadmium stress. J. Exp. Bot. 57, 1785–1793 (2006).
Corpas, F. J., Alche, J. D. & Barroso, J. B. Current overview of S-nitrosoglutathione (GSNO) in higher plants. Front. Plant Sci. 4, 126 (2013).
Espunya, M., De Michele, R., Gomez-Cadenas, A. & Carmen Martinez, M. S-Nitrosoglutathione is a component of wound- and salicylic acid-induced systemic responses in Arabidopsis thaliana. J. Exp. Bot. 63, 3219–3227 (2012).
Bramanti, E. et al. Determination of S-nitrosoglutathione in plasma: comparison of two methods. Talanta 81, 1295–1299 (2010).
Tsikas, D. et al. UPLC-MS/MS measurement of S-nitrosoglutathione (GSNO) in human plasma solves the S-nitrosothiol concentration enigma. J. Chromatogr. B 927, 147–157 (2013).
Pate, J. S., Sharkey, P. J. & Lewis, O. A. M. Phloem bleeding from legume fruits—technique for study of fruit nutrition. Planta 120, 229–243 (1974).
Gopal, M., Shil, S., Gupta, A., Hebbar, K. B. & Arivalagan, M. Metagenomic investigation uncovers presence of probiotic-type microbiome in Kalparasa® (fresh unfermented coconut inflorescence sap). Front. Microbiol. 12, 662783 (2021).
Hebbar, K. B. et al. Nutritional profiling of coconut (Cocos nucifera L.) inflorescence sap collected using novel coco-sap chiller method and its value added products. J. Food Meas. Charact. 14, 2703–2712 (2020).
Hewer, A., Will, T. & van Bel, A. J. E. Plant cues for aphid navigation in vascular tissues. J. Exp. Biol. 213, 4030–4042 (2010).
Kollist, H. et al. Rapid responses to abiotic stress: priming the landscape for the signal transduction network. Trends Plant Sci. 24, 25–37 (2019).
Baxter, A., Mittler, R. & Suzuki, N. ROS as key players in plant stress signalling. J. Exp. Bot. 65, 1229–1240 (2014).
Choi, W.-G. et al. Orchestrating rapid long-distance signaling in plants with Ca2+, ROS and electrical signals. Plant J. 90, 698–707 (2017).
Gilroy, S. et al. ROS, calcium, and electric signals: key mediators of rapid systemic signaling in plants. Plant Physiol. 171, 1606–1615 (2016).
Zandalinas, S. I. & Mittler, R. Vascular and non-vascular transmission of systemic reactive oxygen signals during wounding and heat stress. Plant Physiol. 186, 1721–1733 (2021).
Bellin, D., Asai, S., Delledonne, M. & Yoshioka, H. Nitric oxide as a mediator for defense responses. Mol. Plant Microbe Interact. 26, 271–277 (2013).
Trapet, P. et al. NO signaling in plant immunity: a tale of messengers. Phytochemistry 112, 72–79 (2015).
Tanou, G. et al. Oxidative and nitrosative-based signaling and associated post-translational modifications orchestrate the acclimation of citrus plants to salinity stress. Plant J. 72, 585–599 (2012).
del Rio, L. A. ROS and RNS in plant physiology: an overview. J. Exp. Bot. 66, 2827–2837 (2015).
Feng, Z. et al. Efficient genome editing in plants using a CRISPR/Cas system. Cell Res. 23, 1229–1232 (2013).
Ma, X. et al. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol. Plant 8, 1274–1284 (2015).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Chen, J.-H. et al. Nuclear-encoded synthesis of the D1 subunit of photosystem II increases photosynthetic efficiency and crop yield. Nat. Plants 6, 570–580 (2020).
Wang, Q.-L., Sun, A.-Z., Chen, S.-T., Chen, L.-S. & Guo, F.-Q. SPL6 represses signalling outputs of ER stress in control of panicle cell death in rice. Nat. Plants 4, 280–288 (2018).
Cervera, M. Histochemical and fluorometric assays for uidA (GUS) gene detection. Methods Mol. Biol. 286, 203–213 (2005).
Prunet, N., Jack, T. P. & Meyerowitz, E. M. Live confocal imaging of Arabidopsis flower buds. Dev. Biol. 419, 114–120 (2016).
Lichtenthaler, H. K. Chlorophylls and carotenoids—pigments of photosynthetic biomembranes. Methods Enzymol. 148, 350–382 (1987).
Deeken, R. et al. Identification of Arabidopsis thaliana phloem RNAs provides a search criterion for phloem-based transcripts hidden in complex datasets of microarray experiments. Plant J. 55, 746–759 (2008).
Murray, C. I., Uhrigshardt, H., O’Meally, R. N., Cole, R. N. & Van Eyk, J. E. Identification and quantification of S-nitrosylation by cysteine reactive tandem mass tag switch assay. Mol. Cell. Proteom. 11, M111.013441 (2012).
Jaffrey, S. R., Erdjument-Bromage, H., Ferris, C. D., Tempst, P. & Snyder, S. H. Protein S-nitrosylation: a physiological signal for neuronal nitric oxide. Nat. Cell Biol. 3, 193–197 (2001).
Yang, H. et al. S-Nitrosylation positively regulates ascorbate peroxidase activity during plant stress responses. Plant Physiol. 167, 1604–U753 (2015).
Feng, J. et al. S-Nitrosylation of phosphotransfer proteins represses cytokinin signaling. Nat. Commun. 4, 1529 (2013).
Gao, X. et al. Downregulation of Rubisco activity by non-enzymatic acetylation of RbcL. Mol. Plant 9, 1018–1027 (2016).
Chen, H. et al. Arabidopsis CULLIN4 forms an E3 ubiquitin ligase with RBX1 and the CDD complex in mediating light control of development. Plant Cell 18, 1991–2004 (2006).
Acknowledgements
This work was supported by grants from the Chinese Academy of Sciences (the Strategic Priority Research Program, no. XDB27040105), the Ministry of Science and Technology of China (National Key R&D Program of China, no. 2020YFA0907604) and the National Natural Science Foundation of China (nos U1812401, 31770314 and 31570260). We thank N. Crawford (University of California at San Diego, USA), Z.-M. Pei (Duke University, USA), S. Xu (Northwest A&F University, China) and E. Vierling (University of Massachusetts, USA) for providing plant materials; J.-K. Zhu (Shanghai Center for Plant Stress Biology, CAS, China) and Y.-G. Liu (South China Agricultural University, China) for providing the plasmids of the CRISPR–Cas9 gene-editing system; and the CEMPS Core Facility for technical assistance.
Author information
Authors and Affiliations
Contributions
F.-Q.G. conceived the project. F.-Q.G. and N.-Y.H. designed the experiments. N.-Y.H. performed most of the experimental work with assistance from A.-Z.S., Y.Z. and S.-N.Y. L.-S.C. contributed to the in situ hybridization. N.-Y.H., F.-Q.G. and L.-S.C. analysed the data. F.-Q.G. and N.-Y.H. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Christian Lindermayr, Gary Loake, Nobuhiro Suzuki and Ziqiang Zhu for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Heat stress triggers the transcriptional activation of HsfA2 firstly at shoot apical region, subsequently at young rosette leaves and old leaves.
Intensity and developing patterns of LUC-signals were shown in the transgenic Arabidopsis seedlings (20-d-old) harboring the pHsfA2::LUC construct in which a luciferase reporter gene (LUC) was driven by the HsfA2 promoter under control condition (a) or subjected to heat stress (38 °C, 30 min) (b). Under heat stress conditions, time-lapse imaging showed rapid systemic signal transduction from shoot apical region to young rosette leaves and then old leaves via vascular system (leaf veins). Bars = 1 cm.
Extended Data Fig. 2 Transcriptional activation of HsfA2 is severely inhibited in wild-type flowering plants with shoot tip removed in response to heat stress.
The expression level of HsfA2 was analyzed by qRT-PCR in the rosette leaves of wild-type flowering plants or wild-type flowering plants with inflorescence shoot tip removed when subjected to heat stress (38 °C) for the indicated times. Individual values are means of three independent biological replicates. Bars indicate standard deviation (SD) (*P value<0.05, **P value<0.01, ***P value<0.001, two-sided Student’s t test). AtACTIN2 was used as the internal control.
Extended Data Fig. 3 Heat stress induces burst of nitric oxide (NO) at shoot apex and accumulation in phloem in Arabidopsis seedlings.
a, Detection of endogenous nitric oxide (NO) at shoot apex by the specific probe 4-amino-5-methylamino-2’,7’-difluorofluorescein diacetate (DAF-FM DA) in 5-day-old wild-type seedlings grown under control conditions (Control) or subjected to heat treatment (38 °C, 5 min) (Heat) by a confocal microscopy. YL, young leaf; SAM, shoot apical meristem. Bars = 20 µm. Three times these experiments were repeated independently with similar results. b, Detection of NO by DAF-FM DA using a fluorescence microscope in hand-made stem cross-sections taken from the inflorescence stems of 23-d-old Arabidopsis plants grown under control conditions (Control) or subjected to heat treatment (38 °C, 30 min) (Heat). Bars = 100 µm. Three times these experiments were repeated independently with similar results. c, Close-up view of the image of DAF-FM fluorescence signals in (b). ep, epidermis; co, cortex; ph, phloem; xylem, xy. Stem-cross sections were also stained with propidium iodide (PI). Bar = 100 µm.
Extended Data Fig. 4 The transcriptional activation of HsfA2 was severely inhibited in the NO-deficient mutant noa1 in response to heat stress.
a, Time-lapse imaging of systemic signal transduction from shoot to root to initiate transcriptional activation of a luciferase reporter gene (LUC) driven by the HsfA2 promoter (pHsfA2::LUC) in the transgenic Arabidopsis plants of wild-type (upper panels) and noa1 (lower panels) under control conditions. b, Time-lapse imaging of systemic signal transduction from shoot to root to initiate rapid transcriptional activation of a luciferase reporter gene (LUC) driven by the HsfA2 promoter (pHsfA2::LUC) in the transgenic Arabidopsis plant of wild-type (upper panels), but relatively slower and weaker transcriptional activation in noa1 (lower panels) in response to heat stress (38 °C, 30 min) (b). Bars = 1 cm.
Extended Data Fig. 5 Micrografting with the heat-stressed wild-type or noa1 shoots as scion and the transgenic wild-type or noa1 roots containing pHsfA2::LUC as rootstocks.
a, Time-lapse imaging showed systemic signal transduction from shoot to root when a control wild-type shoot (upper panels and lower panels) or a control NO-deficient mutant noa1 shoot (middle panels) was grafted onto the rootstock taken from the transgenic plant of wild-type or noa1 harboring the pHsfA2::LUC construct. White arrowheads indicate grafting joints. b, Time-lapse imaging showed rapid systemic signal transduction from shoot to root when a heat-treated wild-type shoot (upper panels and lower panels), rather than the heat-treated noa1 shoot (middle panels), was grafted onto the rootstock taken from the transgenic plant of wild-type or noa1 harboring the pHsfA2::LUC construct. White arrowheads indicate grafting joints. Bars = 1 cm.
Extended Data Fig. 6 Characterization of the T-DNA insertion mutant gt-1-1 and heat-sensitive phenotypes of two allelic mutants gt-1-2 and gt-1-3 generated by the CRISPR-cas9 technique.
a, Schematic diagram of the GT-1 gene showing the T-DNA insertion site for gt-1-1 and the editing sites for two allelic mutants gt-1-2 (“TCACAA” inserted at 149 bp from ATG) and gt-1-3 (“TA” deleted at 302 bp from ATG) generated by the CRISPR-cas9 technique. Closed box indicates ORF. Exons (boxes) and introns (lines) were determined by a comparison of the genomic and cDNA sequences. The T-DNA insertion site (931 bp from ATG) and position of the start codon are indicated. b, The specific primers (LP + RP and LBb1.3 + RP), derived from the left (LP) and right (RP) borders of T-DNA insertion and the T-DNA vector sequence (LBb1.3) respectively, were used to amplify the indicated fragments in wild-type and gt-1-1 mutant plants. The amplified fragments were confirmed by sequencing. c, The protein abundance of GT-1 in leaves taken from the plants of wild-type and the gt-1-1 mutant was analyzed by western blots with a polyclonal antibody against GT-1. Ponceau S staining was used as an internal loading control. d, e Expression analysis of GT-1 in the detached fully-expanded leaves of wild-type, the gt-1-1 mutant and the complemented gt-1-1 mutant (gt-1-1 comp) with pGT-1::GT-1 cDNA-GFP (d), and in the detached fully-expanded leaves of wild-type and the CRISPR-cas9-generated gt-1-2 and gt-1-3 mutants (d) by qRT-PCR. It should be noted that around 20% expression level in relative to wild-type was detected in gt-1-2, but the resulted transcript with frameshift mutations had nothing to do with GT-1 when the resulted transcript was sequenced. In gt-1-2, as shown in (a), “TCACAA” was inserted at 149 bp from ATG and this kind of insertion led to a frameshift mutation. No signal was detected in gt-1-3. Individual values are means of three independent biological replicates. Bars indicate standard deviation (SD). ND, non-detectable. ACTIN2 (At3g18780) was used as the internal control. f, g, Phenotypes of the wild-type, gt-1-2 and gt-1-3 as examined with detached fully-expanded leaves (f) and whole plants (g) challenged with heat treatments for the indicated time in dark. Bars = 1 cm in (f) or 2 cm in (g).
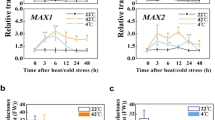
Extended Data Fig. 7 Knockdown of GT-1 inhibits transcriptional activation of the HsfA2-target genes in response to heat stress.
Expression analysis of the representative downstream target genes of HsfA2 including Hsp17.7II, Hsp18.1, GolS1, Hsp70, Hsp25.3, APX2 and Hsp101 in detached fully-expanded leaves of the wild-type and gt-1-1 mutant at the indicated time when subjected to 2-h-heat treatment (38 °C) in dark. Individual values are means of three independent biological replicates. Bars indicate standard deviation (SD). AtACTIN2 was used as the internal control.
Extended Data Fig. 8 Effects of S-nitrosylation of GT-1 by GSNO on its binding activities to the promoter of HsfA2.
a, b, Recombinant TF-GT-1 was incubated with the indicated concentrations of GSNO and subsequently subjected to the EMSA assay for examining the binding activity of TF-GT-1 to the cis-element 1 (cis1) (a) and the cis-element 2 (cis2) (b) identified in the HsfA2 promoter. c, Single residue mutagenesis of Cys324 or Cys347 of GT-1 led to partial losses of its GSNO-dependent binding activity to the cis-element 1 in the promoter of HsfA2. EMSA assay was performed to examine the binding activity of TF-GT-1 and the mutated TF-GT-1 proteins with the site-directed mutagenesis of Cys324 and Cys347 into Ser residues in single residue mutagenesis (GT-1C324S or GT-1C347S), treated with GSNO (10 μM) or plus cPTIO (200 μM). d, In vitro S-nitrosylation of TF-GT-1C324/347 S containing double-residue mutations of Cys324 and Cys347 into Ser residues (only Cys135 residue is available for S-nitrosylation) with GSNO treatment. The purified recombinant TF-GT-1C324/347 S was incubated with the indicated concentrations of GSNO and then subjected to the TMT-switch assay. TMT is referred to as “tandem mass tags“. The S-nitrosylation levels of resulting proteins were examined by western blot analysis using an anti-TMT antibody. Ponceau S staining was used as an internal loading control. e, EMSA assay was performed to examine the binding activity of TF-GT-1C324/347 S with the site-directed mutagenesis of Cys324 and Cys347 into Ser residues, treated with different concentrations of GSNO (5-200 μM). Three times these experiments were repeated independently with similar results as shown in (a-e).
Extended Data Fig. 9 Quantification of NO levels in wild-type and the NO-related mutants such as noa1, hot5-2/gsnor1 and nox1 under control or heat stress conditions.
The levels of NO were detected by the specific probe 4-amino-5-methylamino-2’,7’-difluorofluorescein diacetate (DAF-FM DA) at shoot apical regions of 5-day-old wild-type seedlings and the NO-related mutants such as noa1, hot5-2/gsnor1 and nox1 under control or subjected to heat treatment (38oC, 5 min) (Heat) or subjected to heat treatment (38 °C, 5 min) plus the NO scavenger cPTIO treatment (100 μM) (Heat + cPTIO) by a confocal microscopy (LSM880, Zeiss), respectively. Seventeen individual seedlings were examined for each genotype or treatment. The signal intensity of DAF-FM fluorescence in each image was quantified using ZEN software (Zeiss). Data were shown as mean of the signal intensities from seventeen individual seedlings (n = 17). Error bars indicate SD (***P value<0.001, two-sided Student’s t test).
Extended Data Fig. 10 Relative abundance of the S-nitrosylated peptides of GT-1 identified with Cys135, Cys324 or Cys347 as the S-nitrosylated residues respectively in wild-type and hot5-2 under heat stress.
a, b, c, For quantification of the S-nitrosylated peptides of GT-1 identified with Cys135 (a), Cys324 (b) or Cys347 (c) as the S-nitrosylated residues respectively in 5-d-old Arabidopsis seedlings of wild-type and hot5-2 were subjected to heat treatment (38 °C) and harvested at the indicated time points by the MS/MS-based analysis strategy using isotope-coded cysteine thiol-reactive multiplex regent, referred to as “tandem mass tags” (TMTs). The total proteins extracted from Arabidopsis seedlings were first labeled with TMT. The GT-1 protein was immunoprecipitated with anti-GT-1 antibody. The resultant samples were separated on 10% SDS–PAGE. After slicing the protein band corresponding to GT-1 and destaining, the gel slices were reduced with DTT, alkylated with iodoacetamide, and digested with trypsin (V5280, Promega) at 37 °C overnight. The sample was then purified with a C-18 spin column (PepClean C-18 Spin Columns, Thermo), and the resultant peptides were examined by LC–MS/MS. The MS data were initially searched against the NCBI database with the aid of the Sequest search engine. The resultant MS spectra were used to determine the peptide identity and abundance of each peptide in the same spectrum. Relative abundance of peptide was calculated by comparing the intensity of the corresponding tag. The absolute quantification was calculated by comparing with the internal control peptides, and the sum of the S-nitrosylated and non-nitrosylated peptides for each pair was set to 100%. Data are presented as mean values + /- SD (n = 3 independent biological replicates).
Supplementary information
Supplementary Information
Supplementary Figs. 1–13 (Figs. 11–13 contain only the full scans for Figs. 7b, 9 and 10) and Table 1.
Supplementary Data 1
Statistical source data for Supplementary Fig. 3.
Supplementary Data 2
Statistical source data for Supplementary Fig. 8a,b.
Source data
Source Data Fig. 5
Statistical source data.
Source Data Fig. 5
Unprocessed gels.
Source Data Fig. 6
Unprocessed western blots/gels.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 6
Unprocessed gels/western blots.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Unprocessed western blots.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
He, NY., Chen, LS., Sun, AZ. et al. A nitric oxide burst at the shoot apex triggers a heat-responsive pathway in Arabidopsis. Nat. Plants 8, 434–450 (2022). https://doi.org/10.1038/s41477-022-01135-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-022-01135-9