Abstract
Angiosperm leaves show extensive shape diversity and are broadly divided into two forms; simple leaves with intact lamina and compound leaves with lamina dissected into leaflets. The mechanistic basis of margin dissection and leaflet initiation has been inferred primarily by analysing compound-leaf architecture, and thus whether the intact lamina of simple leaves has the potential to initiate leaflets upon endogenous gene inactivation remains unclear. Here, we show that the CINCINNATA-like TEOSINTE BRANCHED1, CYCLOIDEA, PROLIFERATING CELL FACTORS (CIN-TCP) transcription factors activate the class II KNOTTED1-LIKE (KNOX-II) genes and the CIN-TCP and KNOX-II proteins together redundantly suppress leaflet initiation in simple leaves. Simultaneous downregulation of CIN-TCP and KNOX-II in Arabidopsis leads to the reactivation of the stemness genes KNOX-I and CUPSHAPED COTYLEDON (CUC) and triggers ectopic organogenesis, eventually converting the simple lamina to a super-compound form that appears to initiate leaflets indefinitely. Thus, a conserved developmental mechanism promotes simple leaf architecture in which CIN-TCP–KNOX-II forms a strong differentiation module that suppresses the KNOX-I-CUC network and leaflet initiation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
There are no restrictions on data availability. The transcriptomic raw data used in this study have been deposited to the National Centre for Biotechnology Information (NCBI) and Gene Expression Omnibus (GEO) database under the accession numbers GSE174702 and PRJNA734903. Source data are provided with this paper.
References
Efroni, I., Eshed, Y. & Lifschitz, E. Morphogenesis of simple and compound leaves: a critical review. Plant Cell 22, 1019–1032 (2010).
Vlad, D. et al. Leaf shape evolution through duplication, regulatory diversification, and loss of a homeobox gene. Science 343, 780–783 (2014).
Bar, M. & Ori, N. Leaf development and morphogenesis. Development 141, 4219–4230 (2014).
Hay, A. & Tsiantis, M. The genetic basis for differences in leaf form between Arabidopsis thaliana and its wild relative Cardamine hirsuta. Nat. Genet. 38, 942–947 (2006).
Nikolov, L. A., Runions, A., Das Gupta, M. & Tsiantis, M. Leaf development and evolution. Curr Top Dev Biol 131, 109–139 (2019).
Blein, T. et al. A conserved molecular framework for compound leaf development. Science 322, 1835–1839 (2008).
Barkoulas, M., Hay, A., Kougioumoutzi, E. & Tsiantis, M. A developmental framework for dissected leaf formation in the Arabidopsis relative Cardamine hirsuta. Nat. Genet. 40, 1136–1141 (2008).
Hareven, D., Gutfinger, T., Parnis, A., Eshed, Y. & Lifschitz, E. The making of a compound leaf: genetic manipulation of leaf architecture in tomato. Cell 84, 735–744 (1996).
Vollbrecht, E., Veit, B., Sinha, N. & Hake, S. The developmental gene Knotted-1 is a member of a maize homeobox gene family. Nature 350, 241–243 (1991).
Long, J. A., Moan, E. I., Medford, J. I. & Barton, M. K. A member of the KNOTTED class of homeodomain proteins encoded by the STM gene of Arabidopsis. Nature 379, 66–69 (1996).
Hay, A., Barkoulas, M. & Tsiantis, M. ASYMMETRIC LEAVES1 and auxin activities converge to repress BREVIPEDICELLUS expression and promote leaf development in Arabidopsis. Development 133, 3955–3961 (2006).
Bharathan, G. et al. Homologies in leaf form inferred from KNOXI gene expression during development. Science 296, 1858–1860 (2002).
Shani, E. et al. Stage-specific regulation of Solanum lycopersicum leaf maturation by class 1 KNOTTED1-LIKE HOMEOBOX proteins. Plant Cell 21, 3078–3092 (2009).
Sinha, N. R., Williams, R. E. & Hake, S. Overexpression of the maize homeo box gene, KNOTTED-1, causes a switch from determinate to indeterminate cell fates. Genes Dev. 7, 787–795 (1993).
Kierzkowski, D. et al. A growth-based framework for leaf shape development and diversity. Cell 177, 1405–1418 (2019).
Vuolo, F. et al. Coupled enhancer and coding sequence evolution of a homeobox gene shaped leaf diversity. Genes Dev. 30, 2370–2375 (2016).
Bilsborough, G. D. et al. Model for the regulation of Arabidopsis thaliana leaf margin development. Proc. Natl Acad. Sci. USA 108, 3424–3429 (2011).
Rubio-Somoza, I. et al. Temporal control of leaf complexity by miRNA-regulated licensing of protein complexes. Curr. Biol. 24, 2714–2719 (2014).
Aida, M., Ishida, T., Fukaki, H., Fujisawa, H. & Tasaka, M. Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant. Plant Cell 9, 841–857 (1997).
Nikovics, K. et al. The balance between the MIR164A and CUC2 genes controls leaf margin serration in Arabidopsis. Plant Cell 18, 2929–2945 (2006).
Hasson, A. et al. Evolution and diverse roles of the CUP-SHAPED COTYLEDON genes in Arabidopsis leaf development. Plant Cell 23, 54–68 (2011).
Takada, S., Hibara, K., Ishida, T. & Tasaka, M. The CUP-SHAPED COTYLEDON1 gene of Arabidopsis regulates shoot apical meristem formation. Development 128, 1127–1135 (2001).
Furumizu, C., Alvarez, J. P., Sakakibara, K. & Bowman, J. L. Antagonistic roles for KNOX1 and KNOX2 genes in patterning the land plant body plan following an ancient gene duplication. PLoS Genet. 11, e1004980 (2015).
Nath, U., Crawford, B. C., Carpenter, R. & Coen, E. Genetic control of surface curvature. Science 299, 1404–1407 (2003).
Palatnik, J. F. et al. Control of leaf morphogenesis by microRNAs. Nature 425, 257–263 (2003).
Ori, N. et al. Regulation of LANCEOLATE by miR319 is required for compound-leaf development in tomato. Nat. Genet. 39, 787–791 (2007).
Shleizer-Burko, S., Burko, Y., Ben-Herzel, O. & Ori, N. Dynamic growth program regulated by LANCEOLATE enables flexible leaf patterning. Development 138, 695–704 (2011).
Challa, K. R., Aggarwal, P. & Nath, U. Activation of YUCCA5 by the transcription factor TCP4 integrates developmental and environmental signals to promote hypocotyl elongation in Arabidopsis. Plant Cell 28, 2117–2130 (2016).
Challa, K. R., Rath, M. & Nath, U. The CIN-TCP transcription factors promote commitment to differentiation in Arabidopsis leaf pavement cells via both auxin-dependent and independent pathways. PLoS Genet. 15, e1007988 (2019).
Efroni, I., Blum, E., Goldshmidt, A. & Eshed, Y. A protracted and dynamic maturation schedule underlies Arabidopsis leaf development. Plant Cell 20, 2293–2306 (2008).
Donnelly, P. M., Bonetta, D., Tsukaya, H., Dengler, R. E. & Dengler, N. G. Cell cycling and cell enlargement in developing leaves of Arabidopsis. Dev. Biol. 215, 407–419 (1999).
Nag, A., King, S. & Jack, T. miR319a targeting of TCP4 is critical for petal growth and development in Arabidopsis. Proc. Natl Acad. Sci. USA 106, 22534–22539 (2009).
Aloni, R., Schwalm, K., Langhans, M. & Ullrich, C. I. Gradual shifts in sites of free-auxin production during leaf-primordium development and their role in vascular differentiation and leaf morphogenesis in Arabidopsis. Planta 216, 841–853 (2003).
Balkunde, R., Pesch, M. & Hulskamp, M. Trichome patterning in Arabidopsis thaliana from genetic to molecular models. Curr. Top. Dev. Biol. 91, 299–321 (2010).
Telfer, A., Bollman, K. M. & Poethig, R. S. Phase change and the regulation of trichome distribution in Arabidopsis thaliana. Development 124, 645–654 (1997).
Andriankaja, M. et al. Exit from proliferation during leaf development in Arabidopsis thaliana: a not-so-gradual process. Dev. Cell 22, 64–78 (2012).
Nakata, M. & Okada, K. The leaf adaxial–abaxial boundary and lamina growth. Plants 2, 174–202 (2013).
Zgurski, J. M., Sharma, R., Bolokoski, D. A. & Schultz, E. A. Asymmetric auxin response precedes asymmetric growth and differentiation of asymmetric leaf1 and asymmetric leaf2 Arabidopsis leaves. Plant Cell 17, 77–91 (2005).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Schmid, M. et al. A gene expression map of Arabidopsis thaliana development. Nat. Genet. 37, 501–506 (2005).
Tian, C. et al. A gene expression map of shoot domains reveals regulatory mechanisms. Nat. Commun. 10, 141 (2019).
Schommer, C., Debernardi, J. M., Bresso, E. G., Rodriguez, R. E. & Palatnik, J. F. Repression of cell proliferation by miR319-regulated TCP4. Mol. Plant 7, 1533–1544 (2014).
Aida, M., Ishida, T. & Tasaka, M. Shoot apical meristem and cotyledon formation during Arabidopsis embryogenesis: interaction among the CUP-SHAPED COTYLEDON and SHOOT MERISTEMLESS genes. Development 126, 1563–1570 (1999).
Belles-Boix, E. et al. KNAT6: an Arabidopsis homeobox gene involved in meristem activity and organ separation. Plant Cell 18, 1900–1907 (2006).
Jiang, W. et al. jaw-1D: a gain-of-function mutation responsive to paramutation-like induction of epigenetic silencing. J. Exp. Bot. 70, 459–468 (2019).
Khan, M. et al. Antagonistic interaction of BLADE-ON-PETIOLE1 and 2 with BREVIPEDICELLUS and PENNYWISE regulates Arabidopsis inflorescence architecture. Plant Physiol. 158, 946–960 (2012).
Koyama, T., Mitsuda, N., Seki, M., Shinozaki, K. & Ohme-Takagi, M. TCP transcription factors regulate the activities of ASYMMETRIC LEAVES1 and miR164, as well as the auxin response, during differentiation of leaves in Arabidopsis. Plant Cell 22, 3574–3588 (2010).
Li, Z., Li, B., Shen, W. H., Huang, H. & Dong, A. TCP transcription factors interact with AS2 in the repression of class-I KNOX genes in Arabidopsis thaliana. Plant J. 71, 99–107 (2012).
Koyama, T., Furutani, M., Tasaka, M. & Ohme-Takagi, M. TCP transcription factors control the morphology of shoot lateral organs via negative regulation of the expression of boundary-specific genes in Arabidopsis. Plant Cell 19, 473–484 (2007).
Yamaguchi, N., Winter, C. M., Wellmer, F. & Wagner, D. Identification of direct targets of plant transcription factors using the GR fusion technique. Methods Mol. Biol. 1284, 123–138 (2015).
Vadde, B. V. L., Challa, K. R. & Nath, U. The TCP4 transcription factor regulates trichome cell differentiation by directly activating GLABROUS INFLORESCENCE STEMS in Arabidopsis thaliana. Plant J. 93, 259–269 (2018).
Vadde, B. V. L., Challa, K. R., Sunkara, P., Hegde, A. S. & Nath, U. The TCP4 transcription factor directly activates TRICHOMELESS1 and 2 and suppresses trichome initiation. Plant Physiol. 181, 1587–1599 (2019).
Sarvepalli, K. & Nath, U. Hyper-activation of the TCP4 transcription factor in Arabidopsis thaliana accelerates multiple aspects of plant maturation. Plant J. 67, 595–607 (2011).
Aggarwal, P. et al. Identification of specific DNA binding residues in the TCP family of transcription factors in Arabidopsis. Plant Cell 22, 1174–1189 (2010).
Kubota, A. et al. TCP4-dependent induction of CONSTANS transcription requires GIGANTEA in photoperiodic flowering in Arabidopsis. PLoS Genet. 13, e1006856 (2017).
Waites, R., Selvadurai, H. R., Oliver, I. R. & Hudson, A. The PHANTASTICA gene encodes a MYB transcription factor involved in growth and dorsoventrality of lateral organs in Antirrhinum. Cell 93, 779–789 (1998).
Tsiantis, M., Schneeberger, R., Golz, J. F., Freeling, M. & Langdale, J. A. The maize rough sheath2 gene and leaf development programs in monocot and dicot plants. Science 284, 154–156 (1999).
Byrne, M. E. et al. Asymmetric leaves1 mediates leaf patterning and stem cell function in Arabidopsis. Nature 408, 967–971 (2000).
Guo, M., Thomas, J., Collins, G. & Timmermans, M. C. Direct repression of KNOX loci by the ASYMMETRIC LEAVES1 complex of Arabidopsis. Plant Cell 20, 48–58 (2008).
Gallois, J. L., Woodward, C., Reddy, G. V. & Sablowski, R. Combined SHOOT MERISTEMLESS and WUSCHEL trigger ectopic organogenesis in Arabidopsis. Development 129, 3207–3217 (2002).
Alvarez, J. P., Furumizu, C., Efroni, I., Eshed, Y. & Bowman, J. L. Active suppression of a leaf meristem orchestrates determinate leaf growth. eLife 5, e15023 (2016).
Martin-Trillo, M. & Cubas, P. TCP genes: a family snapshot ten years later. Trends Plant Sci. 15, 31–39 (2010).
Schommer, C. et al. Control of jasmonate biosynthesis and senescence by miR319 targets. PLoS Biol. 6, e230 (2008).
Mandelbrot, B. B., Freeman, W. H. & Company. The Fractal Geometry of Nature (Henry Holt and Company, 1983).
Hamant, O. et al. The KNAT2 homeodomain protein interacts with ethylene and cytokinin signaling. Plant Physiol. 130, 657–665 (2002).
Hibara, K. et al. Arabidopsis CUP-SHAPED COTYLEDON3 regulates postembryonic shoot meristem and organ boundary formation. Plant Cell 18, 2946–2957 (2006).
Blein, T., Pautot, V. & Laufs, P. Combinations of mutations sufficient to alter Arabidopsis leaf dissection. Plants 2, 230–247 (2013).
Zhou, C. et al. STM/BP-like KNOXI is uncoupled from ARP in the regulation of compound leaf development in Medicago truncatula. Plant Cell 26, 1464–1479 (2014).
Lenhard, M., Jurgens, G. & Laux, T. The WUSCHEL and SHOOTMERISTEMLESS genes fulfil complementary roles in Arabidopsis shoot meristem regulation. Development 129, 3195–3206 (2002).
Koyama, T., Sato, F. & Ohme-Takagi, M. Roles of miR319 and TCP transcription factors in leaf development. Plant Physiol. 175, 874–885 (2017).
Serra, L. & Perrot-Rechenmann, C. Spatiotemporal control of cell growth by CUC3 shapes leaf margins. Development https://doi.org/10.1242/dev.183277 (2020).
Efroni, I. et al. Regulation of leaf maturation by chromatin-mediated modulation of cytokinin responses. Dev. Cell 24, 438–445 (2013).
Zhao, M. et al. Arabidopsis BREVIPEDICELLUS interacts with the SWI2/SNF2 chromatin remodeling ATPase BRAHMA to regulate KNAT2 and KNAT6 expression in control of inflorescence architecture. PLoS Genet. 11, e1005125 (2015).
Hay, A. & Tsiantis, M. KNOX genes: versatile regulators of plant development and diversity. Development 137, 3153–3165 (2010).
Yu, H. et al. TCP5 controls leaf margin development by regulating KNOX and BEL-like transcription factors in Arabidopsis. J. Exp. Bot. 72, 1809–1821 (2021).
Beerling, D. J. & Fleming, A. J. Zimmermann’s telome theory of megaphyll leaf evolution: a molecular and cellular critique. Curr. Opin. Plant Biol. 10, 4–12 (2007).
He, L. et al. A molecular framework underlying the compound leaf pattern of Medicago truncatula. Nat. Plants 6, 511–521 (2020).
Trigg, S. A. et al. CrY2H-seq: a massively multiplexed assay for deep-coverage interactome mapping. Nat. Methods 14, 819–825 (2017).
Sabatini, S. et al. An auxin-dependent distal organizer of pattern and polarity in the Arabidopsis root. Cell 99, 463–472 (1999).
Ragni, L., Belles-Boix, E., Gunl, M. & Pautot, V. Interaction of KNAT6 and KNAT2 with BREVIPEDICELLUS and PENNYWISE in Arabidopsis inflorescences. Plant Cell 20, 888–900 (2008).
Larue, C. T., Wen, J. & Walker, J. C. A microRNA-transcription factor module regulates lateral organ size and patterning in Arabidopsis. Plant J. 58, 450–463 (2009).
Ichihashi, Y. et al. Key proliferative activity in the junction between the leaf blade and leaf petiole of Arabidopsis. Plant Physiol. 157, 1151–1162 (2011).
Karidas, P., Challa, K. R. & Nath, U. The tarani mutation alters surface curvature in Arabidopsis leaves by perturbing the patterns of surface expansion and cell division. J. Exp. Bot. 66, 2107–2122 (2015).
Aggarwal, P., Challa, K. R., Rath, M., Sunkara, P. & Nath, U. Generation of inducible transgenic lines of Arabidopsis transcription factors regulated by MicroRNAs. Methods Mol. Biol. 1830, 61–79 (2018).
Sessions, A., Weigel, D. & Yanofsky, M. F. The Arabidopsis thaliana MERISTEM LAYER 1 promoter specifies epidermal expression in meristems and young primordia. Plant J. 20, 259–263 (1999).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Acknowledgements
We thank J. Bowman, T. Imaizumi, H. Tsukaya, V. Pautot, T. Jack, D. Weigel, Y. Eshed, P. Laufs and P. Aggarwal for plant material; C. Tian for raw-read counts of tissue-specific RNA-seq datasets; and Genotypic Technology for the cycloheximide and cycloheximide + dexamethasone microarray experiment. This work was supported by the Ministry of Human Resource Development, Government of India (fellowships to K.R.C., M.R. and A.N.S.), Department of Science & Technology for Improvement of S&T Infrastructure (DST-FIST), University Grants Commission Centre for Advanced Studies, and Department of Biotechnology (DBT)-IISc Partnership Program Phase-II at IISc (sanction no. BT/PR27952/INF/22/212/2018 to U.N.). A.K.B. and S.D. were supported by Shodhaka Life Sciences.
Author information
Authors and Affiliations
Contributions
K.R.C. initiated the project, performed several initial experiments, analysed and interpreted results, organized the figures, wrote the first draft of the manuscript and contributed to its finalization; M.R. designed and performed the majority of the experiments with input from K.R.C., analysed and interpreted data, contributed to making figures and helped finalize the manuscript; A.K.B., S.D. and K.K.A. carried out the alignment of the raw RNA-seq reads to the reference genome and initial transcriptomic analysis; A.N.S. carried out transcriptome analysis of RNA-seq and microarray datasets that yielded Figs. 3a,b and 5d, and helped finalize the manuscript; and U.N. contributed to designing experiments and data interpretation, guided the first three authors, corrected the manuscript and finalized it.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Naomi Ori and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
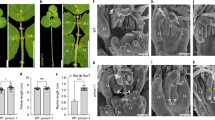
Extended Data Fig. 1 CIN-TCP and KNOX-II redundantly promote cotyledon maturation.
a,b, Cotyledons of 9-day old seedlings grown in the presence of 0 (Mock) or 12 µM (Dex) dexamethasone (a) and their abaxial epidermal cell images (b). Scale bar in (a) 2 mm and in (b) 50 µm. c, Abundance of CIN-TCP and KNOX transcripts in Arabidopsis tissue types (shown in cartoon) analyzed using the expression datasets available in the Genevestigator database (https://genevestigator.com). Sample numbers of microarray experiments are indicated. (d) pTCP4::GUS and pKNAT4::GUS expression in 5 and 9-day old seedlings.
Extended Data Fig. 2 Higher order leaflet emergence in jk-mock leaves.
a, 6th leaf from a 60 DAS jaw-D;pTCP4::mTCP4:GR x knat3;4;5-amiR (jk-mock) plant (shown in Fig. 1c) branches dissected from the main leaf. The order of visible leaflets on the branch is shown as numbers from 1–8. b, 32 DAS rosettes (left) of indicated genotypes grown in the presence of 0 (Mock) or 12 µM (Dex) dexamethasone and their leaves (right). Node numbers of the leaves are indicated at the bottom. Scale bar, 5 mm.
Extended Data Fig. 3 Suppression of ectopic leaflet emergence in jk-Mock leaves upon transient TCP4 induction.
a, A scheme of transient dexamethasone treatment experiment for 24 h in 14 or 16 DAS jk-Mock plants which were further growth till 40 DAS. b, Leaves (numbers indicate node positions) from 40 DAS jk-Mock plants grown without (Mock) or with 12 µM dexamethasone treatment for 24 h at 14 DAS or 16 DAS. Leaflet count of these leaves is shown in Fig. 1g. c, 2nd cauline leaf of jk-Mock plants at 75 DAS grown in the presence of 0 (Mock) or 12 (Dex) µM dexamethasone. d,e, Scanning electron micrographs corresponding to the boxed region in (c). f, Scanning electron micrograph of a jk-Mock 6th leaf (*) at 15 DAS. Scale bar, 5 mm (c), 150 µm (d) and 50 µm (e).
Extended Data Fig. 4 Transient TCP4 induction suppresses ectopic pCyclinB1;1 expression in jk-Mock leaves.
a, pCyclinB1;1::GUS activity in leaves (numbers denote node positions) from 25 DAS plants of indicated genotypes. Scale bar, 1 mm. b, pCyclinB1;1::GUS activity in 5th, 6th and 8th leaves from 32 DAS jk-Mock plants grown without dexamethasone (Mock), with transient dexamethasone treatment for 24 h at 16 DAS (16 DAS-24h Dex), or with continuous dexamethasone (Dex). Scale bar, 1 mm.
Extended Data Fig. 5 pCUC2::GUS expression analysis in CIN-TCP and KNOX-II knockdown leaves.
a, pCUC2::GUS activity in 6th leaves of indicated genotypes. b, pCUC2::GUS activity in the 5th, 6th and 8th leaves of jk-Mock plants grown without dexamethasone (Mock), with transient dexamethasone treatment for 24 h at 16 DAS (16 DAS-24h Dex), or with continuous dexamethasone (Dex). c, pDR5::GUS activity in the 5th leaf from 28 DAS plants. In jk-Mock;pDR5::GUS, the two panels on the right indicate leaflets from the leaf shown on the left. In jk-Dex;pDR5::GUS, a magnified image of the boxed region is shown in the inset. Arrowheads point to GUS expression at leaflet tips. Scale bar, 1 mm.
Extended Data Fig. 6 Leaflet phenotype and KNOX-I expression analysis in jk-Mock leaves.
a, Rosettes of indicated genotypes at 55 DAS. Scale bar, 1 cm. b, 6th leaf from 56 DAS jk-Mock plants grown without (jk-Mock) or with (jk-Dex) 12 µM dexamethasone or 80 µM L-kynurenine (L-Kyn). Scale bar, 5 mm. c, GUS reporter analysis at different developmental stages of leaves from 30 DAS jk-Mock plant. Leaf lengths are indicated. Punctate pKNAT2::GUS and pKNAT6::GUS signal is highlighted with asterisks and arrowheads, respectively. pCUC2::GUS signal is shown for comparison.
Extended Data Fig. 7 Analysis of meristem gene expression in jk-Mock leaves.
a-c, GUS reporter analysis of the 6th leaf from 25 DAS plants of indicated genotypes. Magnified images of the boxed regions are shown as insets on the right. Scale bar, 1 mm (a,b and the left panel in c) and 100 µm (insets in c).
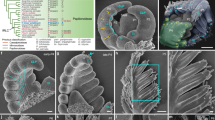
Extended Data Fig. 8 CIN-TCPs activate KNOX-II genes.
a, Heat map of differentially expressed transcripts in the microarray experiment of 9-day old jaw-D;pTCP4::mTCP4:GR seedlings treated with either 40 µM cycloheximide (CHX) or a combination of 40 µM cycloheximide and 20 µM dexamethasone (CHX & Dex) for 4 hours (see Fig. 5c). b, Relative level of KNAT3 and KNAT4 transcripts analyzed by RT-qPCR in 9 DAS p35S::mTCP4:GR seedlings treated with 40 µM cycloheximide (CHX) or a combination of 20 µM dexamethasone and 40 µM CHX (CHX & Dex) for 4 h. Transcript levels were normalized to internal control PP2A. N = 3, Averages of three biological replicates are shown. Error bars indicate mean value SD. P, probability values of unpaired Student’s t-test. c, EMSA gel blots with radio-labeled (Probe) or unlabeled (Competitor) oligonucleotides corresponding to BS1 and BS3 (shown in Fig. 6a) and recombinant MBP-TCP4D protein. d, EMSA with radio-labeled (Probe) or unlabeled (Competitor) oligonucleotides corresponding to BS2 and BS3 (shown in Fig. 6b) and recombinant His-TCP4 protein. 25–150 fold excess competitor was used. e, Average size of abaxial pavement cells of the 1st leaves in 29 DAS seedlings. Increased cell size in the jaw-D;GR x p35S::KNAT3 leaves compared to jaw-D;GR x Col-0 leaves suggests that KNAT3 overexpression partly rescues the small-cell phenotype of jaw-D. Sample number, N = 120–150 cells per leaf were measured and averages of 3–5 leaves are shown. Unpaired Student’s t-test was used. Error bars indicate mean value SD. P, probability values of unpaired Student’s t-test. f, Relative transcript level analyzed by RT-qPCR in 9 DAS seedlings. Reduced levels of TCP2, 3, 10 & 24 in jaw-D;GR x p35S::KNAT3 suggests that the rescue of jaw-D leaf phenotype by KNAT3 overexpression is not due to the co-suppression of the jaw-D locus. Transcript levels were normalized to internal control PP2A. N = 3, Averages of three biological replicates are shown. Error bars indicate mean value SD. P, probability values of unpaired Student’s t-test.
Extended Data Fig. 9 Reproductive organ development in jk-Mock plants.
a,b, Flowers (a) and inflorescence (b) of 42 DAS mock-grown plants. White arrows indicate infertile flowers. Scale bar, 1 mm (a) or 5 mm (b).
Supplementary information
Source data
Source Data Fig. 1
a, Complete gene list of differentially regulated genes in jaw-D;GR × knat3;4;5-amiR (jk-mock) only and not in jaw-D;GR and knat3;4;5-amiR compared to Col-0. b, Full list of division- and age-specific marker genes. c, List of division- and age-specific marker genes differentially regulated in jaw-D;GR × knat3;4;5-amiR (jk-mock) only and not in jaw-D;GR and knat3;4;5-amiR.
Source Data Fig. 2
Common DEGs obtained by comparing the microarray done using 9 DAS jaw-D;pTCP4::mTCP4:GR seedlings treated with 40 µM cycloheximide (CHX) or a combination of 40 µM CHX and 20 µM dexamethasone (CHX DEX) for 4 h with the previously published microarray28.
Source Data Fig. 3
List of primers and oligonucleotides used in this study.
Rights and permissions
About this article
Cite this article
Challa, K.R., Rath, M., Sharma, A.N. et al. Active suppression of leaflet emergence as a mechanism of simple leaf development. Nat. Plants 7, 1264–1275 (2021). https://doi.org/10.1038/s41477-021-00965-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-00965-3
This article is cited by
-
Rewiring of a KNOXI regulatory network mediated by UFO underlies the compound leaf development in Medicago truncatula
Nature Communications (2024)
-
Control of compound leaf patterning by MULTI-PINNATE LEAF1 (MPL1) in chickpea
Nature Communications (2023)
-
Genome-wide characterization of the TALE homeodomain family and the KNOX-BLH interaction network in tomato
Plant Molecular Biology (2022)