Abstract
Microbial symbioses can mitigate drought stress in crops but harnessing these beneficial interactions will require an in-depth understanding of root microbiome responses to drought cycles. Here, by detailed temporal characterization of root-associated microbiomes of rice plants during drought stress and recovery, we find that endosphere communities remained compositionally altered after rewatering, with prolonged droughts leading to decreased resilience. Several endospheric Actinobacteria were significantly enriched during drought and for weeks after rewatering. Notably, the most abundant endosphere taxon during this period was a Streptomyces, and a corresponding isolate promoted root growth. Additionally, drought stress disrupted the temporal dynamics of late-colonizing microorganisms, permanently altering the normal successional trends of root microbiota. These findings reveal that severe drought results in enduring impacts on rice root microbiomes, including enrichment of taxonomic groups that could shape the recovery response of the host, and have implications relevant to drought protection strategies using root microbiota.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw reads have been deposited in the Sequene Read Archive under BioProject PRJNA551661. The SLBN-177 sequence has been deposited within the 100K Project BioProject PRJNA743693 with accession number SRR15049341. The Greengenes database (v.13_8) can be downloaded from http://qiime.org/home_static/dataFiles.html. The SILVA database (v.132) can be downloaded from https://www.arb-silva.de/download/archive/qiime/.
Code availability
All scripts and intermediate files are available in GitHub (https://github.com/cmsantosm/RiceDroughtRecovery).
References
Lesk, C., Rowhani, P. & Ramankutty, N. Influence of extreme weather disasters on global crop production. Nature 529, 84–87 (2016).
Zhang, J. et al. Effect of drought on agronomic traits of rice and wheat: a meta-analysis. Int. J. Environ. Res. Public Health 15, 839 (2018).
Hirasawa, T., in Genetic Improvement of Rice for Water-Limited Environments (eds Ito, O, O’Toole, J. C. & Hardy, B.) 89–98 (International Rice Research Institute, 1999).
Pandey, V. & Shukla, A. Acclimation and tolerance strategies of rice under drought stress. Rice Sci. 22, 147–161 (2015).
Compant, S., van der Heijden, M. G. A. & Sessitsch, A. Climate change effects on beneficial plant-microorganism interactions. FEMS Microbiol. Ecol. 73, 197–214 (2010).
de Vries, F. T., Griffiths, R. I., Knight, C. G., Nicolitch, O. & Williams, A. Harnessing rhizosphere microbiomes for drought-resilient crop production. Science 368, 270–274 (2020).
Busby, P. E. et al. Research priorities for harnessing plant microbiomes in sustainable agriculture. PLoS Biol. 15, e2001793 (2017).
Santos-Medellín, C., Edwards, J., Liechty, Z., Nguyen, B. & Sundaresan, V. Drought stress results in a compartment-specific restructuring of the rice root-associated microbiomes. mBio 8, e00764-17 (2017).
Naylor, D., DeGraaf, S., Purdom, E. & Coleman-Derr, D. Drought and host selection influence bacterial community dynamics in the grass root microbiome. ISME J. https://doi.org/10.1038/ismej.2017.118 (2017).
Fitzpatrick, C. R. et al. Assembly and ecological function of the root microbiome across angiosperm plant species. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.1717617115 (2018).
Edwards, J. A. et al. Compositional shifts in root-associated bacterial and archaeal microbiota track the plant life cycle in field-grown rice. PLoS Biol. 16, e2003862 (2018).
Zhang, J. et al. Root microbiota shift in rice correlates with resident time in the field and developmental stage. Sci. China Life Sci. 61, 613–621 (2018).
Xu, L. et al. Drought delays development of the sorghum root microbiome and enriches for monoderm bacteria. Proc. Natl Acad. Sci. USA 115, E4284–E4293 (2018).
Liechty, Z. et al. Comparative analysis of root microbiomes of rice cultivars with high and low methane emissions reveals differences in abundance of methanogenic archaea and putative upstream fermenters. mSystems 5, e00897-19 (2020).
Rong, X. & Huang, Y. Taxonomic evaluation of the Streptomyces griseus clade using multilocus sequence analysis and DNA–DNA hybridization, with proposal to combine 29 species and three subspecies as 11 genomic species. Int. J. Syst. Evol. Microbiol. 60, 696–703 (2010).
Lin, L. & Xu, X. Indole-3-acetic acid production by endophytic Streptomyces sp. En-1 isolated from medicinal plants. Curr. Microbiol. 67, 209–217 (2013).
Legault, G. S., Lerat, S., Nicolas, P. & Beaulieu, C. Tryptophan regulates thaxtomin A and indole-3-acetic acid production in Streptomyces scabiei and modifies its interactions with radish seedlings. Phytopathology 101, 1045–1051 (2011).
Guo, J. et al. Seed-borne, endospheric and rhizospheric core microbiota as predictor for plant functional traits across rice cultivars are dominated by deterministic processes. New Phytol. https://doi.org/10.1111/nph.17297 (2021).
de Vries, F. T. et al. Soil bacterial networks are less stable under drought than fungal networks. Nat. Commun. 9, 3033 (2018).
de Vries, F. T. & Shade, A. Controls on soil microbial community stability under climate change. Front. Microbiol. 4, 265 (2013).
Borken, W. & Matzner, E. Reappraisal of drying and wetting effects on C and N mineralization and fluxes in soils. Glob. Change Biol. 15, 808–824 (2009).
Lueders, T. & Friedrich, M. W. Effects of amendment with ferrihydrite and gypsum on the structure and activity of methanogenic populations in rice field soil. Appl. Environ. Microbiol. 68, 2484–2494 (2002).
Linquist, B. A. et al. Reducing greenhouse gas emissions, water use, and grain arsenic levels in rice systems. Glob. Change Biol. 21, 407–417 (2015).
Speirs, L. B. M., Rice, D. T. F., Petrovski, S. & Seviour, R. J. The phylogeny, biodiversity, and ecology of the chloroflexi in activated sludge. Front. Microbiol. 10, 2015 (2019).
Thomas, S. H. et al. The mosaic genome of Anaeromyxobacter dehalogenans strain 2CP-C suggests an aerobic common ancestor to the delta-proteobacteria. PLoS ONE 3, e2103 (2008).
Yang, T. H., Coppi, M. V., Lovley, D. R. & Sun, J. Metabolic response of Geobacter sulfurreducens towards electron donor/acceptor variation. Microb. Cell Fact. 9, 90 (2010).
Keller, K. L. & Wall, J. D. Genetics and molecular biology of the electron flow for sulfate respiration in desulfovibrio. Front. Microbiol. 2, 135 (2011).
Zhalnina, K. et al. Dynamic root exudate chemistry and microbial substrate preferences drive patterns in rhizosphere microbial community assembly. Nat. Microbiol. https://doi.org/10.1038/s41564-018-0129-3 (2018).
Williams, A. & de Vries, F. T. Plant root exudation under drought: implications for ecosystem functioning. New Phytol. 225, 1899–1905 (2020).
Vries, F. T. et al. Changes in root-exudate-induced respiration reveal a novel mechanism through which drought affects ecosystem carbon cycling. New Phytol. 224, 132–145 (2019).
Casartelli, A. et al. Exploring traditional aus-type rice for metabolites conferring drought tolerance. Rice 11, 9 (2018).
Pérez-Jaramillo, J. E. et al. Linking rhizosphere microbiome composition of wild and domesticated Phaseolus vulgaris to genotypic and root phenotypic traits. ISME J. https://doi.org/10.1038/ismej.2017.85 (2017).
Kang, D.-J. & Futakuchi, K. Effect of moderate drought-stress on flowering time of interspecific hybrid progenies (Oryza sativa L. × Oryza glaberrima Steud.). J. Crop Sci. Biotechnol. 22, 75–81 (2019).
Guo, X. et al. Host-associated quantitative abundance profiling reveals the microbial load variation of root microbiome. Plant Commun. 1, 100003 (2020).
Varoquaux, N. et al. Transcriptomic analysis of field-droughted sorghum from seedling to maturity reveals biotic and metabolic responses. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.1907500116 (2019).
Li, P. et al. Physiological and transcriptome analyses reveal short-term responses and formation of memory under drought stress in rice. Front. Genet. 10, 55 (2019).
Vandenkoornhuyse, P., Quaiser, A., Duhamel, M., Le Van, A. & Dufresne, A. The importance of the microbiome of the plant holobiont. New Phytol. 206, 1196–1206 (2015).
Toju, H. et al. Core microbiomes for sustainable agroecosystems. Nat. Plants 4, 247–257 (2018).
Shade, A. & Stopnisek, N. Abundance-occupancy distributions to prioritize plant core microbiome membership. Curr. Opin. Microbiol. 49, 50–58 (2019).
Suralta, R. R. et al. Plasticity in nodal root elongation through the hardpan triggered by rewatering during soil moisture fluctuation stress in rice. Sci. Rep. 8, 4341 (2018).
Hamedi, J. & Mohammadipanah, F. Biotechnological application and taxonomical distribution of plant growth promoting actinobacteria. J. Ind. Microbiol. Biotechnol. 42, 157–171 (2015).
Vurukonda, S. S. K. P., Vardharajula, S., Shrivastava, M. & SkZ, A. Enhancement of drought stress tolerance in crops by plant growth promoting rhizobacteria. Microbiol. Res. 184, 13–24 (2016).
Aznar, A. & Dellagi, A. New insights into the role of siderophores as triggers of plant immunity: what can we learn from animals? J. Exp. Bot. 66, 3001–3010 (2015).
Viaene, T., Langendries, S., Beirinckx, S., Maes, M. & Goormachtig, S. Streptomyces as a plant’s best friend? FEMS Microbiol. Ecol. https://doi.org/10.1093/femsec/fiw119 (2016).
Meena, K. K. et al. Abiotic stress responses and microbe-mediated mitigation in plants: the omics strategies. Front. Plant Sci. 8, 172 (2017).
Mukamuhirwa, A. et al. Effect of intermittent drought on grain yield and quality of rice (Oryza sativa L.) grown in Rwanda. J. Agro Crop Sci. 206, 252–262 (2020).
Fleta-Soriano, E. & Munné-Bosch, S. Stress memory and the inevitable effects of drought: a physiological perspective. Front. Plant Sci. 7, 143 (2016).
Ding, Y., Fromm, M. & Avramova, Z. Multiple exposures to drought ‘train’ transcriptional responses in Arabidopsis. Nat. Commun. 3, 740 (2012).
de la Fuente Cantó, C. et al. An extended root phenotype: the rhizosphere, its formation and impacts on plant fitness. Plant J. 103, 951–964 (2020).
Kittas, C., Bartzanas, T. & Jaffrin, A. Temperature gradients in a partially shaded large greenhouse equipped with evaporative cooling pads. Biosyst. Eng. 85, 87–94 (2003).
Edwards, J. et al. Soil domestication by rice cultivation results in plant–soil feedback through shifts in soil microbiota. Genome Biol. 20, 221 (2019).
Edwards, J., Santos-Medellín, C. & Sundaresan, V. Extraction and 16S rRNA sequence analysis of microbiomes associated with rice roots. Bio. Protoc. 8, e2884 (2018).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl Acad. Sci. USA 108, 4516–4522 (2011).
Masella, A. P., Bartram, A. K., Truszkowski, J. M., Brown, D. G. & Neufeld, J. D. PANDAseq: paired-end assembler for illumina sequences. BMC Bioinform. 13, 31 (2012).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Weimer, B. C. 100K Pathogen Genome Project. Genome Announc. 5, e00594-17 (2017).
Kong, N. et al. Draft genome sequences of 1,183 Salmonella strains from the 100K Pathogen Genome Project. Genome Announc. 5, e00518–17 (2017).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Medema, M. H. et al. antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. 39, W339–W346 (2011).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2018); https://www.R-project.org/
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
McMurdie, P. J. & Holmes, S. Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput. Biol. 10, e1003531 (2014).
Paradis, E., Claude, J. & Strimmer, K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004).
Oksanen, J. et al. vegan: Community Ecology Package (2018).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, 2016).
Kuznetsova, A., Brockhoff, P. B. & Christensen, R. H. B. lmerTest package: tests in linear mixed effects models. J. Stat. Softw. 82, 13 (2017).
Lenth, R., Singmann, H., Love, J., Buerkner, P. & Herve, M. Emmeans: estimated marginal means, aka least-squares means. R package v.1, 3 (R Foundation for Statistical Computing, 2018).
Kassambara, A. Rstatix: pipe-friendly framework for basic statistical tests. R package v.0.6.0 (R Foundation for Statistical Computing, 2020).
Graves, S., Piepho, H.-P., Selzer, L. & Dorai-Raj, S. multcompView: visualizations of paired comparisons. R package v.0.1-7 (R Foundation for Statistical Computing, 2015).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Liaw, A. & Wiener, M. Classification and regression by randomForest. R. News 2, 18–22 (2002).
Subramanian, S. et al. Persistent gut microbiota immaturity in malnourished Bangladeshi children. Nature 510, 417–421 (2014).
Acknowledgements
We thank R. Melnyk for helpful advice regarding assembly, annotation and interpretation of the SLBN-177 genome, and A. Bennett for helpful suggestions. This project was supported by grants to V.S. from the National Science Foundation (no. IOS 1444974) and the United States Department of Agriculture (no. NIFA 2021-67013-34607 and Agricultural Experiment Station project no. CA-D-XXX-6973-H). C.S.M. acknowledges support from the University of California Institute for Mexico, Consejo Nacional de Ciencia y Tecnología and Secretaría de Educación Pública (Mexico). C.S.M. and Z.L. also acknowledge partial support from the Elsie Taylor Stocking Memorial Research Fellowship and the Henry A. Jastro Graduate Research Award. J.E. is supported by USDA National Institute of Food and Agriculture Postdoctoral Fellowship (grant no. 2019-67012-2971/project accession no. 1019437). Sequencing was performed by the DNA Technologies and Expression Analysis Cores at the UC Davis Genome Center supported by the National Institutes of Health Shared Instrumentation grant no. 1S10OD010786-01.
Author information
Authors and Affiliations
Contributions
C.S.-M., Z.L. and V.S. conceptualized the study. C.S.M., Z.L., J.E. and B.N. performed the experiments. B.H. and B.C.W. generated the SLBN-177 genome sequence and reviewed the paper. C.S.-M. and Z.L. analysed the data. C.S.-M., Z.L., J.E. and V.S. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Alex Williams, Maggie Wagner and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Compartments harbor compositionally distinct microbial communities.
a, Principal coordinates analysis (PCoA) performed on weighted UniFrac distances across the whole dataset. Colours indicate compartment. b, Distribution of rhizosphere and endosphere within-group distances, that is distances between samples within each treatment and time point combination. The large variation displayed by endospheres in panels a and b is, in part, a result of the strong effect that drought treatments had on this compartment. c-f, PCoA performed on the rhizosphere (c & e) and endosphere (d & f) subsets. In panels c and d, colours indicate plant age. In panels e and f, data points are faceted by plant age and coloured by drought treatment.
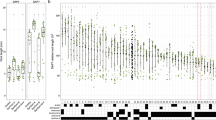
Extended Data Fig. 2 Complete set of drought-responsive OTUs.
Heatmap displaying the log2 fold-changes between water controls and drought treatments: depletion under drought tends towards green while enrichment under drought tends toward brown. Each row represents a differentially abundant OTU detected as significant (Wald test, FDR < 0.05) in at least one pair-wise comparison. Horizontal facets indicate each of the modules detected through hierarchical clustering in the rhizosphere (RS) and endosphere (ES). Clusters shown in Fig. 2b are highlighted in black. Vertical dotted lines delimit the periods of suspended irrigation for each of the drought treatments.
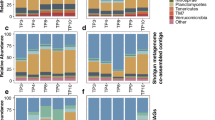
Extended Data Fig. 3 Modules of drought responsive OTUs exhibit a strong taxonomic signature.
Classification of the differentially abundant OTUs in each of the distinct drought modules detected through hierarchical clustering. Each tile indicates the number of classified OTUs in a particular module, while the colour represents membership to a specific Phylum / Proteobacteria class. Orders with only one representative have been excluded to ease visualization.
Extended Data Fig. 4 OTU 1037355 displays reproducible trends in an independent experiment.
a, Timeline of the watering regimes followed by control (WC) and drought-stressed (DS) plants. Horizontal lines represent the watering status during the experiment: solid segments indicate periods of constant irrigation while dotted segments indicate periods of suspended irrigation. Upside down triangles mark each of 5 collection time points. b, Ranked relative abundances of individual community members throughout time. Each ribbon represents a single OTU in the community: for each time point, width indicates its relative abundance while the position across the y axis indicates its rank within the community. The most abundant member of the semipersistent enrichment module, Streptomyces sp. (OTU ID: 1037355), is highlighted. In all panels, the vertical dotted lines delimit the periods of suspended irrigation in each of the drought treatments. c,d, Beta-diversity patterns in the rhizosphere (c) and endosphere (d) communities. In both cases, the y axis displays the position of each sample across the first principal coordinate (PCo) from a weighted UniFrac PCo analysis and the x axis displays the age of the plant at the moment of sample collection. In all panels, the vertical dotted lines delimit the periods of suspended irrigation for the drought treatment.
Extended Data Fig. 5 OTU 1037355 is highly occurring and displays a significant drought enrichment across compositionally distinct soils.
a, Occupancy-abundance curves for rhizosphere and endosphere communities of rice plants grown in three California soils (Arbuckle, Biggs, and Davis), and one Arkansas field. The x axis displays the log-transformed mean relative abundance of each OTU while the y axis displays the percent of samples in which each OTU was detected. OTU 1037355 is highlighted in orange. b, Relative abundance of OTU 1037355 in rhizosphere and endosphere communities of 49-day-old rice plants grown under well irrigated (WC) or drought-stressed (DS) conditions. Asterisks on top indicate a significant difference (PFDR < 0.001) between WC and DS treatments. Statistical significance was determined by negative binomial generalized linear models and pairwise Wald tests (two-sided) corrected with the Benjamini-Hochberg procedure.
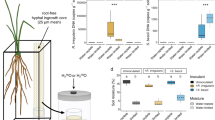
Extended Data Fig. 6 Other phenotypic traits were not impacted by Streptomyces sp. SLBN-177.
a, Distribution of measured plant phenotypes (number of roots, root weight, shoot weight, and shoot length) across microbial treatments and watering regimes. b, Protein alignment of putative iaaM genes from SLBN-177, S. coelicolor, S. scabiei, and S. sp ADI96-02 (refs. 16,17). Black lines indicate an amino acid mismatch with a negative score on the BLOSUM62 matrix, and dark grey bars represent a mismatch with a positive score. The vertical black dashed lines indicate the bounds of the amino oxidase functional domain. c, Relative abundances of inoculated isolates in the endospheres of rice plants. Reads identified as mitochondria or chloroplast (collapsed as organellar reads and represented in black) were not discarded in order to measure the degree of colonization. Reads classified as OTUs other than 1037355 (SLBN-177) and 1108350 (SLBN-111) were collapsed and are represented in gray.
Extended Data Fig. 7 Random forests can identify age discriminant OTUs.
a, Cross validation error as a function of the number of OTUs used in each model. For both compartments, the lowest error was detected when using the 65 most important taxa. b, Overlap between the age-discriminant OTUs and the drought-responsive OTUs detected in each module (Fig. 2b). c, Hierarchical clustering of the relative abundances of the age-discriminant OTUs. The heatmap displays the z-transformed mean relative abundances of each OTU across drought treatments and time points. The colours at the left end of each vertical facet indicate the longitudinal trends exhibited by each taxa. In all panels, the vertical dotted lines delimit the periods of suspended irrigation for the drought treatment.
Extended Data Fig. 8 Taxonomic classification of the age-discriminant OTUs in each of the longitudinal trends.
Each tile indicates the number of classified OTUs in a particular module, while the colour represents membership to a specific Phylum / Proteobacteria class.
Extended Data Fig. 9 Drought stress delayed transition to flowering.
a, Developmental growth stages of well-watered and drought-stressed rice plants through the experiment. Photos of each sample were taken at each time point and defined as either pre-panicle emergence (no portion of the panicle visible), panicle emergence (panicle partially emerged from flag leaf), anthesis (panicle fully emerged and anthers visible), grain filling (panicles bent over instead of standing upright), or maturity (flower colour appears yellow). b, Developmental growth stage and microbiome age predictions of rhizosphere and endosphere communities across drought treatments (D1, D2, and D3). The dashed curve represents the baseline microbiome development under well-irrigated conditions and was calculated by fitting a loess curve between the predicted microbiome age and the chronological plant age in the control (WC) test set.
Extended Data Fig. 10 Calculation of relative microbiome maturity of drought-stressed samples.
a, After applying the sparse random forest models to the validating set of well-watered plants, a loess curve (dashed green line) was fit between host chronological age (x axis) and microbiome predicted age (y axis). b, Using the fitted loess curve as a baseline of microbiome development, relative microbiome maturity was estimated by calculating the difference between the predicted microbiome age of an individual sample and the corresponding baseline microbiome age of well-watered plants. c, Using the approach described in B, relative microbiome maturity was calculated for all drought-stressed samples.
Supplementary information
Supplementary Information
Supplementary Figs. 1–5.
Supplementary Table
Supplementary Tables 1–10.
Rights and permissions
About this article
Cite this article
Santos-Medellín, C., Liechty, Z., Edwards, J. et al. Prolonged drought imparts lasting compositional changes to the rice root microbiome. Nat. Plants 7, 1065–1077 (2021). https://doi.org/10.1038/s41477-021-00967-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-00967-1
This article is cited by
-
Host genotype-specific rhizosphere fungus enhances drought resistance in wheat
Microbiome (2024)
-
Dynamic root microbiome sustains soybean productivity under unbalanced fertilization
Nature Communications (2024)
-
Chromosomal barcodes for simultaneous tracking of near-isogenic bacterial strains in plant microbiota
Nature Microbiology (2024)
-
Belowground microbiota associated with the progression of Verticillium wilt of smoke trees
Plant and Soil (2024)
-
Effect of drought stress on symbiotic nitrogen fixation, soil nitrogen availability and soil microbial diversity in forage legumes
Plant and Soil (2024)