Abstract
During plant development, a precise balance of cytokinin is crucial for correct growth and patterning, but it remains unclear how this is achieved across different cell types and in the context of a growing organ. Here we show that in the root apical meristem, the TMO5/LHW complex increases active cytokinin levels via two cooperatively acting enzymes. By profiling the transcriptomic changes of increased cytokinin at single-cell level, we further show that this effect is counteracted by a tissue-specific increase in CYTOKININ OXIDASE 3 expression via direct activation of the mobile transcription factor SHORTROOT. In summary, we show that within the root meristem, xylem cells act as a local organizer of vascular development by non-autonomously regulating cytokinin levels in neighbouring procambium cells via sequential induction and repression modules.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Upon acceptance, the scRNA-seq data will be made accessible via an on-line browser tool (http://bioit3.irc.ugent.be/plant-sc-atlas/) and raw data can be accessed at NCBI with GEO number GSE179820. All other data are either in the main paper or the Supplementary Material. Source data are provided with this paper. Requests should be directed to the corresponding authors.
References
Lucas, W. J. et al. The plant vascular system: evolution, development and functions. J. Integr. Plant Biol. 55, 294–388 (2013).
De Rybel, B., Mähönen, A. P., Helariutta, Y. & Weijers, D. Plant vascular development: from early specification to differentiation. Nat. Rev. Mol. Cell Biol. 17, 30–40 (2016).
Bishopp, A. et al. A mutually inhibitory interaction between auxin and cytokinin specifies vascular pattern in roots. Curr. Biol. 21, 917–926 (2011).
De Rybel, B. et al. Plant development. Integration of growth and patterning during vascular tissue formation in Arabidopsis. Science 345, 1255215 (2014).
Ohashi-Ito, K. et al. A bHLH complex activates vascular cell division via cytokinin action in root apical meristem. Curr. Biol. 24, 2053–2058 (2014).
De Rybel, B. et al. A bHLH complex controls embryonic vascular tissue establishment and indeterminate growth in Arabidopsis. Dev. Cell 24, 426–437 (2013).
Katayama, H. et al. A negative feedback loop controlling bHLH complexes is involved in vascular cell division and differentiation in the root apical meristem. Curr. Biol. 25, 3144–3150 (2015).
Ohashi-Ito, K. & Bergmann, D. C. Regulation of the Arabidopsis root vascular initial population by LONESOME HIGHWAY. Development 134, 2959–2968 (2007).
Ohashi-Ito, K., Matsukawa, M. & Fukuda, H. An atypical bHLH transcription factor regulates early xylem development downstream of auxin. Plant Cell Physiol. 54, 398–405 (2013).
Ohashi-Ito, K., Oguchi, M., Kojima, M., Sakakibara, H. & Fukuda, H. Auxin-associated initiation of vascular cell differentiation by LONESOME HIGHWAY. Development 140, 765–769 (2013).
Vera-Sirera, F. et al. A bHLH-based feedback loop restricts vascular cell proliferation in plants. Dev. Cell 35, 432–443 (2015).
Miyashima, S. et al. Mobile PEAR transcription factors integrate positional cues to prime cambial growth. Nature 565, 490–494 (2019).
Smet, W. et al. DOF2.1 controls cytokinin-dependent vascular cell proliferation downstream of TMO5/LHW. Curr. Biol. 29, 520–529.e6 (2019).
Wybouw, B. & De Rybel, B. Cytokinin - a developing story. Trends Plant Sci. 24, 177–185 (2019).
Kuroha, T. et al. Functional analyses of LONELY GUY cytokinin-activating enzymes reveal the importance of the direct activation pathway in Arabidopsis. Plant Cell 21, 3152–3169 (2009).
Tokunaga, H. et al. Arabidopsis lonely guy (LOG) multiple mutants reveal a central role of the LOG-dependent pathway in cytokinin activation. Plant J. 69, 355–365 (2012).
Mähönen, A. P. et al. A novel two-component hybrid molecule regulates vascular morphogenesis of the Arabidopsis root. Genes Dev. 14, 2938–2943 (2000).
Wendrich, J. R. et al. Vascular transcription factors guide plant epidermal responses to limiting phosphate conditions. Science 370, eaay4970 (2020).
Xu, Z. et al. Functional genomic analysis of Arabidopsis thaliana glycoside hydrolase family 1. Plant Mol. Biol. 55, 343–367 (2004).
Leah, R., Kigel, J., Svendsen, I. & Mundy, J. Biochemical and molecular characterization of a barley seed beta-glucosidase. J. Biol. Chem. 270, 15789–15797 (1995).
Dharmawardhana, D. P., Ellis, B. E. & Carlson, J. E. A beta-glucosidase from lodgepole pine xylem specific for the lignin precursor coniferin. Plant Physiol. 107, 331–339 (1995).
Brzobohatý, B. et al. Release of active cytokinin by a beta-glucosidase localized to the maize root meristem. Science 262, 1051–1054 (1993).
Weijers, D. et al. An Arabidopsis Minute-like phenotype caused by a semi-dominant mutation in a RIBOSOMAL PROTEIN S5 gene. Development 128, 4289–4299 (2001).
Bartrina, I., Otto, E., Strnad, M., Werner, T. & Schmülling, T. Cytokinin regulates the activity of reproductive meristems, flower organ size, ovule formation, and thus seed yield in Arabidopsis thaliana. Plant Cell 23, 69–80 (2011).
Zurcher, E. et al. A robust and sensitive synthetic sensor to monitor the transcriptional output of the cytokinin signaling network in planta. Plant Physiol. 161, 1066–1075 (2013).
Werner, T., Motyka, V., Strnad, M. & Schmülling, T. Regulation of plant growth by cytokinin. Proc. Natl Acad. Sci. USA 98, 10487–10492 (2001).
Matsumoto-Kitano, M. et al. Cytokinins are central regulators of cambial activity. Proc. Natl Acad. Sci. USA 105, 20027–20031 (2008).
Schmülling, T. et al. Structure and function of cytokinin oxidase/dehydrogenase genes of maize, rice, Arabidopsis and other species. J. Plant Res. 116, 241–252 (2003).
Lee, D. J. et al. Genome-wide expression profiling of ARABIDOPSIS RESPONSE REGULATOR 7(ARR7) overexpression in cytokinin response. Mol. Genet. Genomics 277, 115–137 (2007).
Rashotte, A. M., Carson, S. D., To, J. P. & Kieber, J. J. Expression profiling of cytokinin action in Arabidopsis. Plant Physiol. 132, 1998–2011 (2003).
Cui, H. et al. Genome-wide direct target analysis reveals a role for SHORT-ROOT in root vascular patterning through cytokinin homeostasis. Plant Physiol. 157, 1221–1231 (2011).
Marquès-Bueno, M. D. M. et al. A versatile Multisite Gateway-compatible promoter and transgenic line collection for cell type-specific functional genomics in Arabidopsis. Plant J. 85, 320–333 (2016).
Levesque, M. P. et al. Whole-genome analysis of the SHORT-ROOT developmental pathway in Arabidopsis. PLoS Biol. 4, e143 (2006).
Mellor, N. et al. Theoretical approaches to understanding root vascular patterning: a consensus between recent models. J. Exp. Bot. 68, 5–16 (2017).
Mähönen, A. P. et al. Cytokinin signaling and its inhibitor AHP6 regulate cell fate during vascular development. Science 311, 94–98 (2006).
Help, H., Mahonen, A. P., Helariutta, Y. & Bishopp, A. Bisymmetry in the embryonic root is dependent on cotyledon number and position. Plant Signal. Behav. 6, 1837–1840 (2011).
Helariutta, Y. et al. The SHORT-ROOT gene controls radial patterning of the Arabidopsis root through radial signaling. Cell 101, 555–567 (2000).
Nakajima, K., Sena, G., Nawy, T. & Benfey, P. N. Intercellular movement of the putative transcription factor SHR in root patterning. Nature 413, 307–311 (2001).
Sozzani, R. et al. Spatiotemporal regulation of cell-cycle genes by SHORTROOT links patterning and growth. Nature 466, 128–132 (2010).
Möller, B. K. et al. Auxin response cell-autonomously controls ground tissue initiation in the early Arabidopsis embryo. Proc. Natl Acad. Sci. USA 114, E2533–E2539 (2017).
Karimi, M., Depicker, A. & Hilson, P. Recombinational cloning with plant gateway vectors. Plant Physiol. 145, 1144–1154 (2007).
Lei, Y. et al. CRISPR-P: a web tool for synthetic single-guide RNA design of CRISPR-system in plants. Mol. Plant 7, 1494–1496 (2014).
De Rybel, B. et al. A novel Aux/IAA28 signaling cascade activates GATA23-dependent specification of lateral root founder cell identity. Curr. Biol. 20, 1697–1706 (2010).
Ursache, R., Andersen, T. G., Marhavý, P. & Geldner, N. A protocol for combining fluorescent proteins with histological stains for diverse cell wall components. Plant J. 93, 399–412 (2018).
Hellemans, J., Mortier, G., De Paepe, A., Speleman, F. & Vandesompele, J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol. 8, R19 (2007).
Gong, X. et al. SEUSS integrates gibberellin signaling with transcriptional inputs from the SHR-SCR-SCL3 module to regulate middle cortex formation in the Arabidopsis root. Plant Physiol. 170, 1675–1683 (2016).
Svačinová, J. et al. A new approach for cytokinin isolation from Arabidopsis tissues using miniaturized purification: pipette tip solid-phase extraction. Plant Methods 8, 17 (2012).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Novák, O., Hauserová, E., Amakorová, P., Dolezal, K. & Strnad, M. Cytokinin profiling in plant tissues using ultra-performance liquid chromatography–electrospray tandem mass spectrometry. Phytochemistry 69, 2214–2224 (2008).
Lun, A. T., Bach, K. & Marioni, J. C. Pooling across cells to normalize single-cell RNA sequencing data with many zero counts. Genome Biol. 17, 75 (2016).
McGinnis, C. S., Murrow, L. M. & Gartner, Z. J. DoubletFinder: doublet detection in single-cell RNA sequencing data using artificial nearest neighbors. Cell Syst. 8, 329–337.e324 (2019).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Acknowledgements
We thank D. Weijers for sharing unpublished materials, T. Schmülling for sharing ckx3, ckx5 and ckx3 ckx5 seeds and K. Ljung for stimulating discussions. This work was funded by The Research Foundation – Flanders (FWO; Odysseus II G0D0515N to B.D.R. and post-doc fellowship 1215820N to B.Y.); the Netherlands Organization for Scientific Research (NWO; VIDI 864.13.00) to W.S., J.R.W. and B.D.R.; Ghent University (BOF20/GOA/012 to M.M. and B.D.R., and BOF18/PDO/151 to J.R.W.); the European Research Council (ERC Starting Grant TORPEDO; 714055) to B.Y. and B.D.R.; the China Scholarship Council (file number 202009350010) to Y.S.; the Ministry of Education, Youth and Sports of the Czech Republic (European Regional Development Fund-Project ‘Plants as a tool for sustainable global development’ No. CZ.02.1.01/0.0/0.0/16_019/ 0000827) to O.N.; and the Internal Grant Agency of Palacký University (IGA_PrF_2021_011) to O.N.
Author information
Authors and Affiliations
Contributions
B.D.R. and B.Y. conceived the project and designed experiments; F.B., L.P., I.P., K.H. and O.N. performed enzymatic assays and cytokinin measurements; J.N. produced and purified BGLU44 protein with the help of J.H.; M.M., K.V., T.E. and Y.S. analysed single-cell data; E.F. and A.B. performed the mathematical modelling; B.Y., M.M., Y.S., W.S. and J.R.W performed all other experiments; B.D.R. supervised the project; B.Y. and B.D.R. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Hitoshi Sakakibara, Enrico Scarpella and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 LOG4 single cell co-expression analysis.
Top 50 genes co-expressed with LOG4 according to a single cell atlas of the Arabidopsis root meristem. Expression values are shown for each of the separate tissue types.
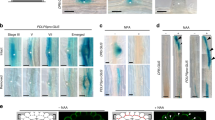
Extended Data Fig. 2 Characterization of BGLU44 expression and function.
a-c, Predicted expression of LOG4 and BGLU44 according to a single cell atlas of the Arabidopsis root meristem. Note the predicted co-expression in the feature plots for both genes. d-g, Expression of pBGLU44::nYFP/GUS in the dGR shoot and root meristems grown on mock medium and transferred to mock or 10 μM DEX for 24h. h, Inflorescence stem and silique phenotypes in the indicated genotypes. i, Relative expression levels as determined by RT-qPCR of BGLU44 and LOG4 in the indicated lines. Boxes represent the 1st and 3rd quartile and the center line represents the median. Asterisks in graphs indicate significance values as determined by standard two-sided t-tests. Black lines indicates mean values and grey boxes indicate data ranges. j, Expression of pRPS5A::BGLU44:YFP in the root meristem grown on mock medium showing that the BGLU44:YFP fusion protein is expressed. k-o, Phenotypes and quantification of the number of vascular cell files in the mentioned genotypes. Boxes represent the 1st and 3rd quartile and the center line represents the median. Statistics in n are performed using pairwise comparisons using a standard two sided t-test (p-value<0,05). p-s, Confocal images of optical cross sections of root meristems expressing pTCSn::ntdTomato reporter in the mentioned genotypes. Scale bars are 10 μm. t-w, Root hair phenotypes of the mentioned genotypes. In all panels, n represents the number of replicates or data points; all data and statistics are summarized in Supplementary Data 4. The experiments in j was repeated three times independently, with similar results.
Extended Data Fig. 3 BGLU44 and SHR are direct TMO5/LHW target genes.
a, b, ChIP-RT-qPCR experiments using anti-GR antibody on Col-0 and dGR lines. Graphs show the values of two biological repeats; black lines indicates mean values and grey boxes indicate data ranges. n = 2 biological replicates. Asterisks indicate significance values as determined by standard two-sided t-tests. c, Relative expression levels as determined by RT-qPCR of LOG4, BGLU44, CKX3 and SHR in the indicated lines in presence or absence of 10μM DEX. n = 3 biological replicates. Asterisks in graphs indicate significance values as determined by standard two-sided t-tests. Black lines indicates mean values and grey boxes indicate data ranges. All Boxes represent the 1st and 3rd quartile and the center line represents the median. In all panels, n represents the.
Extended Data Fig. 4 BGLU44 mutant generation and protein purification.
a, Position of the deletion in the bglu44 mutant line. b, PCR validation of the bglu44 mutant and crosses with log4 and log34 mutants. c, quantification of the number of vascular cell files in the indicated lines. Boxes represent the 1st and 3rd quartile and the center line represents the median. n indicate independent plants, the data and statistics are summarized in Supplementary Data 4. Asterisks in graphs indicate significance values as determined by standard two-sided t-tests (*:p-value<0.05; **: p-value<0.01). d, Coomassie gel showing subsequent fractions of purified BGLU44 protein (arrowhead) after ion exchange chromatography. The experiments in b and d were repeated three times independently, with similar results.
Extended Data Fig. 5 Single cell transcriptional changes in root meristem cells upon cytokinin treatment.
a-c. UMAP representation of the merged (a), mock (b) and cytokinin (c) datasets in which cell identities are shown by clusters of different colours. In panels b-c, the cells from the other dataset are indicated in grey to show contribution to each of the cell clusters. LRC: lateral root cap; PX: protoxylem; MX: metaxylem; SE: sieve element; CC: companion cell. The most central dark blue cell cluster is the initial cell cluster. d. Relative cell identity abundance in the mock and cytokinin (CK) datasets in comparison to a previously published root meristem atlas8. e,f. Violin plots showing normalized gene expression of CKX3 (e) and CKX5 (f) by cell type and split by treatment: left side of the violin is mock dataset in green; right side of the violin is CK-treated dataset in red).
Extended Data Fig. 6 CKX3 and CKX5 are required for normal vascular development.
a,b, Predicted expression of CKX3 and CKX5 according to a previously published single cell atlas of the Arabidopsis root meristem18. Compare to Supplementary Fig. 2a for cell identities. c,d, Expression of pCKX5::nYFP/GUS in the root meristem grown on mock medium. e-k, Microscopic images of xylem differentiation in the mentioned genotypes. p: protoxylem, m: metaxylem. l, Quantification of the different classes of xylem phenotypes shown in panels e-k. m-p, Confocal microscopy images of cross sections through the root meristem showing the vascular cell file number phenotype and quantification of the mentioned genotypes. q, Quantification of the number of vascular cell files shown in panels m-p. Lower-case letters on top of the boxplots indicate significantly different groups as determined by one-way ANOVA with post-hoc Tukey HSD testing (p<0.001). Boxes represent the 1st and 3rd quartile and the center line represents the median. Grey dotted line separates data from different experiments. r, RT-qPCR analysis showing the relative expression levels of CKX3 on wild type Col-0 and tmo5 triple mutant backgrounds when grown on mock (0h) or 100 nM or 5 μM BA for the indicated time. Boxes represent the 1st and 3rd quartile and the center line represents the median. n = 3 biological replicates. Asterisks in graphs indicate significance values as determined by standard two-sided t-tests. Black lines indicates mean values and grey boxes indicate data ranges. In all panels, n represents the number of replicates or data points; all data and statistics are summarized in Supplementary Data 4. The experiments in c-d were repeated three times independently, with similar results.
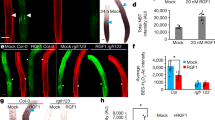
Extended Data Fig. 7 SHR acts downstream of TMO5/LHW.
a,b, Confocal images of root meristems expressing pTCSn::ntdTomato reporter in Col-0 and shr-2 backgrounds. c, Quantification of the pTCSn::ntdTomato mean pixel intensity in the mentioned genotypes. Boxes represent the 1st and 3rd quartile and the center line represents the median. n indicate independent plants. Lower-case letters on top of the boxplots indicate significantly different groups as determined by one-way ANOVA with post-hoc Tukey HSD testing (p<0.001). d-g, Expression of pSHR::SHR:GFP in the dGR root meristem grown on mock medium and transferred to mock or 10 μM DEX for 24 h. h, Relative expression levels of SHR in wild type (Col-0), TMO5/LHW misexpression, and tmo5, tmo5 tmo5like1 (t5 t5l1) and tmo5 tmo5like1 tmo5like3 (t5 t5l1 t5l3) mutant backgrounds. Asterisks indicate significance values as determined by standard two-sided t-tests. Boxes represent the 1st and 3rd quartile and the center line represents the median. n = 3 biological replicates. Black lines indicates mean values and grey boxes indicate data ranges. i-l, Microscopic images of xylem differentiation in the mentioned genotypes. p: protoxylem, m: metaxylem. m, Quantification of the different classes of xylem phenotypes shown in panels i-l. n-o, Expression of pSCR::H2B:nYFP in the dGR root meristem grown on mock medium and transferred to mock or 10 μM DEX for 24h. p-q, Expression of pmiR165a::nYFP/GUS in the dGR root meristem grown on mock medium and transferred to mock or 10 μM DEX for 24h. r-s, Relative expression levels of SCR and miR165a in wild type (Col-0), TMO5/LHW misexpression, and tmo5, tmo5 tmo5like1 (t5 t5l1) and tmo5 tmo5like1 tmo5like3 (t5 t5l1 t5l3) mutant backgrounds. Boxes represent the 1st and 3rd quartile and the center line represents the median. n = 3 biological replicates. Asterisks indicate significance values as determined by standard two-sided t-tests. Black lines indicates mean values and grey boxes indicate data ranges. In panels a-b, d-g, i-l and n-q, scale bars are 50 μm. In all panels, n represents the number of replicates or data points; all data and statistics are summarized in Supplementary Data 4. The experiments in d-g, n-q were repeated three times independently, with similar results.
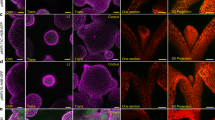
Extended Data Fig. 8 Xylem expressed SHR is sufficient to rescue the shr-2 mutant.
a, Seedling phenotypes of the lines indicated. Note the rescue of root length in the pTMO5::SHR:GFP lines in shr-2 mutant background. b-g, Confocal images of root meristems of the indicated lines counterstained with PI (red). Scale bars are 10 μm. Panels c, e and g are optical cross sections of the images in b, d and f, respectively. Asterisks indicate endodermal cells; x marks mixed ground tissue identity. The experiments here were repeated three times independently, with similar results.
Extended Data Fig. 9 A model for spatiotemporal control of cytokinin levels.
Auxin induces TMO5 expression and as such triggers activation of the TMO5/LHW heterodimer complex. Sequentially, LOG3/4, BGLU44 and SHR expression are induced. SHR itself binds to the promoter region of CKX3 to activate its expression. As such, the same transcription factor complex sequentially increases levels of active cytokinin via LOG3/4 and BGLU44; and represses levels active cytokinin via SHR-CKX3. The combined activity balances levels of active cytokinin to allow normal vascular proliferation and differentiation. The left side of the figures shows the temporal aspect, while the right side shows the spatial aspect.
Supplementary information
Supplementary Information
Methods: Mathematical Modelling, Supplementary Figs. 1–3 and captions.
Supplementary Table 1
Co-expression analysis.
Supplementary Table 2
scRNAseq dataset.
Supplementary Table 3
Primer list.
Source data
Source Data Fig. 1
Source data and statistics.
Source Data Fig. 2
Source data and statistics.
Source Data Fig. 3
Source data and statistics.
Source Data Fig. 4
Source data and statistics.
Source Data Extended Data Fig. 2
Source data and statistics.
Source Data Extended Data Fig. 3
Source data and statistics.
Source Data Extended Data Fig. 4
Source data and statistics.
Source Data Extended Data Fig. 6
Source data and statistics.
Source Data Extended Data Fig. 7
Source data and statistics.
Rights and permissions
About this article
Cite this article
Yang, B., Minne, M., Brunoni, F. et al. Non-cell autonomous and spatiotemporal signalling from a tissue organizer orchestrates root vascular development. Nat. Plants 7, 1485–1494 (2021). https://doi.org/10.1038/s41477-021-01017-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-01017-6
This article is cited by
-
How to explore what is hidden? A review of techniques for vascular tissue expression profile analysis
Plant Methods (2023)
-
Application of single-cell multi-omics approaches in horticulture research
Molecular Horticulture (2023)
-
A redundant transcription factor network steers spatiotemporal Arabidopsis triterpene synthesis
Nature Plants (2023)
-
microRNA165 and 166 modulate response of the Arabidopsis root apical meristem to salt stress
Communications Biology (2023)
-
A SHORTROOT-Mediated Transcriptional Regulatory Network for Vascular Development in the Arabidopsis Shoot
Journal of Plant Biology (2022)