Abstract
Plants tailor their metabolism to environmental conditions, in part through the recognition of a wide array of self and non-self molecules. In particular, the perception of microbial or plant-derived molecular patterns by cell-surface-localized pattern recognition receptors (PRRs) induces pattern-triggered immunity, which includes massive transcriptional reprogramming1. An increasing number of plant PRRs and corresponding ligands are known, but whether plants tune their immune outputs to patterns of different biological origins or of different biochemical natures remains mostly unclear. Here, we performed a detailed transcriptomic analysis in an early time series focused to study rapid-signalling transcriptional outputs induced by well-characterized patterns in the model plant Arabidopsis thaliana. This revealed that the transcriptional responses to diverse patterns (independent of their origin, biochemical nature or type of PRR) are remarkably congruent. Moreover, many of the genes most rapidly and commonly upregulated by patterns are also induced by abiotic stresses, suggesting that the early transcriptional response to patterns is part of the plant general stress response (GSR). As such, plant cells’ response is in the first instance mostly to danger. Notably, the genetic impairment of the GSR reduces pattern-induced antibacterial immunity, confirming the biological relevance of this initial danger response. Importantly, the definition of a small subset of ‘core immunity response’ genes common and specific to pattern response revealed the function of previously uncharacterized GLUTAMATE RECEPTOR-LIKE (GLR) calcium-permeable channels in immunity. This study thus illustrates general and unique properties of early immune transcriptional reprogramming and uncovers important components of plant immunity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The RNA-seq datasets generated and analysed in the current study have been deposited in the ArrayExpress database at EMBL-EBI (www.ebi.ac.uk/arrayexpress) under accession number E-MTAB-9694. Markdowns documenting the steps in filtering, visualizing and analysing the data in all figures and tables are available in Supplementary Note 1. Source data are provided with this paper.
References
Yu, X., Feng, B., He, P. & Shan, L. From chaos to harmony: responses and signaling upon microbial pattern recognition. Annu. Rev. Phytopathol. 55, 109–137 (2017).
Albert, I., Hua, C., Nürnberger, T., Pruitt, R. N. & Zhang, L. Surface sensor systems in plant immunity. Plant Physiol. 182, 1582–1596 (2020).
Saijo, Y., Loo, E. P.-I. & Yasuda, S. Pattern recognition receptors and signaling in plant–microbe interactions. Plant J. 93, 592–613 (2018).
Denoux, C. et al. Activation of defense response pathways by OGs and Flg22 elicitors in Arabidopsis seedlings. Mol. Plant 1, 423–445 (2008).
Wan, W.-L. et al. Comparing Arabidopsis receptor kinase and receptor protein-mediated immune signaling reveals BIK1-dependent differences. N. Phytol. 221, 2080–2095 (2019).
Stringlis, I. A. et al. Root transcriptional dynamics induced by beneficial rhizobacteria and microbial immune elicitors reveal signatures of adaptation to mutualists. Plant J. 93, 166–180 (2018).
Wan, J. et al. A LysM receptor-like kinase plays a critical role in chitin signaling and fungal resistance in Arabidopsis. Plant Cell 20, 471–481 (2008).
Zipfel, C. et al. Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation. Cell 125, 749–760 (2006).
Gómez-Gómez, L. & Boller, T. FLS2: an LRR receptor-like kinase involved in the perception of the bacterial elicitor flagellin in Arabidopsis. Mol. Cell 5, 1003–1011 (2000).
Krol, E. et al. Perception of the Arabidopsis danger signal peptide 1 involves the pattern recognition receptor AtPEPR1 and its close homologue AtPEPR2. J. Biol. Chem. 285, 13471–13479 (2010).
Yamaguchi, Y., Huffaker, A., Bryan, A. C., Tax, F. E. & Ryan, C. A. PEPR2 is a second receptor for the Pep1 and Pep2 peptides and contributes to defense responses in Arabidopsis. Plant Cell 22, 508–522 (2010).
Yamaguchi, Y., Pearce, G. & Ryan, C. A. The cell surface leucine-rich repeat receptor for AtPep1, an endogenous peptide elicitor in Arabidopsis, is functional in transgenic tobacco cells. Proc. Natl Acad. Sci. USA 103, 10104–10109 (2006).
Albert, I. et al. An RLP23–SOBIR1–BAK1 complex mediates NLP-triggered immunity. Nat. Plants 1, 15140 (2015).
Cao, Y. et al. The kinase LYK5 is a major chitin receptor in Arabidopsis and forms a chitin-induced complex with related kinase CERK1. eLife 3, e03766 (2014).
Kutschera, A. et al. Bacterial medium-chain 3-hydroxy fatty acid metabolites trigger immunity in Arabidopsis plants. Science 364, 178–181 (2019).
Ranf, S. et al. A lectin S-domain receptor kinase mediates lipopolysaccharide sensing in Arabidopsis thaliana. Nat. Immunol. 16, 426–433 (2015).
Brutus, A., Sicilia, F., Macone, A., Cervone, F. & De Lorenzo, G. A domain swap approach reveals a role of the plant wall-associated kinase 1 (WAK1) as a receptor of oligogalacturonides. Proc. Natl Acad. Sci. USA 107, 9452–9457 (2010).
Navarro, L. et al. The transcriptional innate immune response to flg22: interplay and overlap with Avr gene-dependent defense responses and bacterial pathogenesis. Plant Physiol. 135, 1113–1128 (2004).
Libault, M., Wan, J., Czechowski, T., Udvardi, M. & Stacey, G. Identification of 118 Arabidopsis transcription factor and 30 ubiquitin-ligase genes responding to chitin, a plant-defense elicitor. Mol. Plant Microbe Interact. 20, 900–911 (2007).
Hu, X. Y., Neill, S. J., Cai, W. M. & Tang, Z. C. Induction of defence gene expression by oligogalacturonic acid requires increases in both cytosolic calcium and hydrogen peroxide in Arabidopsis thaliana. Cell Res. 14, 234–240 (2004).
Lex, A., Gehlenborg, N., Strobelt, H., Vuillemot, R. & Pfister, H. Upset: visualization of intersecting sets. IEEE Trans. Vis. Comput. Graph. 20, 1983–1992 (2014).
Jeworutzki, E. et al. Early signaling through the Arabidopsis pattern recognition receptors FLS2 and EFR involves Ca-associated opening of plasma membrane anion channels. Plant J. 62, 367–378 (2010).
Bjornson, M., Dandekar, A. & Dehesh, K. Determinants of timing and amplitude in the plant general stress response. J. Integr. Plant Biol. 58, 119–126 (2016).
Varala, K. et al. Temporal transcriptional logic of dynamic regulatory networks underlying nitrogen signaling and use in plants. Proc. Natl Acad. Sci. USA 115, 6494–6499 (2018).
Birkenbihl, R. P. et al. Principles and characteristics of the Arabidopsis WRKY regulatory network during early MAMP-triggered immunity. Plant J. 96, 487–502 (2018).
Doherty, C. J., Van Buskirk, H. A., Myers, S. J. & Thomashow, M. F. Roles for Arabidopsis CAMTA transcription factors in cold-regulated gene expression and freezing tolerance. Plant Cell 21, 972–984 (2009).
Benn, G. et al. A key general stress response motif is regulated non-uniformly by CAMTA transcription factors. Plant J. 80, 82–92 (2014).
Walley, J. W. et al. Mechanical stress induces biotic and abiotic stress responses via a novel cis-element. PLoS Genet. 3, 1800–1812 (2007).
Kilian, J. et al. The AtGenExpress global stress expression data set: protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses. Plant J. 50, 347–363 (2007).
Bilgin, D. D. et al. Biotic stress globally downregulates photosynthesis genes. Plant Cell Environ. 33, 1597–1613 (2010).
Göhre, V., Jones, A. M. E., Sklenář, J., Robatzek, S. & Weber, A. P. M. Molecular crosstalk between PAMP-triggered immunity and photosynthesis. Mol. Plant Microbe Interact. 25, 1083–1092 (2012).
Lolle, S. et al. Matching NLR immune receptors to autoimmunity in camta3 mutants using antimorphic NLR alleles. Cell Host Microbe 21, 518–529 (2017).
Jacob, F. et al. A dominant-interfering camta3 mutation compromises primary transcriptional outputs mediated by both cell surface and intracellular immune receptors in Arabidopsis thaliana. N. Phytol. 217, 1667–1680 (2018).
Yuan, P., Du, L. & Poovaiah, B. W. Ca2+/calmodulin-dependent AtSR1/CAMTA3 plays critical roles in balancing plant growth and immunity. Int. J. Mol. Sci. 19, 1764 (2018).
Du, L. et al. Ca2+/calmodulin regulates salicylic-acid-mediated plant immunity. Nature 457, 1154–1158 (2009).
Jiang, X., Hoehenwarter, W., Scheel, D. & Lee, J. Phosphorylation of the CAMTA3 transcription factor triggers its destabilization and nuclear export. Plant Physiol. https://doi.org/10.1104/pp.20.00795 (2020).
Gladman, N. P., Marshall, R. S., Lee, K.-H. & Vierstra, R. D. The proteasome stress regulon is controlled by a pair of NAC transcription factors in Arabidopsis. Plant Cell 28, 1279–1296 (2016).
van Veen, H. et al. Transcriptomes of eight Arabidopsis thaliana accessions reveal core conserved, genotype- and organ-specific responses to flooding stress. Plant Physiol. 172, 668–689 (2016).
Ding, F. et al. Genome-wide analysis of alternative splicing of pre-mRNA under salt stress in Arabidopsis. BMC Genomics 15, 431 (2014).
Chiu, J. C. et al. Phylogenetic and expression analysis of the glutamate-receptor-like gene family in Arabidopsis thaliana. Mol. Biol. Evol. 19, 1066–1082 (2002).
Toyota, M. et al. Glutamate triggers long-distance, calcium-based plant defense signaling. Science 361, 1112–1115 (2018).
Shao, Q., Gao, Q., Lhamo, D., Zhang, H. & Luan, S. Two glutamate- and pH-regulated Ca2+ channels are required for systemic wound signaling in Arabidopsis. Sci. Signal. 13, eaba1453 (2020).
Mousavi, S. A. R., Chauvin, A., Pascaud, F., Kellenberger, S. & Farmer, E. E. Glutamate Receptor-Like genes mediate leaf-to-leaf wound signalling. Nature 500, 422–426 (2013).
Kwaaitaal, M., Huisman, R., Maintz, J., Reinstädler, A. & Panstruga, R. Ionotropic glutamate receptor (iGluR)-like channels mediate MAMP-induced calcium influx in Arabidopsis thaliana. Biochem. J. 440, 355–365 (2011).
Schwessinger, B. et al. Phosphorylation-dependent differential regulation of plant growth, cell death, and innate immunity by the regulatory receptor-like kinase BAK1. PLoS Genet. 7, e1002046 (2011).
Thor, K. et al. The calcium-permeable channel OSCA1.3 regulates plant stomatal immunity. Nature 585, 569–573 (2020).
Moeder, W., Phan, V. & Yoshioka, K. Ca2+ to the rescue—Ca2+ channels and signaling in plant immunity. Plant Sci. 279, 19–26 (2019).
Tian, W. et al. A calmodulin-gated calcium channel links pathogen patterns to plant immunity. Nature 572, 131–135 (2019).
Lorek, J., Griebel, T., Jones, A. M., Kuhn, H. & Panstruga, R. The role of Arabidopsis heterotrimeric G-protein subunits in MLO2 function and MAMP-triggered immunity. Mol. Plant Microbe Interact. 26, 991–1003 (2013).
Gruner, K., Zeier, T., Aretz, C. & Zeier, J. A critical role for Arabidopsis mildew resistance locus O2 in systemic acquired resistance. Plant J. 94, 1064–1082 (2018).
Lu, H., Rate, D. N., Song, J. T. & Greenberg, J. T. ACD6, a novel ankyrin protein, is a regulator and an effector of salicylic acid signaling in the Arabidopsis defense response. Plant Cell 15, 2408–2420 (2003).
Liu, J. et al. Heterotrimeric G proteins serve as a converging point in plant defense signaling activated by multiple receptor-like kinases. Plant Physiol. 161, 2146–2158 (2013).
Zipfel, C. et al. Bacterial disease resistance in Arabidopsis through flagellin perception. Nature 428, 764–767 (2004).
Huffaker, A., Pearce, G. & Ryan, C. A. An endogenous peptide signal in Arabidopsis activates components of the innate immune response. Proc. Natl Acad. Sci. USA 103, 10098–10103 (2006).
Ridley, B. L., O’Neill, M. A. & Mohnen, D. Pectins: structure, biosynthesis, and oligogalacturonide-related signaling. Phytochemistry 57, 929–967 (2001).
Kumar, R. et al. A high-throughput method for Illumina RNA-seq library preparation. Front. Plant Sci. 3, 202 (2012).
Townsley, B. T., Covington, M. F., Ichihashi, Y., Zumstein, K. & Sinha, N. R. BrAD-seq: breath adapter directional sequencing: a streamlined, ultra-simple and fast library preparation protocol for strand specific mRNA library construction. Front. Plant Sci. 6, 366 (2015).
Picelli, S. et al. Tn5 transposase and tagmentation procedures for massively scaled sequencing projects. Genome Res. 24, 2033–2040 (2014).
Bjornson, M., Kajala, K., Zipfel, C. & Ding, P. Low-cost and high-throughput RNA-seq library preparation for Illumina sequencing from plant tissue. Bio-protocol 10, e3799 (2020).
Rohland, N. & Reich, D. Cost-effective, high-throughput DNA sequencing libraries for multiplexed target capture. Genome Res. 22, 939–946 (2012).
Leggate, J., Allain, R., Isaac, L. & Blais, B. W. Microplate fluorescence assay for the quantification of double stranded DNA using SYBR Green I dye. Biotechnol. Lett. 28, 1587–1594 (2006).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
FastQC: a quality control tool for high throughput sequence data (Babraham Bioinformatics Institute, 2010); http://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
R Core Team. R: a language and environment for statistical computing (R Foundation for Statistical Computing, 2020).
RStudio: integrated development environment for R (RStudio Team, 2020).
Wickham, H. et al. Welcome to the tidyverse. J. Open Source Softw. 4, 1686 (2019).
Conway, J. R., Lex, A. & Gehlenborg, N. UpSetR: an R package for the visualization of intersecting sets and their properties. Bioinformatics 33, 2938–2940 (2017).
Wang, M., Zhao, Y. & Zhang, B. Efficient test and visualization of multi-set intersections. Sci. Rep. 5, 16923 (2015).
Galili, T. dendextend: an R package for visualizing, adjusting and comparing trees of hierarchical clustering. Bioinformatics 31, 3718–3720 (2015).
Xiao, S.-J., Zhang, C., Zou, Q. & Ji, Z.-L. TiSGeD: a database for tissue-specific genes. Bioinformatics 26, 1273–1275 (2010).
Julca, I. et al. Comparative transcriptomic analysis reveals conserved transcriptional programs underpinning organogenesis and reproduction in land plants. Preprint at https://doi.org/10.1101/2020.10.29.361501 (2020).
Alexa, A. & Rahnenfuhrer, J. topGO: Enrichment analysis for gene ontology. R package version 2.42.0 (2020).
McLeay, R. C. & Bailey, T. L. Motif Enrichment Analysis: a unified framework and an evaluation on ChIP data. BMC Bioinform. 11, 165 (2010).
O’Malley, R. C. et al. Cistrome and epicistrome features shape the regulatory DNA landscape. Cell 165, 1280–1292 (2016).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Fan, J., Crooks, C. & Lamb, C. High-throughput quantitative luminescence assay of the growth in planta of Pseudomonas syringae chromosomally tagged with Photorhabdus luminescens luxCDABE. Plant J. 53, 393–399 (2008).
Lenth, R. emmeans: Estimated marginal means, aka least-squares means. R package version 1.5.4 (2020).
Mustroph, A. et al. Profiling translatomes of discrete cell populations resolves altered cellular priorities during hypoxia in Arabidopsis. Proc. Natl Acad. Sci. USA 106, 18843–18848 (2009).
Bates, G. W. et al. A comparative study of the Arabidopsis thaliana guard-cell transcriptome and its modulation by sucrose. PLoS ONE 7, e49641 (2012).
Ribeiro, D. M., Araújo, W. L., Fernie, A. R., Schippers, J. H. M. & Mueller-Roeber, B. Translatome and metabolome effects triggered by gibberellins during rosette growth in Arabidopsis. J. Exp. Bot. 63, 2769–2786 (2012).
Yang, Y., Costa, A., Leonhardt, N., Siegel, R. S. & Schroeder, J. I. Isolation of a strong Arabidopsis guard cell promoter and its potential as a research tool. Plant Methods 4, 6 (2008).
Acknowledgements
We thank P. Ding for sharing protocols and material related to Tn-5 tagmentation, S. Ranf for providing the sd1-29 mutant and 3-OH-FA pattern prior to publication, G. Stacey for providing the lyk4/5 mutant, C. J. S. Moreira for assistance in genotyping the glr2.7 2.8 2.9 CRISPR line, and past and present members of the Zipfel laboratory for helpful discussions. This research was supported by the Gatsby Charitable Foundation, the University of Zurich, the European Research Council under grant agreement nos. 309858 and 773153 (grants ‘PHOSPHOinnATE’ and ‘IMMUNO-PEPTALK’ to C.Z.), and the Swiss National Science Foundation (grant agreement no. 31003A_182625 to C.Z.). M.B. was partially supported by the European Union’s Horizon 2020 Research and Innovation Program under Marie Skłodowska-Curie Actions (grant agreement no. 703954).
Author information
Authors and Affiliations
Contributions
C.Z., T.N. and M.B. conceived and designed the experiments. C.Z. and M.B. obtained the funding. M.B. and P.P. performed the experiments and analysed the data. T.N. contributed conceptually to the study and provided the reagents. M.B. and C.Z. wrote the manuscript with feedback from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Susannah Tringe and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Quality control and exploratory analysis of RNA-seq data.
Expression changes in this study at a, 30 min and b, 3 h are plotted against previously published results for flg22, elf18/26, and chitooctaose (CO8). Linear correlation shown in red, with R2 (linear regression) shown on each plot. c, PCA analysis of log2(FC) of differentially expressed genes, showing (left) minimal changes in receptor-mutant treated plants, mostly corresponding with later time points, and rays of response (right) corresponding with plants at 30, 90, or 180 min post-treatment. d, Pearson correlation heatmap of DESeq2-calculated log2(FC) showing clustering largely by time point, with the strongest correlations at 30 min.
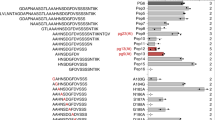
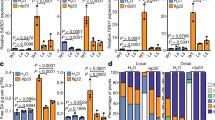
Extended Data Fig. 2 There is little specificity in pattern-induced genes.
Among induced genes, for each pattern a specificity measure (expression in response to pattern/total expression in experiment) was calculated, and genes with at least one SPM>0.33 (one pattern treatment responsible for approximately 1/3 total expression in study, n=412) were gathered. flg22 is the only pattern treatment with a large number of pattern-selective genes expressed (flg22: 332, elf18: 8, Pep1: 33, nlp20:31, OGs: 8, CO8 and 3-OH-FA:0).
Extended Data Fig. 3 Complete complement of set sizes among collapsed pattern-induced and pattern-repressed gene sets.
Each circular ‘track’ represents one pattern treatment; when filled the pattern in question alters the expression of the gene set shown at the perimeter. Gene set size is shown via bar height of bars surrounding pattern tracks, and bar color shows deviation: indicating whether the set size is larger or smaller than would be expected by chance. Large diagrams show the overall set complement of genes induced or repressed by patterns taking all time points into account, whereas smaller diagrams to the left and right are specific for the complement of genes induced or repressed at the indicated time point. No genes were significantly repressed at five minutes post-treatment. Selected pattern subset of a priori interest are highlighted through open arrows on large combined plots; none has deviation far from 0.
Extended Data Fig. 4 Pattern-responsive genes tend to be repressed by single patterns, though there does exist a core set of 93 genes repressed by all tested patterns.
A single set of genes repressed [log2(FC)<-1, p<0.05] by each pattern treatment was found through combining the lists genes repressed at each time. a, UpSet diagram showing the size of ‘collapsed’ gene sets repressed by each pattern (left) and the top 15 intersections (bottom right) by size (top right), colored by deviation from set size predicted by random mixing. b, Heat map of expression of the 93 genes repressed by all tested patterns. Genes are hierarchically clustered according to their behavior across all pattern/time combinations, and cut into three clusters. c, Visualization of average log2(FC) patterns of the three clusters identified in b, showing different patterns of expression. Error bars represent standard error of the mean.
Extended Data Fig. 5 Pattern-triggered transcriptional repression acts in time-resolved waves.
a, GO term and b, cis-element enrichment analysis of repressed genes, categorized according to the time point at which they first passed significance threshold, regardless of which pattern caused repression. The top three GO terms for each time point are indicated. c, Distribution of repressed genes. Each gene repressed in this study was plotted according to the time it is first repressed (panels from top to bottom), the number of tested patterns which repress it (x axis) and the number of abiotic stresses in the AtGenExpress dataset which also repress it within the first 3 h (y axis). The color of each dot indicates the most negative log2(FC) observed in this study.
Extended Data Fig. 6 CRISPR deletes the majority of the GLR2.7/2.8/2.9 genomic region in assayed lines.
Schematic of the GLR2.7/2.8/2.9 genomic region, with deletions in a Col-0, b and c YC3.6 background. In each case, a ‘fusion protein’ may be transcribed, consisting of approximately 90 (92, 92, 89) amino acids of GLR2.7, fused to approximately 12 (12, 13, 12) nonsense amino acids from the GLR2.9 genomic region. The potential fusion protein does not encode any transmembrane domains. GLR exons are represented by colored boxes, introns by grey boxes, and intergenic regions by black lines. Neighboring genes shown in black. Arrows represent direction of transcription.
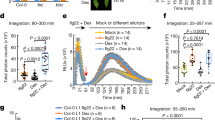
Extended Data Fig. 7 Characterization of glr2.7/2.8/2.9 lines.
a, b, c, Increase in intracellular Ca2+ concentration in response to treatment in seedlings (a, c) or leaf discs (b). Shown are mean corrected YFP/CFP ratio within 25 min (a, b), 1 min, or 5 min (c) post-treatment (timepoint 0) +/1 standard error of the mean. Data were collected every 30 s (a, b), 5 s, or 12 s (c). For a and c corresponding peak values are shown in Fig. 3, for b peak values are shown to the right of response curve. In (b), each point represents peak ratio of YFP to CFP (proportional to Ca2+ concentration) for a single seedling, normalized to initial ratio. Different shapes represent 2 independent experiments, n=11-62 for each experiment/line/treatment combination. Statistical tests were performed in R, two-way ANOVA blocking by experiment. d, Stomatal aperture of WT, glr2.7/2.8/2.9, or flg22-hyporesponsive bak1-5 plants treated with water, 5 µM flg22, or 10 µM ABA. Each point represents one stoma, and plot represents stomata from a total of 12 plants assayed over 5 experiments (n=36-178 stomata per genotype/treatment/experiment). Statistical tests were performed in R, two-way ANOVA blocking by experiment. Post-hoc tests were performed using the emmeans package in R: within each genotype, stomatal aperture was compared with mock treatment with dunnettx multiple testing correction. In spray infection assays glr2.7/2.8/2.9 are not more susceptible to e WT Pto DC3000, or f Pto COR−, deficient in the stomata-opening toxin coronatine. Bacteria were harvested from leaf discs two days post-inoculation; each point represents one plant, and shapes represent three independent experiments (n=6 plants per genotype/treatment/experiment). Statistics were performed in R: one-way ANOVA blocking by experiment followed by dunnettx multiple comparison to Col-0 performed using the emmeans package. Box plots center on the median, with box extending to the first and third quartile, and whiskers extending to the lesser value of the furthest point or 1.5x the inter-quartile range.
Extended Data Fig. 8 Leaf tissue expression patterns of genes encoding calcium-permeable channels implicated in PTI.
Data collected from Genevestigator, and scaled by each experiment.
Extended Data Fig. 9 AT3G32090 is likely miscalled as expressed in response to patterns.
AT3G032090 is among the CIR set, (a), but all reads assigned to this gene map to a single exon (b, image from integrated genomics viewer), not the pattern expected from poly-A purification of mRNA. c, The top BLAST hit for each exon of AT3G032090 are shown, with strong similarity to WRKY40 (AT1G80840) in the ‘expressed’ exon of AT3G032090. d, WRKY40 is strongly expressed (note y axis) and strongly pattern-induced. A small fraction of mis-aligned reads likely account for the observed pattern AT3G032090.
Supplementary information
Supplementary Information
Supplementary Note 1.
Supplementary Table 1
log2(FC) (relative to time 0 for each genotype–treatment combination) across the genome for each genotype–treatment–time combination in this experiment.
Supplementary Table 2
Adjusted P value (FDR corrected, calculated by DESeq2) across the genome for each genotype–treatment–time combination in this experiment.
Supplementary Tables 3–6
Information on 1,000 genes commonly upregulated by all tested elicitors, 100 genes commonly downregulated by all tested elicitors, specificity measure of genes induced selectively by only one elicitor and CIR gene set.
Source data
Source Data Fig. 2
Count elicitors and abiotic stresses for Fig. 2c.
Source Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 5
Count elicitors and abiotic stresses for Fig. 5c.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Bjornson, M., Pimprikar, P., Nürnberger, T. et al. The transcriptional landscape of Arabidopsis thaliana pattern-triggered immunity. Nat. Plants 7, 579–586 (2021). https://doi.org/10.1038/s41477-021-00874-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-00874-5
This article is cited by
-
Modulation of early gene expression responses to water deprivation stress by the E3 ubiquitin ligase ATL80: implications for retrograde signaling interplay
BMC Plant Biology (2024)
-
Mechanisms of calcium homeostasis orchestrate plant growth and immunity
Nature (2024)
-
Signal Peptide Peptidase and PI4Kβ1/2 play opposite roles in plant ER stress response and immunity
Stress Biology (2024)
-
Rice NLR protein XinN1, induced by a pattern recognition receptor XA21, confers enhanced resistance to bacterial blight
Plant Cell Reports (2024)
-
How plants manage pathogen infection
EMBO Reports (2023)