Abstract
Photosynthesis is readily impaired by high light (HL) levels. Photosynthetic organisms have therefore evolved various mechanisms to cope with the problem. Here, we have dramatically enhanced the light tolerance of the cyanobacterium Synechocystis by adaptive laboratory evolution (ALE). By combining repeated mutagenesis and exposure to increasing light intensities, we generated strains that grow under extremely HL intensities. HL tolerance was associated with more than 100 mutations in proteins involved in various cellular functions, including gene expression, photosynthesis and metabolism. Co-evolved mutations were grouped into five haplotypes, and putative epistatic interactions were identified. Two representative mutations, introduced into non-adapted cells, each confer enhanced HL tolerance, but they affect photosynthesis and respiration in different ways. Mutations identified by ALE that allow photosynthetic microorganisms to cope with altered light conditions could be employed in assisted evolution approaches and could strengthen the robustness of photosynthesis in crop plants.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All data supporting the findings of this study are available within the paper and its Supplementary Information files. The biological material is available upon reasonable request. Source data are provided with this paper.
References
Apel, K. & Hirt, H. Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu. Rev. Plant Biol. 55, 373–399 (2004).
Asada, K. The water–water cycle in chloroplasts: scavenging of active oxygens and dissipation of excess photons. Annu. Rev. Plant Physiol. Plant Mol. Biol. 50, 601–639 (1999).
Chaux, F., Peltier, G. & Johnson, X. A security network in PSI photoprotection: regulation of photosynthetic control, NPQ and O2 photoreduction by cyclic electron flow. Front. Plant Sci. 6, 875 (2015).
Nishiyama, Y., Allakhverdiev, S. I. & Murata, N. A new paradigm for the action of reactive oxygen species in the photoinhibition of photosystem II. Biochim. Biophys. Acta 1757, 742–749 (2006).
Sonoike, K. Photoinhibition of photosystem I. Physiol. Plant. 142, 56–64 (2011).
Jarvi, S., Suorsa, M. & Aro, E. M. Photosystem II repair in plant chloroplasts—regulation, assisting proteins and shared components with photosystem II biogenesis. Biochim. Biophys. Acta 1847, 900–909 (2015).
de Bianchi, S., Ballottari, M., Dall’osto, L. & Bassi, R. Regulation of plant light harvesting by thermal dissipation of excess energy. Biochem. Soc. Trans. 38, 651–660 (2010).
Kirilovsky, D. & Kerfeld, C. A. Cyanobacterial photoprotection by the orange carotenoid protein. Nat. Plants 2, 16180 (2016).
Komenda, J. & Sobotka, R. Cyanobacterial high-light-inducible proteins—protectors of chlorophyll-protein synthesis and assembly. Biochim. Biophys. Acta 1857, 288–295 (2016).
Muramatsu, M. & Hihara, Y. Acclimation to high-light conditions in cyanobacteria: from gene expression to physiological responses. J. Plant Res. 125, 11–39 (2012).
Niyogi, K. K. & Truong, T. B. Evolution of flexible non-photochemical quenching mechanisms that regulate light harvesting in oxygenic photosynthesis. Curr. Opin. Plant Biol. 16, 307–314 (2013).
Ruban, A. V. Nonphotochemical chlorophyll fluorescence quenching: mechanism and effectiveness in protecting plants from photodamage. Plant Physiol. 170, 1903–1916 (2016).
Sonoike, K., Hihara, Y. & Ikeuchi, M. Physiological significance of the regulation of photosystem stoichiometry upon high light acclimation of Synechocystis sp. PCC 6803. Plant Cell Physiol. 42, 379–384 (2001).
Garcia-Molina, A. & Leister, D. Accelerated relaxation of photoprotection impairs biomass accumulation in Arabidopsis. Nat. Plants 6, 9–12 (2020).
Kromdijk, J. et al. Improving photosynthesis and crop productivity by accelerating recovery from photoprotection. Science 354, 857–861 (2016).
Exposito-Rodriguez, M., Laissue, P. P., Yvon-Durocher, G., Smirnoff, N. & Mullineaux, P. M. Photosynthesis-dependent H2O2 transfer from chloroplasts to nuclei provides a high-light signalling mechanism. Nat. Commun. 8, 49 (2017).
Schierenbeck, L. et al. Fast forward genetics to identify mutations causing a high light tolerant phenotype in Chlamydomonas reinhardtii by whole-genome-sequencing. BMC Genom. 16, 57 (2015).
Jensen, P. E. & Leister, D. Cyanobacteria as an experimental platform for modifying bacterial and plant photosynthesis. Front. Bioeng. Biotechnol. 2, 7 (2014).
Leister, D. Experimental evolution in photoautotrophic microorganisms as a means of enhancing chloroplast functions. Essays Biochem. 62, 77–84 (2018).
Heidorn, T. et al. Synthetic biology in cyanobacteria: engineering and analyzing novel functions. Methods Enzymol. 497, 539–579 (2011).
Lopo, M. et al. Experimental and modeling analysis of Synechocystis sp. PCC 6803 growth. J. Mol. Microbiol. Biotechnol. 22, 71–82 (2012).
Tillich, U. M., Lehmann, S., Schulze, K., Duhring, U. & Frohme, M. The optimal mutagen dosage to induce point-mutations in Synechocystis sp. PCC6803 and its application to promote temperature tolerance. PLoS ONE 7, e49467 (2012).
Tillich, U. M., Wolter, N., Franke, P., Duhring, U. & Frohme, M. Screening and genetic characterization of thermo-tolerant Synechocystis sp. PCC6803 strains created by adaptive evolution. BMC Biotechnol. 14, 66 (2014).
Kaneko, T. et al. Sequence analysis of the genome of the unicellular cyanobacterium Synechocystis sp. strain PCC6803. II. Sequence determination of the entire genome and assignment of potential protein-coding regions. DNA Res. 3, 109–136 (1996).
Hagemann, M. et al. Identification of the DNA methyltransferases establishing the methylome of the cyanobacterium Synechocystis sp. PCC 6803. DNA Res. 25, 343–352 (2018).
Hu, L. et al. Transgenerational epigenetic inheritance under environmental stress by genome-wide DNA methylation profiling in cyanobacterium. Front. Microbiol. 9, 1479 (2018).
Dann, M. & Leister, D. Evidence that cyanobacterial Sll1217 functions analogously to PGRL1 in enhancing PGR5-dependent cyclic electron flow. Nat. Commun. 10, 5299 (2019).
Munekage, Y. et al. PGR5 is involved in cyclic electron flow around photosystem I and is essential for photoprotection in Arabidopsis. Cell 110, 361–371 (2002).
Battchikova, N., Eisenhut, M. & Aro, E. M. Cyanobacterial NDH-1 complexes: novel insights and remaining puzzles. Biochim. Biophys. Acta 1807, 935–944 (2011).
Ogawa, T. & Mi, H. Cyanobacterial NADPH dehydrogenase complexes. Photosynth. Res. 93, 69–77 (2007).
Peltier, G., Aro, E. M. & Shikanai, T. NDH-1 and NDH-2 plastoquinone reductases in oxygenic photosynthesis. Annu. Rev. Plant Biol. 67, 55–80 (2016).
Gao, F. et al. The NDH-1L-PSI supercomplex is important for efficient cyclic electron transport in cyanobacteria. Plant Physiol. 172, 1451–1464 (2016).
Yeremenko, N. et al. Open reading frame ssr2016 is required for antimycin A-sensitive photosystem I-driven cyclic electron flow in the cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol. 46, 1433–1436 (2005).
Kojima, K. et al. Oxidation of elongation factor G inhibits the synthesis of the D1 protein of photosystem II. Mol. Microbiol. 65, 936–947 (2007).
Ejima, K., Kawaharada, T., Inoue, S., Kojima, K. & Nishiyama, Y. A change in the sensitivity of elongation factor G to oxidation protects photosystem II from photoinhibition in Synechocystis sp. PCC 6803. FEBS Lett. 586, 778–783 (2012).
Tajima, N. et al. Genomic structure of the cyanobacterium Synechocystis sp. PCC 6803 strain GT-S. DNA Res. 18, 393–399 (2011).
Trautmann, D., Voss, B., Wilde, A., Al-Babili, S. & Hess, W. R. Microevolution in cyanobacteria: re-sequencing a motile substrain of Synechocystis sp. PCC 6803. DNA Res. 19, 435–448 (2012).
Zavrel, T., Ocenasova, P. & Cerveny, J. Phenotypic characterization of Synechocystis sp. PCC 6803 substrains reveals differences in sensitivity to abiotic stress. PLoS ONE 12, e0189130 (2017).
Stanier, R. Y., Kunisawa, R., Mandel, M. & Cohen-Bazire, G. Purification and properties of unicellular blue-green algae (order Chroococcales). Bacteriol. Rev. 35, 171–205 (1971).
Yoshihara, S. & Ikeuchi, M. Phototactic motility in the unicellular cyanobacterium Synechocystis sp. PCC 6803. Photochem. Photobiol. Sci. 3, 512–518 (2004).
Ohad, I., Raanan, H., Keren, N., Tchernov, D. & Kaplan, A. Light-induced changes within photosystem II protects Microcoleus sp. in biological desert sand crusts against excess light. PLoS ONE 5, e11000 (2010).
Raanan, H. et al. Towards clarifying what distinguishes cyanobacteria able to resurrect after desiccation from those that cannot: the photosynthetic aspect. Biochim. Biophys. Acta 1857, 715–722 (2016).
Ungerer, J., Wendt, K. E., Hendry, J. I., Maranas, C. D. & Pakrasi, H. B. Comparative genomics reveals the molecular determinants of rapid growth of the cyanobacterium Synechococcus elongatus UTEX 2973. Proc. Natl Acad. Sci. USA 115, E11761–E11770 (2018).
Gaitan-Espitia, J. D. & Hobday, A. J. Evolutionary principles and genetic considerations for guiding conservation interventions under climate change. Glob. Change Biol. 27, 475–488 (2021).
Harvey, B. J., Nash, K. L., Blanchard, J. L. & Edwards, D. P. Ecosystem-based management of coral reefs under climate change. Ecol. Evol. 8, 6354–6368 (2018).
van Oppen, M. J., Oliver, J. K., Putnam, H. M. & Gates, R. D. Building coral reef resilience through assisted evolution. Proc. Natl Acad. Sci. USA 112, 2307–2313 (2015).
Leister, D. Genetic engineering, synthetic biology and the light reactions of photosynthesis. Plant Physiol. 179, 778–793 (2019).
Rippka, R., Deruelles, J., Waterbury, J. B., Herdman, M. & Stanier, R. Y. Generic assignments, strain histories and properties of pure cultures of cyanobacteria. Microbiology 111, 1–61 (1979).
Tillett, D. & Neilan, B. A. Xanthogenate nucleic acid isolation from cultured and environmental cyanobacteria. J. Phycol. 36, 251–258 (2000).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Wick, R. R., Schultz, M. B., Zobel, J. & Holt, K. E. Bandage: interactive visualization of de novo genome assemblies. Bioinformatics 31, 3350–3352 (2015).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Bushnell, B. BBTools: a suite of bioinformatic tools used for DNA and RNA sequence data analysis. (Joint Genome Institute, 2016); http://jgi.doe.gov/data-and-tools/bbtools
Bushnell, B., Rood, J. & Singer, E. BBMerge—accurate paired shotgun read merging via overlap. PLoS ONE 12, e0185056 (2017).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963 (2014).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Barrick, J. E. et al. Identifying structural variation in haploid microbial genomes from short-read resequencing data using breseq. BMC Genom. 15, 1039–1017 (2014).
Deatherage, D. E. & Barrick, J. E. Identification of mutations in laboratory-evolved microbes from next-generation sequencing data using breseq. Methods Mol. Biol. 1151, 165–188 (2014).
Garrison, E. & Marth G. Haplotype-based variant detection from short-read sequencing. Preprint at https://arxiv.org/abs/1207.3907v2 (2012).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Viola, S., Rühle, T. & Leister, D. A single vector-based strategy for marker-less gene replacement in Synechocystis sp. PCC 6803. Microb. Cell Fact. 13, 4 (2014).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Schägger, H. & von Jagow, G. Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal. Biochem. 166, 368–379 (1987).
Heinz, S. et al. Thylakoid membrane architecture in Synechocystis depends on CurT, a homolog of the granal CURVATURE THYLAKOID1 proteins. Plant Cell 28, 2238–2260 (2016).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Zavřel, T., Sinetova, M. A. & Červený, J. Measurement of chlorophyll a and carotenoids concentration in cyanobacteria. Bio-Protoc. 5, e1467 (2015).
Watanabe, M. et al. Attachment of phycobilisomes in an antenna–photosystem I supercomplex of cyanobacteria. Proc. Natl Acad. Sci. USA 111, 2512–2517 (2014).
Murakami, A. Quantitative analysis of 77K fluorescence emission spectra in Synechocystis sp. PCC 6714 and Chlamydomonas reinhardtii with variable PS I/PS II stoichiometries. Photosynth. Res. 53, 141–148 (1997).
Rakhimberdieva, M. G., Boichenko, V. A., Karapetyan, N. V. & Stadnichuk, I. N. Interaction of phycobilisomes with photosystem II dimers and photosystem I monomers and trimers in the cyanobacterium Spirulina platensis. Biochemistry 40, 15780–15788 (2001).
Acknowledgements
We thank P. Hardy for his critical reading of the manuscript, and we thank the German Science Foundation (DFG, grant nos. TR175 Z1, GRK2062 and EXC2089/1 (e-conversion, funding ID: 390776260) to D.L.) and the European Research Council (ERC Synergy Grant ‘PhotoRedesign’) for financial support.
Author information
Authors and Affiliations
Contributions
D.L., M.D. and A.G. designed the adaptive evolution experiments and, together with M.T., performed them. E.M.O. and H.S. designed the sequencing strategy and analysed the next-generation sequencing data. M.L. performed the carotenoid profiling. D.L. was responsible for the conceptualization and management of the entire study and wrote the paper with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Diana Kirilovsky and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
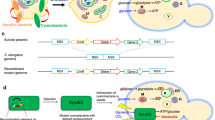
Extended Data Fig. 1 Synechocystis genomic allele-swapping strategy.
Candidate point mutations were introduced into the Synechocystis LT genome employing a modified version of our established single-vector-based, marker-less gene replacement strategy62. The complete ORF of interest (5’ CDS+ 3’ CDS) and a second copy of the 3’ CDS were cloned into a non-replicative plasmid on either side of the double-selection cassette (DSC)62. SNPs were introduced by Q5 site-directed mutagenesis. The endogenous genomic ORF was replaced with the mutated ORF by homologous recombination upon positive selection on kanamycin (KanR; nptI), and the original gene structure was re-established by removal of the DSC by intra-chromosomal homologous recombination upon negative selection on sucrose (SucrS; sacB).
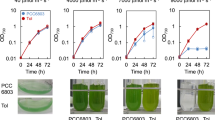
Extended Data Fig. 2 Growth curves for mutant and control strains at 50, 700 and 1200 μmol photons m-2 s-1 and HL tolerance at 2000 μmol photons m-2 s-1.
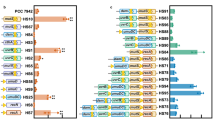
a, Growth of Synechocystis strains over the course of 7 days post inoculation was measured as apparent optical density at λ=720 nm (OD720nm). LT, our laboratory-adapted Synechocystis PCC6803 strain; WT, the original motile Synechocystis PCC6803 strain obtained from the Pasteur Collection; #1, NdhF1F124L; #2, EF-G2R461C; #12, NdhF1F124L EF-G2R461C. Note that cultures were inoculated to reach an OD730nm = 0.05, and that cultivation of strains in Multi-Cultivator OD-1000 devices and direct measurement with the in-built system leads to underestimation of the actual OD720nm if actual values exceed ~0.5. b, Duplication times during the exponential phase of growth curves shown in (a). Duplication time data was inferred from growth curve slopes over eight hours starting at OD = 0.1 (that is after the first full duplication of culture OD). c, HL tolerance at 2000 μmol photons m-2 s-1 of NdhF1F124L, EF-G2R461C, NdhF1F124L EF-G2R461C cells, LT and WT strains, analysed as in Fig. 5. Data are derived from six replicates (two biological replicates of three independent clones per genotype). Error bars indicated SDs from respective average values. Crosses in boxplots indicate average values, letters signify statistically significant differences with p ≤ 0.05 according to post-hoc Bonferroni-Holm simultaneous comparison of all measurements after significant among-group differences were detected by one-factorial ANOVA (two-sided).
Extended Data Fig. 3 Cell culture absorbance spectra.
Absorbance spectra shown are averages of six biological replicates of Synechocystis cultures 7 days post inoculation. Data have been baseline-corrected, such that absorbance at 750 nm equals 0, and were used to estimate phycocyanin-to-chlorophyll a (PC/Chl) molar ratios70.
Extended Data Fig. 4 Low temperature (77 K) fluorescence spectra.
a, 77 K fluorescence emission spectra upon excitation at 435 nm. b, 77 K fluorescence emission spectra upon excitation at 600 nm. With the exception of the data for LT at 700 μmol photons m-2 s-1 (for which n = 4), all spectra shown are averages of six biological replicates of Synechocystis cultures measured 7 days post inoculation. Data has been baseline corrected for emission at 600 nm (a) and 620 nm (b), and normalized to input culture OD730nm, respectively. Spectra were corrected for dark-signal (dark-offset ON) and the blank value (BG11 medium prior to inoculation) was subtracted.
Extended Data Fig. 5 HL tolerance and motility of the original Synechocystis sp. PCC6803 strain.
Time course of the growth of the original (motile) Synechocystis sp PCC6803 strain at 2000 µmol photons m-2 s-1 and 23 °C over 7 days, after inoculation at OD730nm = 0.05. The white arrowheads indicate the initial level of the liquid culture; the dashed outline indicates an area of reduced light exposure owing to the absence of LEDs in the cultivator design. Note that the motile cells migrate out of the liquid medium and only begin to return at 6 days post inoculation (dpi). This observation indicates that this strain requires an extended period of acclimation before it can cope with such high light intensities.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2.
Supplementary Data 1
Processed sequencing data output with identified potentially HL-adaptive mutations categorized by types, frequencies and occurrences in epistatic series and haplotypes.
Supplementary Data 2
Sequencing data SNP analysis output—all candidate SNPs identified in the HL ALE experiment.
Source data
Source Data Fig. 2
Dry mass raw data and statistical analyses for ALE final batch cultures.
Source Data Fig. 3
FluorCam raw data for ALE isolated clones.
Source Data Fig. 5
OD, dry mass, pigment raw data and statistical analyses for ALE reconstituted point mutants.
Source Data Fig. 6
77 K fluorescence and immunoblot raw data and statistical analyses for ALE reconstituted point mutants.
Source Data Fig. 7
Chlorophyll fluorescence, oxygen evolution, P700 redox kinetics raw data and statistical analyses for ALE reconstituted point mutants.
Source Data Extended Data Fig. 2
Growth curves and WT PCC6803 final OD and dry mass at 2,000 µmol photons per m2 per s for ALE reconstituted point mutants.
Source Data Extended Data Fig. 3
77 K fluorescence spectra at excitation wavelengths 435 and 600 nm for ALE reconstituted point mutants.
Rights and permissions
About this article
Cite this article
Dann, M., Ortiz, E.M., Thomas, M. et al. Enhancing photosynthesis at high light levels by adaptive laboratory evolution. Nat. Plants 7, 681–695 (2021). https://doi.org/10.1038/s41477-021-00904-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-00904-2
This article is cited by
-
Light-driven biosynthesis of volatile, unstable and photosensitive chemicals from CO2
Nature Synthesis (2023)
-
Engineered hypermutation adapts cyanobacterial photosynthesis to combined high light and high temperature stress
Nature Communications (2023)
-
Asymmetric survival in single-cell lineages of cyanobacteria in response to photodamage
Photosynthesis Research (2023)
-
Absolute quantification of cellular levels of photosynthesis-related proteins in Synechocystis sp. PCC 6803
Photosynthesis Research (2023)