Abstract
The unparalleled performance of Chlorella ohadii under irradiances of twice full sunlight underlines the gaps in our understanding of how the photosynthetic machinery operates, and what sets its upper functional limit. Rather than succumbing to photodamage under extreme irradiance, unique features of photosystem II function allow C. ohadii to maintain high rates of photosynthesis and growth, accompanied by major changes in composition and cellular structure. This remarkable resilience allowed us to investigate the systems response of photosynthesis and growth to extreme illumination in a metabolically active cell. Using redox proteomics, transcriptomics, metabolomics and lipidomics, we explored the cellular mechanisms that promote dissipation of excess redox energy, protein S-glutathionylation, inorganic carbon concentration, lipid and starch accumulation, and thylakoid stacking. C. ohadii possesses a readily available capacity to utilize a sudden excess of reducing power and carbon for growth and reserve formation, and post-translational redox regulation plays a pivotal role in this rapid response. Frequently the response in C. ohadii deviated from that of model species, reflecting its life history in desert sand crusts. Comparative global and case-specific analyses provided insights into the potential evolutionary role of effective reductant utilization in this extreme resistance of C. ohadii to extreme irradiation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Chorella ohadii genome annotation has been deposited in the NCBI/Genbank database with accession no. PRJNA573576. Mass spectrometry proteomics data have been deposited in the ProteomeXchange Consortium via the PRIDE partner repository, with the dataset identifier PXD015681. Metabolite and lipid profiling data are provided in the Supplementary Information as indicated in the main text.
References
Treves, H. et al. A newly isolated Chlorella sp. from desert sand crusts exhibits a unique resistance to excess light intensity. FEMS Microbiol. Ecol. 86, 373–380 (2013).
Treves, H. et al. The mechanisms whereby the green alga Chlorella ohadii, isolated from desert soil crust, exhibits unparalleled photodamage resistance. N. Phytol. 210, 1229–1243 (2016).
Treves, H. et al. Metabolic flexibility underpins growth capabilities of the fastest growing alga. Curr. Biol. 27, 2559–2567 (2017).
Ohad, I. et al. Light-induced changes within photosystem II protects Microcoleus sp. in biological desert sand crusts against excess light. PLoS ONE 5, e11000 (2010).
Aro, E.-M., Virgin, I. & Andersson, B. Photoinhibition of photosystem II. Inactivation, protein damage and turnover. Biochim. Biophys. Acta 1143, 113–134 (1993).
Keren, N. & Krieger-Liszkay, A. Photoinhibition: molecular mechanisms and physiological significance. Physiol. Plant. 142, 1–5 (2011).
Raven, J. A. The cost of photoinhibition. Physiol. Plant. 142, 87–104 (2011).
Miller, G. & Mittler, R. Could heat shock transcription factors function as hydrogen peroxide sensors in plants? Ann. Bot. 98, 279–288 (2006).
Ahn, S. G. & Thiele, D. J. Redox regulation of mammalian heat shock factor 1 is essential for Hsp gene activation and protection from stress. Genes Dev. 17, 516–528 (2003).
Fedoroff, N. Redox regulatory mechanisms in cellular stress responses. Ann. Bot. 98, 289–300 (2006).
Sorger, P. K. & Pelham, H. R. B. Yeast heat-shock factor is an essential DNA-binding protein that exhibits temperature-dependent phosphorylation. Cell 54, 855–864 (1988).
Park, H. O. & Craig, E. A. Positive and negative regulation of basal expression of a yeast HSP70 gene. Mol. Cell. Biol. 9, 2025–2033 (1989).
Morimoto, R. I. Regulation of the heat shock transcriptional response: cross talk between a family of heat shock factors, molecular chaperones, and negative regulators. Genes Dev. 12, 3788–3796 (1998).
Foyer, C. H. & Noctor, G. Redox regulation in photosynthetic organisms: signaling, acclimation, and practical implications. Antioxid. Redox Signal. 11, 861–905 (2009).
Suzuki, N. et al. ROS and redox signalling in the response of plants to abiotic stress. Plant Cell Environ. 35, 259–270 (2012).
Ananyev, G. et al. Photosystem II-cyclic electron flow powers exceptional photoprotection and record growth in the microalga Chlorella ohadii. Biochim. Biophys. Acta Bioenerg. 1858, 873–883 (2017).
Murik, O., Elboher, A. & Kaplan, A. Dehydroascorbate: a possible surveillance molecule of oxidative stress and programmed cell death in the green alga Chlamydomonas reinhardtii. N. Phytol. 202, 471–484 (2014).
Khatoon, M. et al. Quality control of photosystem II: thylakoid unstacking is necessary to avoid further damage to the D1 protein and to facilitate D1 degradation under light stress in spinach thylakoids. J. Biol. Chem. 284, 25343–25352 (2009).
Anderson, J. M. Photoregulation of the composition, function, and structure of thylakoid membranes. Annu. Rev. Plant Physiol. 37, 93–136 (1986).
Kirchhoff, H. Architectural switches in plant thylakoid membranes. Photosynth. Res. 116, 481–487 (2013).
Dormann, P. et al. Isolation and characterization of an Arabidopsis mutant deficient in the thylakoid lipid digalactosyl diacylglycerol. Plant Cell 7, 1801–1810 (1995).
Hölzl, G. et al. The role of diglycosyl lipids in photosynthesis and membrane lipid homeostasis in Arabidopsis. Plant Physiol. 150, 1147–1159 (2009).
Deme, B. et al. Contribution of galactoglycerolipids to the 3-dimensional architecture of thylakoids. FASEB J. 28, 3373–3383 (2014).
Kaplan, A. & Reinhold, L. The CO2 concentrating mechanisms in photosynthetic microorganisms. Annu. Rev. Plant Physiol. Plant Mol. Biol. 50, 539–570 (1999).
Giordano, M., Beardall, J. & Raven, J. A. CO2 concentrating mechanisms in algae: mechanisms, environmental modulation, and evolution. Ann. Rev. Plant Biol. 56, 99–131 (2005).
Ramazanov, Z. et al. The induction of the CO2-concentrating mechanism is correlated with the formation of the starch sheath around the pyrenoid of Chlamydomonas reinhardtii. Planta 195, 210–216 (1994).
Küken, A. et al. Effects of microcompartmentation on flux distribution and metabolic pools in Chlamydomonas reinhardtii chloroplasts. eLife 7, e37960 (2018).
Mettler, T. et al. Systems analysis of the response of photosynthesis, metabolism, and growth to an increase in irradiance in the photosynthetic model organism Chlamydomonas reinhardtii. Plant Cell 26, 2310–2350 (2014).
Schmollinger, S. et al. Nitrogen-sparing mechanisms in Chlamydomonas affect the transcriptome, the proteome, and photosynthetic metabolism. Plant Cell 26, 1410–1435 (2014).
Blaby, I. K. et al. Systems-level analysis of nitrogen starvation-induced modifications of carbon metabolism in a Chlamydomonas reinhardtii starchless mutant. Plant Cell 25, 4305–4323 (2013).
Kimura, M. et al. Identification of Arabidopsis genes regulated by high light-stress using cDNA microarray. Photochem. Photobiol. 77, 226–233 (2003).
Murchie, E. H. et al. Acclimation of photosynthesis to high irradiance in rice: gene expression and interactions with leaf development. J. Exp. Bot. 56, 449–460 (2005).
Im, C. S. et al. Analysis of light and CO2 regulation in Chlamydomonas reinhardtii using genome-wide approaches. Photosynth. Res. 75, 111–125 (2003).
Ritz, M. et al. Kinetics of photoacclimation in response to a shift to high light of the red alga Rhodella violacea adapted to low irradiance. Plant Physiol. 123, 1415–1425 (2000).
Deblois, C. P., Marchand, A. & Juneau, P. Comparison of photoacclimation in twelve freshwater photoautotrophs (Chlorophyte, Bacillaryophyte, Cryptophyte and Cyanophyte) isolated from a natural community. PLoS ONE 8, e57139 (2013).
Roach, T., Na, C. S. & Krieger-Liszkay, A. High light-induced hydrogen peroxide production in Chlamydomonas reinhardtii is increased by high CO2 availability. Plant J. 81, 759–766 (2015).
Armbruster, U. et al. Ion antiport accelerates photosynthetic acclimation in fluctuating light environments. Nat. Commun. 5, 5439 (2014).
Yokotani, N. et al. A novel chloroplast protein, CEST induces tolerance to multiple environmental stresses and reduces photooxidative damage in transgenic Arabidopsis. J. Exp. Bot. 62, 557–569 (2011).
Calderon, R. H. et al. A conserved rubredoxin is necessary for photosystem II accumulation in diverse oxygenic photoautotrophs. J. Biol. Chem. 288, 26688–26696 (2013).
Michelet, L. et al. Redox regulation of the Calvin–Benson cycle: something old, something new. Front. Plant Sci. 4, 470 (2013).
Pérez-Pérez, M. E. et al. The deep thioredoxome in Chlamydomonas reinhardtii: new insights into redox regulation. Mol. Plant 10, 1107–1125 (2017).
Scarpeci, T. E., Zanor, M. I. & Valle, E. M. Investigating the role of plant heat shock proteins during oxidative stress. Plant Signal. Behav. 3, 856–857 (2008).
Vignols, F. et al. Redox control of Hsp70–Co-chaperone interaction revealed by expression of a thioredoxin-like Arabidopsis protein. J. Biol. Chem. 278, 4516–4523 (2003).
Jung, H.-S. & Niyogi, K. K. Mutations in Arabidopsis YCF20-like genes affect thermal dissipation of excess absorbed light energy. Planta 231, 923–937 (2010).
Herdean, A. et al. A voltage-dependent chloride channel fine-tunes photosynthesis in plants. Nat. Commun. 7, 11654 (2016).
Gao, X.-H. et al. Methods for analysis of protein glutathionylation and their application to photosynthetic organisms. Mol. Plant 2, 218–235 (2009).
Dixon, D. P. et al. Stress-induced protein S-glutathionylation in Arabidopsis. Plant Physiol. 138, 2233–2244 (2005).
Tibshirani, R., Walther, G. & Hastie, T. Estimating the number of clusters in a data set via the gap statistic. J. R. Stat. Soc. Series B 63, 411–423 (2001).
Martins, M. C. et al. Feedback inhibition of starch degradation in Arabidopsis leaves mediated by trehalose 6-phosphate. Plant Physiol. 163, 1142–1163 (2013).
Mackinder, L. C. M. et al. A repeat protein links Rubisco to form the eukaryotic carbon-concentrating organelle. Proc. Natl Acad. Sci. USA 113, 5958–5963 (2016).
Brueggeman, A. J. et al. Activation of the carbon concentrating mechanism by CO2 deprivation coincides with massive transcriptional restructuring in Chlamydomonas reinhardtii. Plant Cell 24, 1860–1875 (2012).
Fang, W. et al. Transcriptome-wide changes in Chlamydomonas reinhardtii gene expression regulated by carbon dioxide and the CO2-concentrating mechanism regulator CIA5/CCM1. Plant Cell 24, 1876–1893 (2012).
Barber, J. & Andersson, B. Too much of a good thing: light can be bad for photosynthesis. Trends Biochem. Sci. 17, 61–66 (1992).
Tikkanen, M. et al. Thylakoid protein phosphorylation in higher plant chloroplasts optimizes electron transfer under fluctuating light. Plant Physiol. 152, 723–735 (2010).
Allahverdiyeva, Y. et al. Flavodiiron proteins Flv1 and Flv3 enable cyanobacterial growth and photosynthesis under fluctuating light. Proc. Natl Acad. Sci. USA 110, 4111–4116 (2013).
Wagner, H., Jakob, T. & Wilhelm, C. Balancing the energy flow from captured light to biomass under fluctuating light conditions. New Phytol. 169, 95–108 (2006).
Beeby, M. et al. The genomics of disulfide bonding and protein stabilization in thermophiles. PLoS Biol. 3, e309 (2005).
Jorda, J. Y. & O Yeates, Todd Widespread disulfide bonding in proteins from thermophilic Archaea. Archaea 2011, 409156 (2011).
Raanan, H. et al. Towards clarifying what distinguishes cyanobacteria able to resurrect after desiccation from those that cannot: the photosynthetic aspect. Biochim. Biophys. Acta Bioenerg. 1857, 715–722 (2016).
Murik, O. et al. What distinguishes cyanobacteria able to revive after desiccation from those that cannot: the genome aspect. Environ. Microbiol. 19, 535–550 (2016).
De-Bashan, L. E. et al. Chlorella sorokiniana UTEX 2805, a heat and intense, sunlight-tolerant microalga with potential for removing ammonium from wastewater. Bioresour. Technol. 99, 4980–4989 (2008).
Estevez, M. S., Malanga, G. & Puntarulo, S. Iron-dependent oxidative stress in Chlorella vulgaris. Plant Sci. 161, 9–17 (2001).
Badger, M. R. & Price, G. D. The role of carbonic anhydrase in photosynthesis. Annu. Rev. Plant Physiol. Plant Mol. Biol. 45, 369–392 (1994).
Villarejo, A. et al. The induction of the CO2 concentrating mechanism in a starch-less mutant of Chlamydomonas reinhardtii. Physiol. Plant. 98, 798–802 (1996).
Kuchitsu, K., Tsuzuki, M. & Miyachi, S. Changes of starch localization within the chloroplast induced by changes in CO2 concentration during growth of Chlamydomonas reinhardtii: independent regulation of pyrenoid starch and stroma starch. Plant Cell Physiol. 29, 1269–1278 (1988).
Cerveny, J. et al. Ultradian metabolic rhythm in the diazotrophic cyanobacterium Cyanothece sp. ATCC 51142. Proc. Natl Acad. Sci. USA 110, 13210–13215 (2013).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Gotz, S. et al. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 36, 3420–3435 (2008).
Jüppner, J. et al. Dynamics of lipids and metabolites during the cell cycle of Chlamydomonas reinhardtii. Plant J. 92, 331–343 (2017).
Luedemann, A. et al. TagFinder for the quantitative analysis of gas chromatography-mass spectrometry (GC–MS)-based metabolite profiling experiments. Bioinformatics 24, 732–737 (2008).
Kopka, J. et al. GMD@CSB.DB: the Golm metabolome database. Bioinformatics 21, 1635–1638 (2005).
Hummel, J. et al. Ultra performance liquid chromatography and high resolution mass spectrometry for the analysis of plant lipids. Front. Plant Sci. 2, 54 (2011).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Hendriks, J. H. M. et al. ADP-glucose pyrophosphorylase is activated by posttranslational redox-modification in response to light and to sugars in leaves of Arabidopsis and other plant species. Plant Physiol. 133, 838–849 (2003).
Stitt, M., Lilley, R., Gerhardt, R. & Heldt, H. W. in Methods in Enzymology Vol. 174 518–552 (Academic Press, 1989).
Guo, J. et al. Resin-assisted enrichment of thiols as a general strategy for proteomic profiling of cysteine-based reversible modifications. Nat. Protoc. 9, 64–75 (2013).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Kong, F. et al. Interorganelle communication: peroxisomal MALATE DEHYDROGENASE2 connects lipid catabolism to photosynthesis through redox coupling in Chlamydomonas. Plant Cell 30, 1824–1847 (2018).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Stacklies, W. et al. pcaMethods—a bioconductor package providing PCA methods for incomplete data. Bioinformatics 23, 1164–1167 (2007).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Strimmer, K. fdrtool: a versatile R package for estimating local and tail area-based false discovery rates. Bioinformatics 24, 1461–1462 (2008).
Acknowledgements
We thank B. Gontero from BIP-Marseille for constructive discussions regarding the CP12 protein. We thank the Technion Genome Centre (Haifa, Israel) for their support during transcriptome sequencing and analysis. We thank the Human Frontiers Scholarship programme for financial support given to H.T.
Author information
Authors and Affiliations
Contributions
H.T. and M.S. designed the research. H.T., B.S., U.L. and A.E. performed research. H.T., O.M., A.E., U.A., Y.B., J.K., S.A.R. and J.S. analysed data. H.T. and M.S. wrote the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks G. Charles Dismukes, Arthur Grossman the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 NADPH fluorescence in C. ohadii cultures under LL and EIL.
NADPH fluorescence measured at shifts from dark to 100, 1000 and 3000 µmol photons m−2 s−1, in LL (black) and t15EIL (red) treated C. ohadii cells. Similar results were obtained from 6 biologically independent replicates.
Extended Data Fig. 2 Principle Component analyses of nuclear-encoded transcripts, metabolites and lipids following shift to EIL.
Principle Component analyses of log2-normalized nuclear-encoded transcripts (a), metabolites (b) and lipids (c) following shift to EIL, and the corresponding loading analysis of individual traits (d–f). tXEIL indicate the minutes relative to time point zero under EIL, Cont. - continuous EIL, and LL, low light. For a given treatment, each biological replicate is shown separately (n = 3). The dashed line in d separates upper positive quartile of PC1.
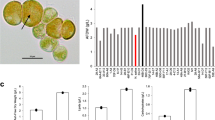
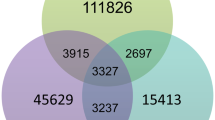
Extended Data Fig. 3 C. ohadii genomic cysteine content analysis.
a, Enriched GO terms (Fisher’s exact test, P value < 0.05, corrected for multiple hypotheses testing by the Benjamini-Hochberg correction procedure) identified in the redox-responsive proteome (n = 4 biologically independent experiments) of t3EIL (red), using a reference (black) of the entire thiol enriched proteome. Values are presented as % of proteins associated with a given GO term from the entire list of proteins in the t3EIL redox-responsive proteome (test) and the entire thiol-enriched proteome (reference). b, Cysteine counts in 4341 shared by the 5 algae presented. c, Average cysteine counts per gene of different KOG categories in C. ohadii (red) and C. reinhardtii (black).
Extended Data Fig. 4 Functional analysis of changes in the transcriptome.
a–c, Gene ontology (GO) bias word clouds (biological process). Word clouds of genes upregulated (n = 3 biologically independent experiments) in t15EIL (a), t120EIL (b) and continuous EIL (c). Font size correlates with significance (see Supplementary Table 4); red terms are depleted and green terms enriched (Fisher’s exact test, P value < 0.05, corrected for multiple hypotheses testing by the Benjamini-Hochberg correction procedure).
Extended Data Fig. 5 Changes of primary metabolites levels in C. ohadii cultures under LL and EIL.
Changes of primary metabolites levels - Heatmap representation of metabolite levels of C. ohadii cells at LL, t120EIL and continuous EIL. Hierarchical clustering was performed on log-transformed, mean-centered and unit variance-scaled data using Pearson’s correlation optimized for sample and metabolite leaf order (see methods). Only significantly changing metabolites were considered (n = 3 biologically independent experiments, one-way ANOVA <0.05). The row Z-score value of each metabolite is plotted in red–blue color scale. GAP statistics indicated 5 metabolite clusters (Supplementary Fig. 2), which are indicated by transparent blue triangles.
Extended Data Fig. 6 Detailed relative metabolites levels in C. ohadii cultures under LL and EIL.
Detailed Log2 normalized values of detected metabolites at LL (blue), t120EIL (orange) and continuous EIL (red). Data are mean ± SD (n = 3 biologically independent experiments, paired two-tailed t-test; one asterisk, P < 0.05; two asterisks, P < 0.01; three asterisks, P < 0.001).
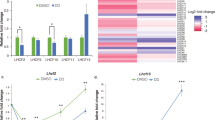
Extended Data Fig. 7 Photorespiratory gene expression in C. ohadii cultures under LL and EIL.
Log2 normalized fold change of expression of photorespiratory genes between t15EIL (yellow), t120EIL (orange) and continuous EIL (red) to LL (n = 3 biologically independent experiments, Cuffdiff, T-statistics, Benjamini-Hochberg corrected P values are presented; one asterisk, P < 0.05; two asterisks, P < 0.01). Expression and corrected P values are further detailed in Supplementary Table 7.
Extended Data Fig. 8 C. ohadii morphology after mid-term exposure to EIL and during growth in continuous EIL.
C. ohadii morphology after mid-term exposure to EIL and during growth in continuous EIL. a, Electron micrographs showing C. ohadii cells grown in TAP (Tris, acetate, phosphate) medium at 35 °C under LL (center), t120EIL (left panel) and continuous EIL (right panel). Bar indicates 500 nm. b, Higher magnifications of C. ohadii continuous EIL treated cells. Arrows indicate altered morphology of pyrenoid starch sheath where in several other cells sheath is absent. Large amounts of starch are also deposited at other sites in the plastid. Similar results were obtained from 3 biologically independent replicates. Bars indicate 2000 and 1000 nm in the upper and lower panels, respectively.
Extended Data Fig. 9 Multi-omics illustration of C. ohadii response to EIL.
Multi-omics illustration of C. ohadii temporal responses to EIL treatments associated with thylakoid morphology. Genes, proteins, metabolites and lipids presented showed significantly higher (red) or lower (blue) levels of expression, redox-response, and accumulation, respectively. Arrows illustrate potential positive (red) and negative (blue), direct (solid line), and indirect (dashed line) effects, according to the model suggested in the discussion.
Extended Data Fig. 10 Desert BSC diurnal cycle.
Schematic view of desert BSC diurnal cycle. Main water supply is by dew formation during the night, followed by morning dehydration. Through most of the day, crusts are dry and exposed to ~2000 µmol photons m−2 s−1.
Supplementary information
Supplementary Information
Supplementary text, Figs. 1–11 and Tables 1, 3 and 5–9.
Supplementary Tables
Supplementary Tables 1–10. Legends are provided for each table within each tab in the collated file.
Supplementary Data 1
Comparative cysteine count in C. ohadii proteins with homologues in other algae. Gene names and descriptions are detailed in accession no. PRJNA573576 (NCBI).
Supplementary Data 2
NADP-MDH sequence alignment.
Supplementary Data 3
Cytosolic aldolase sequence alignment.
Supplementary Data 4
NADP-GAPDH sequence alignment.
Supplementary Data 5
NADP-ME sequence alignment.
Supplementary Data 6
LPAT sequence alignment.
Supplementary Data 7
DGAT2 sequence alignment.
Supplementary Data 8
CP47 sequence alignment.
Supplementary Data 9
Cytochrome f sequence alignment.
Rights and permissions
About this article
Cite this article
Treves, H., Siemiatkowska, B., Luzarowska, U. et al. Multi-omics reveals mechanisms of total resistance to extreme illumination of a desert alga. Nat. Plants 6, 1031–1043 (2020). https://doi.org/10.1038/s41477-020-0729-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-0729-9
This article is cited by
-
Increased genetic diversity loss and genetic differentiation in a model marine diatom adapted to ocean warming compared to high CO2
The ISME Journal (2022)
-
Carbon flux through photosynthesis and central carbon metabolism show distinct patterns between algae, C3 and C4 plants
Nature Plants (2021)
-
Cryo-EM photosystem I structure reveals adaptation mechanisms to extreme high light in Chlorella ohadii
Nature Plants (2021)
-
Juggling Lightning: How Chlorella ohadii handles extreme energy inputs without damage
Photosynthesis Research (2021)