Abstract
Meiotic crossovers are tightly restricted in most eukaryotes, despite an excess of initiating DNA double-strand breaks. The majority of plant crossovers are dependent on class I interfering repair, with a minority formed via the class II pathway. Class II repair is limited by anti-recombination pathways; however, similar pathways repressing class I crossovers have not been identified. Here, we performed a forward genetic screen in Arabidopsis using fluorescent crossover reporters to identify mutants with increased or decreased recombination frequency. We identified HIGH CROSSOVER RATE1 (HCR1) as repressing crossovers and encoding PROTEIN PHOSPHATASE X1. Genome-wide analysis showed that hcr1 crossovers are increased in the distal chromosome arms. MLH1 foci significantly increase in hcr1 and crossover interference decreases, demonstrating an effect on class I repair. Consistently, yeast two-hybrid and in planta assays show interaction between HCR1 and class I proteins, including HEI10, PTD, MSH5 and MLH1. We propose that HCR1 plays a major role in opposition to pro-recombination kinases to restrict crossovers in Arabidopsis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Genome sequencing data of F2 plants can be found at the ArrayExpress repository hosted by the European Bioinformatics Institute under accessions E-MTAB-9621 and E-MTAB-10168. Source data are provided with this paper.
References
Villeneuve, A. M. & Hillers, K. J. Whence meiosis? Cell 106, 647–650 (2001).
Mercier, R., Mézard, C., Jenczewski, E., Macaisne, N. & Grelon, M. The molecular biology of meiosis in plants. Annu. Rev. Plant Biol. 66, 297–327 (2015).
Grelon, M., Vezon, D., Gendrot, G. & Pelletier, G. AtSPO11-1 is necessary for efficient meiotic recombination in plants. EMBO J. 20, 589–600 (2001).
Robert, T. et al. The TopoVIB-Like protein family is required for meiotic DNA double-strand break formation. Science 351, 943–949 (2016).
Hartung, F. et al. The catalytically active tyrosine residues of both SPO11-1 and SPO11-2 are required for meiotic double-strand break induction in Arabidopsis. Plant Cell 19, 3090–3099 (2007).
Hunter, N. Meiotic recombination: the essence of heredity. Cold Spring Harb. Perspect. Biol. 7, a016618 (2015).
Ferdous, M. et al. Inter-homolog crossing-over and synapsis in Arabidopsis meiosis are dependent on the chromosome axis protein AtASY3. PLoS Genet. 8, e1002507 (2012).
Rowan, B. A. et al. An ultra high-density Arabidopsis thaliana crossover map that refines the influences of structural variation and epigenetic features. Genetics 213, 771–787 (2019).
Girard, C. et al. AAA-ATPase FIDGETIN-LIKE 1 and helicase FANCM antagonize meiotic crossovers by distinct mechanisms. PLoS Genet. 11, e1005369 (2015).
Cifuentes, M., Rivard, M., Pereira, L., Chelysheva, L. & Mercier, R. Haploid meiosis in Arabidopsis: double-strand breaks are formed and repaired but without synapsis and crossovers. PLoS ONE 8, e72431 (2013).
Berchowitz, L. E. & Copenhaver, G. P. Genetic interference: don’t stand so close to me. Curr. Genomics 11, 91–102 (2010).
Li, Y. et al. HEIP1 regulates crossover formation during meiosis in rice. Proc. Natl Acad. Sci. USA 115, 10810–10815 (2018).
Pyatnitskaya, A., Borde, V. & De Muyt, A. Crossing and zipping: molecular duties of the ZMM proteins in meiosis. Chromosoma 128, 181–198 (2019).
Mercier, R. et al. Two meiotic crossover classes cohabit in Arabidopsis: one is dependent on MER3,whereas the other one is not. Curr. Biol. 15, 692–701 (2005).
Séguéla-Arnaud, M. et al. Multiple mechanisms limit meiotic crossovers: TOP3α and two BLM homologs antagonize crossovers in parallel to FANCM. Proc. Natl Acad. Sci. USA 112, 4713–4718 (2015).
Serra, H. et al. Massive crossover elevation via combination of HEI10 and recq4a recq4b during Arabidopsis meiosis. Proc. Natl Acad. Sci. USA 115, 2437–2442 (2018).
Marston, A. L. & Amon, A. Meiosis: cell-cycle controls shuffle and deal. Nat. Rev. Mol. Cell Biol. 5, 983–997 (2004).
Yang, C. et al. The Arabidopsis Cdk1/Cdk2 homolog CDKA;1 controls chromosome axis assembly during plant meiosis. EMBO J. 39, 1–19 (2020).
Lange, J. et al. The landscape of mouse meiotic double-strand break formation, processing, and repair. Cell 167, 695–708 (2016).
Garcia, V., Gray, S., Allison, R. M., Cooper, T. J. & Neale, M. J. Tel1(ATM)-mediated interference suppresses clustered meiotic double-strand-break formation. Nature 520, 114–118 (2015).
Lange, J. et al. ATM controls meiotic double-strand-break formation. Nature 479, 237–240 (2011).
Serrentino, M.-E., Chaplais, E., Sommermeyer, V. & Borde, V. Differential association of the conserved SUMO ligase Zip3 with meiotic double-strand break sites reveals regional variations in the outcome of meiotic recombination. PLoS Genet. 9, e1003416 (2013).
He, W. et al. Regulated proteolysis of MutSγ controls meiotic crossing over. Mol. Cell 78, 168–183 (2020).
Carballo, J. A., Johnson, A. L., Sedgwick, S. G. & Cha, R. S. Phosphorylation of the axial element protein Hop1 by Mec1/Tel1 ensures meiotic interhomolog recombination. Cell 132, 758–770 (2008).
Brar, G. A. et al. Rec8 phosphorylation and recombination promote the step-wise loss of cohesins in meiosis. Nature 441, 532–536 (2006).
Hustedt, N. et al. Yeast PP4 interacts with ATR homolog Ddc2–Mec1 and regulates checkpoint signaling. Mol. Cell 57, 273–289 (2015).
Lee, D. H. et al. A PP4 phosphatase complex dephosphorylates RPA2 to facilitate DNA repair via homologous recombination. Nat. Struct. Mol. Biol. 17, 365–372 (2010).
Wang, S. et al. The PROTEIN PHOSPHATASE4 complex promotes transcription and processing of primary microRNAs in Arabidopsis. Plant Cell 31, 486–501 (2019).
Wu, G., Rossidivito, G., Hu, T., Berlyand, Y. & Poethig, R. S. Traffic lines: new tools for genetic analysis in Arabidopsis thaliana. Genetics 200, 35–45 (2015).
Melamed-Bessudo, C., Yehuda, E., Stuitje, A. R. & Levy, A. A. A new seed-based assay for meiotic recombination in Arabidopsis thaliana. Plant J. 43, 458–466 (2005).
Berchowitz, L. E. & Copenhaver, G. P. Fluorescent Arabidopsis tetrads: a visual assay for quickly developing large crossover and crossover interference data sets. Nat. Protoc. 3, 41–50 (2008).
Ziolkowski, P. A. et al. Juxtaposition of heterozygous and homozygous regions causes reciprocal crossover remodelling via interference during Arabidopsis meiosis. eLife 4, e03708 (2015).
Yelina, N. E. et al. DNA methylation epigenetically silences crossover hot spots and controls chromosomal domains of meiotic recombination in Arabidopsis. Genes Dev. 29, 2183–2202 (2015).
Lawrence, E. J. et al. Natural variation in TBP-ASSOCIATED factor 4b controls meiotic crossover and germline transcription in Arabidopsis. Curr. Biol. 29, 2676–2686 (2019).
Choi, K. et al. Arabidopsis meiotic crossover hot spots overlap with H2A.Z nucleosomes at gene promoters. Nat. Genet. 45, 1327–1336 (2013).
Ziolkowski, P. A. et al. Natural variation and dosage of the HEI10 meiotic E3 ligase control Arabidopsis crossover recombination. Genes Dev. 31, 306–317 (2017).
Crismani, W. et al. FANCM limits meiotic crossovers. Science 336, 1588–1590 (2012).
Allen, R., Nakasugi, K., Doran, R. L., Millar, A. A. & Waterhouse, P. M. Facile mutant identification via a single parental backcross method and application of whole genome sequencing based mapping pipelines. Front. Plant Sci. 4, 362 (2013).
Shi, Y. Serine/threonine phosphatases: mechanism through structure. Cell 139, 468–484 (2009).
Gingras, A. C. et al. A novel, evolutionarily conserved protein phosphatase complex involved in cisplatin sensitivity. Mol. Cell. Proteom. 4, 1725–1740 (2005).
Ramos, F., Villoria, M. T., Alonso-Rodríguez, E. & Clemente-Blanco, A. Role of protein phosphatases PP1, PP2A, PP4 and Cdc14 in the DNA damage response. Cell Stress 3, 70–85 (2019).
Nakada, S., Chen, G. I., Gingras, A. C. & Durocher, D. PP4 is a γH2AX phosphatase required for recovery from the DNA damage checkpoint. EMBO Rep. 9, 1019–1026 (2008).
Chowdhury, D. et al. PP4–phosphatase complex dephosphorylates γ-H2AX generated during DNA replication. Mol. Cell 31, 33–46 (2008).
Merigliano, C. et al. A role for the twins protein phosphatase (PP2A–B55) in the maintenance of Drosophila genome integrity. Genetics 205, 1151–1167 (2017).
Keogh, M. C. et al. A phosphatase complex that dephosphorylates γH2AX regulates DNA damage checkpoint recovery. Nature 439, 497–501 (2006).
O’Neill, B. M. et al. Pph3–Psy2 is a phosphatase complex required for Rad53 dephosphorylation and replication fork restart during recovery from DNA damage. Proc. Natl Acad. Sci. USA 104, 9290–9295 (2007).
Liu, J. et al. Protein phosphatase PP4 is involved in NHEJ-mediated repair of DNA double-strand breaks. Cell Cycle 11, 2643–2649 (2012).
Falk, J. E., Chan, A. C., Hoffmann, E. & Hochwagen, A. A Mec1- and PP4-dependent checkpoint couples centromere pairing to meiotic recombination. Dev. Cell 19, 599–611 (2010).
Pérez-Callejón, E. et al. Identification and molecular cloning of two homologues of protein phosphatase X from Arabidopsis thaliana. Plant Mol. Biol. 23, 1177–1185 (1993).
Moorhead, G. B. G., De Wever, V., Templeton, G. & Kerk, D. Evolution of protein phosphatases in plants and animals. Biochem. J. 417, 401–409 (2009).
Su, C. et al. The protein phosphatase 4 and SMEK1 complex dephosphorylates HYL1 to promote miRNA biogenesis by antagonizing the MAPK cascade in Arabidopsis. Dev. Cell 41, 527–539 (2017).
de Felippes, F. F., Wang, J. & Weigel, D. MIGS: miRNA-induced gene silencing. Plant J. 70, 541–547 (2012).
Klimyuk, V. I. & Jones, J. D. AtDMC1, the Arabidopsis homologue of the yeast DMC1 gene: characterization, transposon-induced allelic variation and meiosis-associated expression. Plant J. 11, 1–14 (1997).
Lim, E. C. et al. DeepTetrad: high-throughput image analysis of meiotic tetrads by deep learning in Arabidopsis thaliana. Plant J. 101, 473–483 (2020).
Rowan, B. A., Patel, V., Weigel, D. & Schneeberger, K. Rapid and inexpensive whole-genome genotyping-by-sequencing for crossover localization and fine-scale genetic mapping. G3 (Bethesda) 5, 385–398 (2015).
Choi, K. et al. Recombination rate heterogeneity within Arabidopsis disease resistance genes. PLoS Genet. 12, e1006179 (2016).
Choi, K. et al. Nucleosomes and DNA methylation shape meiotic DSB frequency in Arabidopsis thaliana transposons and gene regulatory regions. Genome Res. 28, 532–546 (2018).
Zhang, J. et al. A multiprotein complex regulates interference-sensitive crossover formation in rice. Plant Physiol. 181, 221–235 (2019).
Macaisne, N., Vignard, J. & Mercier, R. SHOC1 and PTD form an XPF–ERCC1-like complex that is required for formation of class I crossovers. J. Cell Sci. 124, 2687–2691 (2011).
Ueki, Y. et al. A consensus binding motif for the PP4 protein phosphatase. Mol. Cell 76, 953–964 (2019).
Fernandes, J. B., Seguela-Arnaud, M., Larcheveque, C., Lloyd, A. H. & Mercier, R. Unleashing meiotic crossovers in hybrid plants. Proc. Natl Acad. Sci. USA 115, 2431–2436 (2017).
Wijnker, E. et al. The Cdk1/Cdk2 homolog CDKA;1 controls the recombination landscape in Arabidopsis. Proc. Natl Acad. Sci. USA 116, 12534–12539 (2019).
He, Y. et al. Genomic features shaping the landscape of meiotic double-strand-break hotspots in maize. Proc. Natl Acad. Sci. USA 114, 12231–12236 (2017).
Liu, S. et al. Mu transposon insertion sites and meiotic recombination events co-localize with epigenetic marks for open chromatin across the maize genome. PLoS Genet. 5, e1000733 (2009).
Underwood, C. J. et al. Epigenetic activation of meiotic recombination near Arabidopsis thaliana centromeres via loss of H3K9me2 and non-CG DNA methylation. Genome Res. 28, 519–531 (2018).
Chelysheva, L. et al. The Arabidopsis HEI10 is a new ZMM protein related to Zip3. PLoS Genet. 8, e1002799 (2012).
Wang, K. et al. The role of rice HEI10 in the formation of meiotic crossovers. PLoS Genet. 8, e1002809 (2012).
Reynolds, A. et al. RNF212 is a dosage-sensitive regulator of crossing-over during mammalian meiosis. Nat. Genet. 45, 269–278 (2013).
Qiao, H. et al. Antagonistic roles of ubiquitin ligase HEI10 and SUMO ligase RNF212 regulate meiotic recombination. Nat. Genet. 46, 194–199 (2014).
Woglar, A. & Villeneuve, A. M. Dynamic architecture of DNA repair complexes and the synaptonemal complex at sites of meiotic recombination. Cell 173, 1678–1691 (2018).
Snowden, T., Acharya, S., Butz, C., Berardini, M. & Fishel, R. hMSH4–hMSH5 recognizes Holliday junctions and forms a meiosis-specific sliding clamp that embraces homologous chromosomes. Mol. Cell 15, 437–451 (2004).
Jessop, L., Rockmill, B., Roeder, G. S. & Lichten, M. Meiotic chromosome synapsis-promoting proteins antagonize the anti-crossover activity of sgs1. PLoS Genet. 2, e155 (2006).
Oh, S. D., Lao, J. P., Taylor, A. F., Smith, G. R. & Hunter, N. RecQ helicase, Sgs1, and XPF family endonuclease, Mus81–Mms4, resolve aberrant joint molecules during meiotic recombination. Mol. Cell 31, 324–336 (2008).
Manhart, C. M. et al. The mismatch repair and meiotic recombination endonuclease Mlh1–Mlh3 is activated by polymer formation and can cleave DNA substrates in trans. PLoS Biol. 15, e2001164 (2017).
Zakharyevich, K., Tang, S., Ma, Y. & Hunter, N. Delineation of joint molecule resolution pathways in meiosis identifies a crossover-specific resolvase. Cell 149, 334–347 (2012).
Ranjha, L., Anand, R. & Cejka, P. The Saccharomyces cerevisiae Mlh1–Mlh3 heterodimer is an endonuclease that preferentially binds to Holliday junctions. J. Biol. Chem. 289, 5674–5686 (2014).
Wijeratne, A. J., Chen, C., Zhang, W., Timofejeva, L. & Ma, H. The Arabidopsis thaliana parting dancers gene encoding a novel protein is required for normal meiotic homologous recombination. Mol. Biol. Cell 17, 1331–1343 (2006).
Lu, P., Wijeratne, A. J., Wang, Z., Copenhaver, G. P. & Ma, H. Arabidopsis PTD is required for type I crossover formation and affects recombination frequency in two different chromosomal regions. J. Genet. Genomics 41, 165–175 (2014).
De Muyt, A. et al. A meiotic XPF–ERCC1-like complex recognizes joint molecule recombination intermediates to promote crossover formation. Genes Dev. 32, 283–296 (2018).
Arora, K. & Corbett, K. D. The conserved XPF:ERCC1-like Zip2:Spo16 complex controls meiotic crossover formation through structure-specific DNA binding. Nucleic Acids Res. 47, 2365–2376 (2019).
Sato-Carlton, A. et al. Protein phosphatase 4 promotes chromosome pairing and synapsis, and contributes to maintaining crossover competence with increasing age. PLoS Genet. 10, e1004638 (2014).
Henderson, K. A., Kee, K., Maleki, S., Santini, P. A. & Keeney, S. Cyclin-dependent kinase directly regulates initiation of meiotic recombination. Cell 125, 1321–1332 (2006).
Lam, I. & Keeney, S. Mechanism and regulation of meiotic recombination initiation. Cold Spring Harb. Perspect. Biol. 7, a016634 (2014).
Valentin, G., Schwob, E. & Della Seta, F. Dual role of the Cdc7-regulatory protein Dbf4 during yeast meiosis. J. Biol. Chem. 281, 2828–2834 (2006).
Sasanuma, H. et al. Cdc7-dependent phosphorylation of Mer2 facilitates initiation of yeast meiotic recombination. Genes Dev. 22, 398–410 (2008).
Wan, L. et al. Cdc28–Clb5 (CDK-S) and Cdc7–Dbf4 (DDK) collaborate to initiate meiotic recombination in yeast. Genes Dev. 22, 386–397 (2008).
Matos, J. et al. Dbf4-Dependent Cdc7 kinase links DNA replication to the segregation of homologous chromosomes in meiosis I. Cell 135, 662–678 (2008).
Chen, X. et al. Phosphorylation of the synaptonemal complex protein Zip1 regulates the crossover/noncrossover decision during yeast meiosis. PLoS Biol. 13, e1002329 (2015).
Keeney, S., Lange, J. & Mohibullah, N. Self-organization of meiotic recombination initiation: general principles and molecular pathways. Annu. Rev. Genet. 48, 187–214 (2014).
Nowack, M. K. et al. Genetic framework of cyclin-dependent kinase function in Arabidopsis. Dev. Cell 22, 1030–1040 (2012).
Culligan, K. M. & Britt, A. B. Both ATM and ATR promote the efficient and accurate processing of programmed meiotic double-strand breaks. Plant J. 55, 629–638 (2008).
Garcia, V. et al. AtATM is essential for meiosis and the somatic response to DNA damage in plants. Plant Cell 15, 119–132 (2003).
Yao, Y. et al. ATM promotes RAD51-mediated meiotic DSB repair by inter-sister-chromatid recombination in Arabidopsis. Front. Plant Sci. 11, 839 (2020).
Villoria, M. T. et al. PP4 phosphatase cooperates in recombinational DNA repair by enhancing double-strand break end resection. Nucleic Acids Res. 47, 10706–10727 (2019).
Chelysheva, L. et al. Zip4/Spo22 is required for class I CO formation but not for synapsis completion in Arabidopsis thaliana. PLoS Genet. 3, e83 (2007).
van Tol, N., Rolloos, M., van Loon, P. & van der Zaal, B. J. MeioSeed: a CellProfiler-based program to count fluorescent seeds for crossover frequency analysis in Arabidopsis thaliana. Plant Methods 14, 32 (2018).
Carpenter, A. E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Zhang, X., Henriques, R., Lin, S. S., Niu, Q. W. & Chua, N. H. Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method. Nat. Protoc. 1, 641–646 (2006).
Chelysheva, L. et al. An easy protocol for studying chromatin and recombination protein dynamics during Arabidopsis thaliana meiosis: immunodetection of cohesins, histones and MLH1. Cytogenet. Genome Res. 129, 143–153 (2010).
Lambing, C., Kuo, P. C., Tock, A. J., Topp, S. D. & Henderson, I. R. ASY1 acts as a dosage-dependent antagonist of telomere-led recombination and mediates crossover interference in Arabidopsis. Proc. Natl Acad. Sci. USA 24, 13647–13658 (2020).
Sanchez-Moran, E., Santos, J.-L., Jones, G. H. & Franklin, F. C. H. ASY1 mediates AtDMC1-dependent interhomolog recombination during meiosis in Arabidopsis. Genes Dev. 21, 2220–2233 (2007).
Higgins, J. D., Sanchez-Moran, E., Armstrong, S. J., Jones, G. H. & Franklin, F. C. H. The Arabidopsis synaptonemal complex protein ZYP1 is required for chromosome synapsis and normal fidelity of crossing over. Genes Dev. 19, 2488–2500 (2005).
Hwang, I. & Sheen, J. Two-component circuitry in Arabidopsis cytokinin signal transduction. Nature 413, 383–389 (2001).
Xue, Y. et al. GPS 2.1: Enhanced prediction of kinase-specific phosphorylation sites with an algorithm of motif length selection. Protein Eng. Des. Sel. 24, 255–260 (2011).
Acknowledgements
We thank G. Copenhaver, A. Levy and S. Poethig for FTLs/CTLs, R. Mercier for fancm-1, L. Ziolkowska and C. Underwood for helping grow the EMS population, M. Grelon for MLH1 antibodies, C. Franklin for ASY1, ZYP1 and RAD51 antibodies and the Gurdon Institute for access to microscopes. This work was funded by the Suh Kyungbae Foundation (Jaeil Kim, Juhyun Kim, J.P., E.-J.K., H.K., D.B., Y.M.P. and K.C.), Next-Generation BioGreen 21 Program PJ01337001 (Jaeil Kim, Juhyun Kim, J.P., E.-J.K., H.K., D.B., Y.M.P. and K.C.) and PJ01342301 (H.S.C., S.L. and I.H.), Rural Development Administration, Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education NRF-2020R1A2C2007763 (H.K., D.B. and K.C.), Marie Curie International Training Network ‘COMREC’ (DN), Biotechnology and Biological Sciences Research Council grant EpiSpiX BB/N007557/1 (X.Z. and I.H.), Biotechnology and Biological Sciences Research Council ERA-CAPs grant BB/M004937/1 (C.L. and I.H.) and European Research Council Consolidator Award ERC-2015-CoG-681987 ‘SynthHotSpot’ (C.L., A.J.T. and I.H.).
Author information
Authors and Affiliations
Contributions
Design and conception of experiments: D.C.N., Jaeil Kim, C.L., Juhyun Kim, J.P., E.-J.K., P.K., K.C. and I.H. Acquisition and analysis of data: D.C.N., Jaeil Kim, C.L., Juhyun Kim, J.P., H.S.C., H.K., D.B., Y.M.P., P.K., S.L., A.J.T., X.Z., I.H. and K.C. Writing of the manuscript: D.C.N., Jaeil Kim, C.L., Juhyun Kim, K.C. and I.H.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks the anonymous reviewers for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 420 crossover frequency in wild type and M2 plants derived from the EMS population.
Box and whisker plot showing 420 crossover frequency (cM) for wild type (Col/Col) 420/++ plants (n=75) and EMS-treated M2 420/++ plants (n=1,217). Black dots indicate 420 crossover frequency in individual plants. Horizontal lines of black (wild type, Col) and red (EMS M2) box plots represent maximum, 3rd quartile, median, 1st quartile and minimum in 420 cM, respectively. In this study, wild type plants show a mean value of 19.5 cM (standard deviation=1.5) within 420, and the majority (81.4%, 991/1,217) of M2 plants display 420 crossover frequency within the range of 18-22 cM (Mean=21.4 cM, SD=1.5). 420 crossover frequency in M2 plants was significantly increased compared to wild type (one-sided Welch’s t-test P=2.2×10−16), which may have been caused by heterozygous EMS polymorphisms.
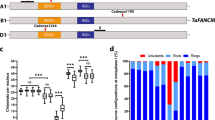
Extended Data Fig. 2 EMS mutations identified in FANCM (hcr4) and TAF4b (lcr1).
a, FANCM gene structure is shown, including the EMS mutation site in hcr4/fancm-11. The red arrow indicates the G to A substitution within exon 15, which causes a G to S amino acid substitution. Exons are shown as boxes (black=CDS, grey=UTR). Scale bar=0.5 kb. b, Multiple sequence alignment of the DEHDc (blue line) and HELICC (green line) domains of FANCM in different species. The mutation positions of the fancm-1 to fancm-10 alleles that were previously identified, and fancm-11 (hcr4), are shown. The fancm-11 mutation is located in a conserved motif within the SF2 helicase domain (bold arrow). c, Gene structure of TAF4b is shown with the location of the lcr1 (taf4b-3) mutation indicated in exon 3 (red arrow), which causes a premature stop codon.
Extended Data Fig. 3 T-DNA insertions in Arabidopsis PP4/PPX complex genes.
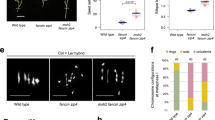
a, The gene structures of PPX1 (At4g26720), PPX2 (At5g55260) and PP4R2 (At5g17070) are shown. Exons are shown as boxes (black=CDS, grey=UTR). Scale bar=0.5 kb. The EMS induced hcr1-1 mutation is located at the splice donor site of the 3rd intron, shown by the asterisk. The red arrows indicate the location of primers for RT-qPCR in PPX1 and PPX2. The hcr1-2 T-DNA (GK_651B07) insertion position in the 5´-UTR is indicated. The position of the ppx2-1 (GK_488H09), ppx2-2 (SALK_049725), and pp4r2 (SALK_093051) T-DNA insertions are shown, which are located in the 4th intron, 8th exon and 7th intron, respectively. The arrows spanning the ppx2 and pp4r2 T-DNA insertions indicate primer positions used for RT-PCR. b, RT-PCR amplification and quantification for PPX1, PPX2 and PP4R2 mRNA expression in wild type Col, hcr1-1, ppx1-2, ppx2-1 and pp4r2. Floral cDNA from two biological replicates were evaluated by RT-PCR amplification for PPX1, PPX2, PP4R2 (shown in a) and GAPC expression. RT-PCR amplicon sizes for wild type, hcr1-1, ppx1-2, ppx2-1, pp4r2 cDNAs and wild type genomic DNA (positive/negative control) are shown. c, Plot showing RT-qPCR enrichment of PPX1 and PPX2 in hcr1-1 and ppx2-1. Relative transcript levels of PPX1 and PPX2 were measured in wild type, hcr1-1, and ppx2-1 using qRT-PCR. TUB2 was used for normalization. The y axis indicates fold-enrichment of PPX1 and PPX2 transcript levels, compared to PPX1 and PPX2 in wild type. RT-qPCR reactions of two technical replicates for each of four biological samples were shown as dots. Mean values are indicated by horizontal lines. Significance between wild type and mutants was assessed by one-sided Welch’s t-tests. The P values between Col and hcr1-1 for PPX1 was 0.186, for PPX2 was 3.49×10−5; between Col and ppx2-1 for PPX1 was 1.77×10−10 and for PPX2 was 3.64×10−9. Asterisks indicate P<0.001. d, Photograph showing developmental phenotypes of wild type, hcr1-2, hcr1-1, ppx2-1, hcr1-2 ppx2-1 and hcr1-1 ppx2-1 grown alongside one another. e, Photograph showing seeds of wild type and hcr1-1/+ ppx2-2/+ plants. Asterisks indicate defective seeds. f, Photograph showing F2 seedlings grown from self-fertilization of F1 hcr1-1/+ ppx2-2/+ plants, with asterisks indicating developmentally delayed seedlings.
Extended Data Fig. 4 Alignment of PP4 homolog protein sequences from diverse eukaryotes.
a, Amino acid sequence alignment of AtPPX1, the predicted hcr1-1 truncated protein, AtPPX2 and PP4 homologs from different eukaryotic species. The predicted hcr1-1 truncated protein consisting of 143 residues is shown. The underlined region indicates amino acids generated due to the retention of the 3rd intron. Hash symbols indicate the locations of conserved PP4 catalytic motifs (GDXHG, GDXVDRG and GNHE) and the histidine (H) residues required for metal binding in C-terminal region. b, As for a, but showing percent identity of amino acid sequence between PP4 homologs.
Extended Data Fig. 5 Meiosis-specific knockdown of PPX1 and PPX2 in meiMIGS transgenic plants.
a, qRT-PCR analysis of PPX1/HCR1 and PPX2 transcripts in floral buds of wild type and meiMIGS-PPX1, meiMIGS-PPX2 and meiMIGS-PPX1-PPX2 T2 transgenic lines. The y axis indicates fold-enrichment of PPX1 and PPX2 transcripts, compared to PPX1 in wild type. DMC1 was used as a meiotic gene for normalization. Replicate measurements are shown as dots and mean values shown by horizontal lines. b, Correlation between PPX1 and PPX2 transcript levels in wild type, meiMIGS-PPX1, meiMIGS-PPX2, and meiMIGS-PPX1-PPX2 lines. The x and y axis indicate relative PPX1 and PPX2 transcript levels in meiMIGS-PPX1 (blue), meiMIGS-PPX2 (red), and meiMIGS-PPX1-PPX2 (green) lines respectively, compared to PPX1 and PPX2 expressions in wild type Col plant. The correlation coefficient between PPX1 and PPX2 expression values across these samples was r=0.80, which was significantly different than expected if there were no relationship (P value=1.21×10−5).
Extended Data Fig. 6 Crossover frequency and interference measured in wild type and hcr1-1 using fluorescent pollen.
a, Crossover frequency measured using the pollen FTLs I1bc and I3bc from wild type and hcr1-1. Crossover frequency in each interval of the three-color FTLs was measured using the DeepTetrad pipeline3 (Supplementary Table 20). b, Crossover interference ratio measured using FTL pollen tetrads in wild type and hcr1-1. Crossover interference ratio (IFR) were calculated using the DeepTetrad pipeline. c, Plots showing the % of tetrads containing double crossovers, using data from the three-color FTL intervals in wild type and hcr1-1. d, As for c, but showing FTL data from the I1bc, I1fg, I3bc and I5ab intervals in wild type and meiMIGS-PPX1-PPX2. Tetrads were classified into 12 fluorescence classes (A-L) by DeepTetrad. Mean values are indicated by horizontal lines.
Extended Data Fig. 7 SPO11-1-oligonucleotides and nucleosome occupancy around wild type and meiMIGS-PPX1-PPX2 crossovers.
10 kb windows surrounding crossover midpoints identified from wild type or meiMIGS-PPX1-PPX2 plants, or the same number of randomly selected positions, were analysed for SPO11-1-oligos (log2(SPO11-1-oligos/gDNA), red) or nucleosome occupancy (log2(MNase-seq/gDNA), blue).
Extended Data Fig. 8 Yeast two hybrid assays showing interactions of HCR1/PPX1 with meiotic proteins.
a, Yeast two hybrid assays testing interaction between HCR1/PPX1 and Class I (ZMM) proteins. The yeast co-transformants were grown until OD600 = 1 and spotted on synthetic dropout media (SD) lacking leucine/tryptophan (-LT) and leucine/trptophan/histidine/adenine (-LTHA) for 3, 5 or 7 days. b, Yeast two hybrid assays of HCR1/PPX1 and meiotic proteins involved in axis formation, DSB formation and DNA repair. The yeast transformants were grown until OD600 = 1, then diluted 10-, 100- and 1,000-fold in water, and spotted on SD (-LT) and SD (-LTHA) plates to examine growth in 3, 5, or 7 days (Supplementary Table 23).
Extended Data Fig. 9 The EVH1 domain of Arabidopsis PP4R3A interacts with meiotic proteins.
a, Yeast two-hybrid assays testing interaction between the PP4R3A EVH1 domain and meiotic proteins. PP4R3A-N indicates the PP4R3A N-terminal region (1-166 aa) containing the EVH1 domain. The yeast co-transformants were grown until OD600 = 1 and spotted on synthetic dropout media (SD) lacking leucine/tryptophan (-LT) and leucine/trptophan/histidine/adenine (-LTHA) for 3 and 5 days. The yeast transformants were grown until OD600 = 1, then diluted 10-, 100- and 1,000-fold in water, and spotted on SD (-LT) and SD (-LTHA) plates to examine growth. b, Venn diagram summarizing yeast two hybrid assays of meiotic proteins that interact with HCR1/PPX1 and the PP4R3A EVH1 domain. c, A schematic model of Arabidopsis PP4 holoenzyme complex that recognizes target protein HEI10 for dephosphorylation via the PP4R3A EVH1 domain and PPX1.
Extended Data Fig. 10 Genome-wide prediction of PP4 complex target proteins during meiosis.
a, Protein domain (green) structure of Arabidopsis PP4 subunits PPX1, PPX2, PP4R2 and PP4R3. b, Amino acid alignment of the PP4R3A homolog EVH1 domain (red box). Hash symbols (#) indicate conserved tyrosine (Y) and tryptophan (W) residues. c, As for a, but showing the positions of FxxP motifs and phosphorylation consensus sites in PTD, HEI10, MSH5 and MLH1. d, Venn diagram showing overlap of meiotically expressed, nuclear proteins with FxxP motifs. e, Venn diagram showing overlap of candidate PP4 target proteins with CDK, DDK or ATM/ATR kinase consensus motifs, predicted using GPS 5.0. The location of HCR1 Y2H interactors are indicated within the Venn diagram. f, Histogram showing a significant enrichment of proteins containing phosphorylation sites in the predicted 1,367 PP4 targets, compared to 1,000 sets of randomly chosen genes (n=1,367). The vertical red line indicates observed predicted PP4 target proteins containing phosphorylation sites, compared to the random sets (black lines). g, Gene ontology (GO) enrichment analysis of the predicted PP4 targets, using PANTHER (http://pantherdb.org/). Benjamini-Hochberg False Discovery Rate (FDR) correction was used for enrichment test.
Supplementary information
Supplementary Information
Supplementary Tables 1–24.
Source data
Source Data Fig. 6
Unprocessed western blots for Fig. 6f.
Source Data Extended Data Fig. 3
Unprocessed DNA gels for Extended Data Fig. 3b.
Rights and permissions
About this article
Cite this article
Nageswaran, D.C., Kim, J., Lambing, C. et al. HIGH CROSSOVER RATE1 encodes PROTEIN PHOSPHATASE X1 and restricts meiotic crossovers in Arabidopsis. Nat. Plants 7, 452–467 (2021). https://doi.org/10.1038/s41477-021-00889-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-00889-y
This article is cited by
-
Control of meiotic crossover interference by a proteolytic chaperone network
Nature Plants (2024)
-
The effect of DNA polymorphisms and natural variation on crossover hotspot activity in Arabidopsis hybrids
Nature Communications (2023)
-
Why do plants need the ZMM crossover pathway? A snapshot of meiotic recombination from the perspective of interhomolog polymorphism
Plant Reproduction (2023)
-
FANCM promotes class I interfering crossovers and suppresses class II non-interfering crossovers in wheat meiosis
Nature Communications (2022)