Abstract
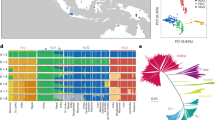
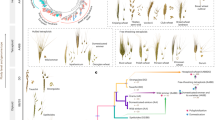
Rice (Oryza sativa) is one of the world’s most important food crops, and is comprised largely of japonica and indica subspecies. Here, we reconstruct the history of rice dispersal in Asia using whole-genome sequences of more than 1,400 landraces, coupled with geographic, environmental, archaeobotanical and paleoclimate data. Originating around 9,000 yr ago in the Yangtze Valley, rice diversified into temperate and tropical japonica rice during a global cooling event about 4,200 yr ago. Soon after, tropical japonica rice reached Southeast Asia, where it rapidly diversified, starting about 2,500 yr bp. The history of indica rice dispersal appears more complicated, moving into China around 2,000 yr bp. We also identify extrinsic factors that influence genome diversity, with temperature being a leading abiotic factor. Reconstructing the dispersal history of rice and its climatic correlates may help identify genetic adaptations associated with the spread of a key domesticated species.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw FASTQ reads for 178 accessions whose genomes were resequenced for this study have been deposited in the SRA under Bioproject accession numbers PRJNA422249 and PRJNA557122. Sources for all downloaded data are referred to in the Supplementary Information.
Code availability
Code repositories are available at: https://github.com/bocinsky/gutaker2020, https://github.com/grafau/discretize, https://github.com/grafau/NextGatkSNPs and https://github.com/em-bellis/riceTravelTime.
References
Purugganan, M. D. & Fuller, D. Q. The nature of selection during plant domestication. Nature 457, 843–848 (2009).
Meyer, R. S. & Purugganan, M. D. Evolution of crop species: genetics of domestication and diversification. Nat. Rev. Genet. 14, 840–852 (2013).
Wang, W. et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557, 43–49 (2018).
Glaszmann, J. C. Isozymes and classification of Asian rice varieties. Theor. Appl. Genet. 74, 21–30 (1987).
Fuller, D. Q. et al. The contribution of rice agriculture and livestock pastoralism to prehistoric methane levels: an archaeological assessment. Holocene 21, 743–759 (2011).
Fuller, D. Q. & Qin, L. Water management and labour in the origins and dispersal of Asian rice. World Archaeol. 41, 88–111 (2009).
Fuller, D. Q. et al. The domestication process and domestication rate in rice: spikelet bases from the Lower Yangtze. Science 323, 1607–1610 (2009).
Allaby, R. G., Stevens, C., Lucas, L., Maeda, O. & Fuller, D. Q. Geographic mosaics and changing rates of cereal domestication. Philos. Trans. R. Soc. Lond. B 372, 20160429 (2017).
Silva, F. et al. A tale of two rice varieties: modelling the prehistoric dispersals of japonica and proto-indica rices. Holocene 28, 1745–1758 (2018).
Fuller, D. Q. Pathways to Asian civilizations: tracing the origins and spread of rice and rice cultures. Rice 4, 78–92 (2011).
Huang, X. et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 490, 497–501 (2012).
Choi, J. Y. & Purugganan, M. D. Multiple origin but single domestication led to Oryza sativa. G3 8, 797–803 (2018).
Choi, J. Y. et al. The rice paradox: multiple origins but single domestication in Asian rice. Mol. Biol. Evol. 34, 969–979 (2017).
Fuller, D. Q., Castillo, C. C. & Murphy, C. in The Routledge Handbook of Archaeology and Globalization (ed. Hodos, T.) 711–729 (Routledge, 2016).
Silva, F. et al. Modelling the geographical origin of rice cultivation in Asia using the rice archaeological database. PLoS ONE 10, e0137024 (2015).
Li, J.-Y., Wang, J. & Zeigler, R. S. The 3,000 rice genomes project: new opportunities and challenges for future rice research. Gigascience 3, 2047–217X–3–8 (2014).
Excoffier, L., Smouse, P. E. & Quattro, J. M. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131, 479–491 (1992).
Rousseeuw, P. J. Silhouettes: a graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 20, 53–65 (1987).
Petkova, D., Novembre, J. & Stephens, M. Visualizing spatial population structure with estimated effective migration surfaces. Nat. Genet. 48, 94–100 (2016).
Peter, B. M., Petkova, D. & Novembre, J. Genetic landscapes reveal how human genetic diversity aligns with geography. Mol. Biol. Evol. 37, 943–951 (2020).
Slayton, E. R. Seascape Corridors: Modeling Routes to Connect Communities Across the Caribbean Sea. (Sidestone Press, 2018).
Peres-Neto, P. R., Legendre, P., Dray, S. & Borcard, D. Variation partitioning of species data matrices: estimation and comparison of fractions. Ecology 87, 2614–2625 (2006).
Lasky, J. R. et al. Genome–environment associations in sorghum landraces predict adaptive traits. Sci. Adv. 1, e1400218 (2015).
Lasky, J. R. et al. Characterizing genomic variation of Arabidopsis thaliana: the roles of geography and climate. Mol. Ecol. 21, 5512–5529 (2012).
Haefele, S. M., Nelson, A. & Hijmans, R. J. Soil quality and constraints in global rice production. Geoderma 235-236, 250–259 (2014).
Kaufmann, L & Rousseeuw, P. J. in Reports of the Faculty of Technical Mathematics and Informatics Vol. 87 (Delft University of Technology, 1987).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
An, C.-B., Tang, L., Barton, L. & Chen, F.-H. Climate change and cultural response around 4000 cal yr B.P. in the western part of Chinese Loess Plateau. Quat. Res 63, 347–352 (2005).
Walker, M. J. C. et al. Formal subdivision of the Holocene series/epoch: a discussion paper by a working group of INTIMATE (integration of ice-core, marine and terrestrial records) and the subcommission on Quaternary stratigraphy (International Commission on Stratigraphy). J. Quat. Sci 27, 649–659 (2012).
Lanehart, R. E. et al. Dietary adaptation during the Longshan period in China: stable isotope analyses at Liangchengzhen (southeastern Shandong). J. Archaeol. Sci. 38, 2171–2181 (2011).
Guedes, J. D., Jiang, M., He, K., Wu, X. & Jiang, Z. Site of Baodun yields earliest evidence for the spread of rice and foxtail millet agriculture to south-west China. Antiquity 87, 758–771 (2013).
Guedes, J. D. & Butler, E. E. Modeling constraints on the spread of agriculture to Southwest China with thermal niche models. Quat. Int. 349, 29–41 (2014).
Dal Martello, R. et al. Early agriculture at the crossroads of China and Southeast Asia: archaeobotanical evidence and radiocarbon dates from Baiyangcun, Yunnan. J. Archaeol. Sci. Rep. 20, 711–721 (2018).
Fuller, D. Q., Weisskopf, A. R. & Castillo, C. Pathways of rice diversification across Asia. Archaeol. Int. 19, 84–96 (2016).
d’Alpoim Guedes, J., Jin, G. & Bocinsky, R. K. The impact of climate on the spread of rice to north-eastern China: a new look at the data from Shandong province. PLoS ONE 10, e0130430 (2015).
Crawford, G. W. & Lee, G.-A. Agricultural origins in the Korean Peninsula. Antiquity 77, 87–95 (2003).
Ahn, S.-M. The emergence of rice agriculture in Korea: archaeobotanical perspectives. Archaeol. Anthropol. Sci. 2, 89–98 (2010).
Yang, X. et al. New radiocarbon evidence on early rice consumption and farming in South China. Holocene 27, 1045–1051 (2017).
Marcott, S. A., Shakun, J. D., Clark, P. U. & Mix, A. C. A reconstruction of regional and global temperature for the past 11,300 years. Science 339, 1198–1201 (2013).
d’Alpoim Guedes, J. & Bocinsky, R. K. Climate change stimulated agricultural innovation and exchange across Asia. Sci. Adv. 4, eaar4491 (2018).
Castillo, C. C., Fuller, D. Q., Piper, P. J., Bellwood, P. & Oxenham, M. Hunter-gatherer specialization in the late Neolithic of southern Vietnam—the case of Rach Nui. Quat. Int. 489, 63–79 (2018).
Higham, C. F. W. Debating a great site: Ban Non Wat and the wider prehistory of Southeast Asia. Antiquity 89, 1211–1220 (2015).
Higham, C. The Bronze Age of Southeast Asia (Cambridge Univ. Press, 1996).
Castillo, C. C. et al. Social responses to climate change in Iron Age north-east Thailand: new archaeobotanical evidence. Antiquity 92, 1274–1291 (2018).
McColl, H. et al. The prehistoric peopling of Southeast Asia. Science 361, 88–92 (2018).
Lipson, M. et al. Ancient genomes document multiple waves of migration in Southeast Asian prehistory. Science 361, 92–95 (2018).
Calò, A. The Distribution of Bronze Drums in Early Southeast Asia: Trade Routes and Cultural Spheres. (Archaeopress, 2009).
Castillo, C. C., Bellina, B. & Fuller, D. Q. Rice, beans and trade crops on the early maritime Silk Route in Southeast Asia. Antiquity 90, 1255–1269 (2016).
Hung, H.-C. et al. Ancient jades map 3,000 years of prehistoric exchange in Southeast Asia. Proc. Natl Acad. Sci. USA 104, 19745–19750 (2007).
Takamiya, H., Hudson, M. J., Yonenobu, H., Kurozumi, T. & Toizumi, T. An extraordinary case in human history: prehistoric hunter-gatherer adaptation to the islands of the Central Ryukyus (Amami and Okinawa archipelagos), Japan. Holocene 26, 408–422 (2016).
Zürcher, E. in The Buddhist conquest of China (Brill, 1972).
Deng, Z. et al. From early domesticated rice of the middle Yangtze basin to millet, rice and wheat agriculture: archaeobotanical macro-remains from Baligang, Nanyang Basin, central China (6700–500 BC). PLoS ONE 10, e0139885 (2015).
Verdugo, M. P. et al. Ancient cattle genomics, origins, and rapid turnover in the Fertile Crescent. Science 365, 173–176 (2019).
Gibbons, A. How the Akkadian empire was hung out to dry. Science 261, 985 (1993).
Wang, J. et al. The abrupt climate change near 4,400 yr BP on the cultural transition in Yuchisi, China and its global linkage. Sci. Rep. 6, 27723 (2016).
Harlan, J. R. Our vanishing genetic resources. Science 188, 617–621 (1975).
Villa, T. C. C., Maxted, N., Scholten, M. & Ford-Lloyd, B. Defining and identifying crop landraces. Plant Genet. Resour. 3, 373–384 (2005).
McLaren, C. G., Bruskiewich, R. M., Portugal, A. M. & Cosico, A. B. The International Rice Information System. A platform for meta-analysis of rice crop data. Plant Physiol. 139, 637–642 (2005).
Huke, R. E. & Huke, E. H. Rice Area by Type of Culture: South, Southeast, and East Asia: A Revised and Updated Data Base (International Rice Research Institute, 1997).
Maclean, J., Hardy, B. & Hettel, G. Rice Almanac: Source Book for One of the Most Important Economic Activities on Earth 4th edn (International Rice Research Institute, 2013).
Laborte, A. G. et al. RiceAtlas, a spatial database of global rice calendars and production. Sci. Data 4, 170074 (2017).
Kim, H. et al. Population dynamics among six major groups of the Oryza rufipogon species complex, wild relative of cultivated Asian rice. Rice 9, 56 (2016).
Hirano, H. Y., Eiguchi, M. & Sano, Y. A single base change altered the regulation of the Waxy gene at the posttranscriptional level during the domestication of rice. Mol. Biol. Evol. 15, 978–987 (1998).
Hammarström, H., Forkel, R. & Haspelmath, M. Glottolog 4.0 (2019); https://doi.org/10.5281/zenodo.3260726
Hijmans, R. J. & van Etten, J. raster: Geographic data analysis and modeling v.2 (2014).
Karger, D. N. et al. Climatologies at high resolution for the earth’s land surface areas. Sci. Data 4, 170122 (2017).
Zomer, R. J. et al. Trees and Water: Smallholder Agroforestry on Irrigated Lands in Northern India. (International Water Management Institute, 2007).
Zomer, R. J., Trabucco, A., Bossio, D. A. & Verchot, L. V. Climate change mitigation: A spatial analysis of global land suitability for clean development mechanism afforestation and reforestation. Agric. Ecosyst. Environ. 126, 67–80 (2008).
Kummu, M., de Moel, H., Ward, P. J. & Varis, O. How close do we live to water? A global analysis of population distance to freshwater bodies. PLoS ONE 6, e20578 (2011).
Hijmans, R. J., Cameron, S. E., Parra, J. L., Jones, P. G. & Jarvis, A. Very high resolution interpolated climate surfaces for global land areas. Int. J. Climatol. 25, 1965–1978 (2005).
Global Soil Data Task Group. Global gridded surfaces of selected soil characteristics (IGBP-DIS) (2002); https://doi.org/10.3334/ORNLDAAC/569
Dunne, K. A. & Willmott, C. J. Global distribution of plant-extractable water capacity of soil. Int. J. Climatol. 16, 841–859 (1996).
Fan, Y., Li, H. & Miguez-Macho, G. Global patterns of groundwater table depth. Science 339, 940–943 (2013).
Di Tommaso, P. et al. Nextflow enables reproducible computational workflows. Nat. Biotechnol. 35, 316–319 (2017).
Du, H. et al. Sequencing and de novo assembly of a near complete indica rice genome. Nat. Commun. 8, 15324 (2017).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at https://arxiv.org/abs/1303.3997 (2013).
Van der Auwera, G. A. et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr. Protoc. Bioinformatics 43, 11.10.1–11.10.33 (2013).
Li, H. Tabix: fast retrieval of sequence features from generic TAB-delimited files. Bioinformatics 27, 718–719 (2011).
McCouch, S. R. et al. Open access resources for genome-wide association mapping in rice. Nat. Commun. 7, 10532 (2016).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
R Core Team. R: A Language and Environment for Statistical Computing http://www.R-project.org/ (R Foundation for Statistical Computing, 2013).
Oksanen, J. Vegan: An Introduction to Ordination https://cran.r-project.org/web/packages/vegan/vignettes/intro-vegan.pdf (2015).
Browning, S. R. & Browning, B. L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007).
van Etten, J. R package gdistance: distances and routes on geographical grids. J. Stat. Softw. 76, v76i13 (2017).
Tobler, W. Three Presentations on Geographical Analysis and Modeling: Non-isotropic Geographic Modeling; Speculations on the Geometry of Geography; And Global Spatial Analysis (National Center for Geographic Information and Analysis, 1993).
White, D. A. & Surface-Evans, S. L. Least Cost Analysis of Social Landscapes: Archaeological Case Studies (Univ. Utah Press, 2012).
Irwin, G., Bickler, S. & Quirke, P. Voyaging by canoe and computer: experiments in the settlement of the Pacific Ocean. Antiquity 64, 34–50 (1990).
Clarke, R. T., Rothery, P. & Raybould, A. F. Confidence limits for regression relationships between distance matrices: estimating gene flow with distance. J. Agric. Biol. Environ. Stat. 7, 361 (2002).
Akaike, H. A new look at the statistical model identification. IEEE Trans. Automat. Contr. 19, 716–723 (1974).
Shirk, A. J., Landguth, E. L. & Cushman, S. A. A comparison of regression methods for model selection in individual-based landscape genetic analysis. Mol. Ecol. Resour. 18, 55–67 (2018).
Peterman, W. E. ResistanceGA: An R package for the optimization of resistance surfaces using genetic algorithms. Methods Ecol. Evol. 9, 1638–1647 (2018).
Peterman, W. E., Connette, G. M., Semlitsch, R. D. & Eggert, L. S. Ecological resistance surfaces predict fine-scale genetic differentiation in a terrestrial woodland salamander. Mol. Ecol. 23, 2402–2413 (2014).
Bauman, D., Drouet, T., Fortin, M.-J. & Dray, S. Optimizing the choice of a spatial weighting matrix in eigenvector-based methods. Ecology 99, 2159–2166 (2018).
Dray, S. et al. adespatial: Multivariate Multiscale Spatial Analysis. R package v.0 (2019).
Mardia, K. V. Some properties of classical multi-dimesional scaling. Comm. Stat. Theory Methods 7, 1233–1241 (1978).
Schubert, E. & Rousseeuw, P. J. in Similarity Search and Applications. SISAP 2019. Lecture Notes in Computer Science Vol 11807 (eds Amato, G. et al.) 171–187 (Springer, 2019).
Kahle, D. & Wickham, H. ggmap: spatial visualization with ggplot2. R J. 5, 144–161 (2013).
Leppälä, K., Nielsen, S. V. & Mailund, T. admixturegraph: an R package for admixture graph manipulation and fitting. Bioinformatics 33, 1738–1740 (2017).
Terhorst, J., Kamm, J. A. & Song, Y. S. Robust and scalable inference of population history from hundreds of unphased whole genomes. Nat. Genet. 49, 303–309 (2017).
Choi, J. Y. & Purugganan, M. D. Evolutionary epigenomics of retrotransposon-mediated methylation spreading in rice. Mol. Biol. Evol. 35, 365–382 (2018).
Gaut, B. S., Morton, B. R., McCaig, B. C. & Clegg, M. T. Substitution rate comparisons between grasses and palms: synonymous rate differences at the nuclear gene Adh parallel rate differences at the plastid gene rbcL. Proc. Natl. Acad Sci. USA 93, 10274–10279 (1996).
Global Historical Climatology Network-DAILY (GHCN-Daily) version 3 (NOAA National Climatic Data Center, 2012); https://www.ncdc.noaa.gov/data-access/land-based-station-data/land-based-datasets/global-historical-climatology-network-monthly-version-3
Edwards, M. Data Announcement 88-MGG-02: Digital Relief of the Surface of the Earth (National Oceanic and Atmospheric Administration and National Geophysical Data Center, 1988).
Acknowledgements
We thank our colleagues for helpful discussions on this project. This work was supported in part by Zegar Family Foundation grant A16-0051-004 and US National Science Foundation Plant Genome Research Program grant IOS-1546218 to M.D.P., Portugal Fundação para a Ciência e a Tecnologia grant EXPL/BIA-BIC/0947/2012 to S.N., SFRH/BD/68835/2010 to I.S.P. and UID/Multi/04551/2013 to M.M.O., Gordon and Betty Moore Foundation and Life Sciences Research Foundation grant GBMF2550.06 to S.C.G., US National Science Foundation grant PRFB 1711950 to E.S.B., Natural Environment Research Council UK grant NE/N010957/1 to C.C.C. and D.Q.F., US National Science Foundation grant BCS-1632207 to J.A.d.G. and United States Department of Agriculture and National Institute of Food and Agriculture grant 2019-67009-29006 to J.R.L.
Author information
Authors and Affiliations
Contributions
R.M.G. and M.D.P. conceived and designed the study with input from J.R.L. and S.C.G. J.Y.C., I.S.P. and O.W. generated sequencing data. M.D.P., S.N. and M.M.O. supervised laboratory work. R.M.G. assembled and processed the sequencing data. S.C.G. and E.S.B. assembled and processed the environmental data with input from J.R.L. J.R.L. led the spatial analyses with input from R.M.G. E.S.B. and E.R.S. carried out travel-time analyses with input from J.R.L. J.R.L. carried out R.D.A. analyses. R.M.G. carried out population-structure, admixture-graph and coalescence analyses. R.K.B. and J.A.d.G. conducted thermal-niche modelling. D.Q.F., C.C.C. and J.A.d.G. provided archaeological context. M.D.P., R.M.G. and J.R.L. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Laura Botigué and Angé́lica Cibrián-Jaramillo and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–24.
Supplementary Video 1
Spatio-temporal distribution of rice thermal niche. Video illustrating the probability of rice being in niche based on the minimum and maximum growing degree days requirement for tropical japonica landraces. Map of Asia with plotted thermal niche probabilities, colour-coded as indicated below the map. White lines denote the border under which niche probability drops below 75%. Below the map is a plot of probabilities averaged across spatial scale. The thick black line represents mean, thin lines show interquartile range, and grey shaded area represents 25% to 75% probability of being in the thermal niche (n = 477,708 cells). The thin black lines are the mean probabilities of being in the thermal niche across the study area when modelled using the 1σ uncertainty intervals as provided by the Northern Hemisphere temperature reconstruction (n = 73 datasets)40. Moving vertical red line indicates time before present.
Supplementary Table 1
Supplementary Tables 1–3.
Source data
Source Data Fig. 1
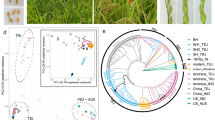
Geographic coordinates, migration data, statistical data and Canonical coordinates.
Source Data Fig. 2
Dimension (MDS) coordinates, country codes and graph dot format.
Source Data Fig. 3
Statistical data points, geographic coordinates, graph trajectories, statistical data and raster data.
Source Data Fig. 4
Geographic coordinates and statistical data.
Source Data Fig. 5
Geographic coordinates and statistical data.
Rights and permissions
About this article
Cite this article
Gutaker, R.M., Groen, S.C., Bellis, E.S. et al. Genomic history and ecology of the geographic spread of rice. Nat. Plants 6, 492–502 (2020). https://doi.org/10.1038/s41477-020-0659-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-0659-6
This article is cited by
-
Biofortified Rice Provides Rich Sakuranetin in Endosperm
Rice (2024)
-
Exploring Genetic Diversity within aus Rice Germplasm: Insights into the Variations in Agro-morphological Traits
Rice (2024)
-
Are cereal grasses a single genetic system?
Nature Plants (2024)
-
Promotive effect of mechanochemically crushed straw on rice growth by improving soil properties and modulating bacterial communities
Plant Growth Regulation (2024)
-
Human influence on the distribution of cacao: insights from remote sensing and biogeography
Biodiversity and Conservation (2024)