Abstract
Rubisco sustains the biosphere through the fixation of CO2 into biomass. In plants and cyanobacteria, form I Rubisco is structurally comprised of large and small subunits, whereas all other Rubisco forms lack small subunits. The rise of the form I complex through the innovation of small subunits represents a key, yet poorly understood, transition in Rubisco’s evolution. Through metagenomic analyses, we discovered a previously uncharacterized clade sister to form I Rubisco that evolved without small subunits. This clade diverged before the evolution of cyanobacteria and the origin of the small subunit; thus, it provides a unique reference point to advance our understanding of form I Rubisco evolution. Structural and kinetic data presented here reveal how a proto-form I Rubisco assembled and functioned without the structural stability imparted from small subunits. Our findings provide insight into a key evolutionary transition of the most abundant enzyme on Earth and the predominant entry point for nearly all global organic carbon.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Form I' RbcL amino acid sequences are included as Supplementary Data 1. Sequences used to generate Fig. 1a were uploaded to figshare (https://doi.org/10.6084/m9.figshare.9980630) along with the associated phylogenetic tree. Representative MAG genbank scaffolds are included as Supplementary Data 2. Site-directed mutagenesis primers and synthesized candidate form I' rbcL genes are included as Supplementary Data 3. The structural coordinates of 2CABP-bound P. breve Rubisco have been deposited in the PDB under the accession ID 6URA. The crystal structure of Syn6301 Rubisco can be found on the PDB under the accession ID 1RBL. Publicly available databases used in this study include: PDB (https://www.rcsb.org/), pfam (https://pfam.xfam.org/), TIGRfams (www.tigrfams.jcvi.org) and KEGG database (https://www.genome.jp/).Two Chloroflexi genomes identified in this study are available at: https://ggkbase.berkeley.edu/Chloroflexi_Rubisco_PatrickShih/organisms.
References
Nisbet, E. G. et al. The age of Rubisco: the evolution of oxygenic photosynthesis. Geobiology 5, 311–335 (2007).
Tabita, F. R. et al. Function, structure, and evolution of the RubisCO-like proteins and their RubisCO homologs. Microbiol. Mol. Biol. Rev. 71, 576–599 (2007).
Tabita, F. R., Satagopan, S., Hanson, T. E., Kreel, N. E. & Scott, S. S. Distinct form I, II, III, and IV Rubisco proteins from the three kingdoms of life provide clues about Rubisco evolution and structure/function relationships. J. Exp. Bot. 59, 1515–1524 (2007).
Andrews, T. J. Catalysis by cyanobacterial ribulose-bisphosphate carboxylase large subunits in the complete absence of small subunits. J. Biol. Chem. 263, 12213–12219 (1988).
Morell, M. K., Wilkin, J. M., Kane, H. J. & Andrews, T. J. Side reactions catalyzed by ribulose-bisphosphate carboxylase in the presence and absence of small subunits. J. Biol. Chem. 272, 5445–5451 (1997).
Spreitzer, R. J. Role of the small subunit in ribulose-1,5-bisphosphate carboxylase/oxygenase. Arch. Biochem. Biophys. 414, 141–149 (2003).
Joshi, J., Mueller-Cajar, O., Tsai, Y.-C. C., Hartl, F. U. & Hayer-Hartl, M. Role of small subunit in mediating assembly of red-type form I rubisco. J. Biol. Chem. 290, 1066–1074 (2015).
Liu, C. et al. Coupled chaperone action in folding and assembly of hexadecameric Rubisco. Nature 463, 197–202 (2010).
Grabsztunowicz, M., Górski, Z., Luciński, R. & Jackowski, G. A reversible decrease in ribulose 1,5-bisphosphate carboxylase/oxygenase carboxylation activity caused by the aggregation of the enzyme’s large subunit is triggered in response to the exposure of moderate irradiance-grown plants to low irradiance. Physiol. Plant. 154, 591–608 (2015).
Kusian, B. & Bowien, B. Organization and regulation of cbb CO2 assimilation genes in autotrophic bacteria. FEMS Microbiol. Rev. 21, 135–155 (1997).
Tabita, F. R. Microbial ribulose 1,5-bisphosphate carboxylase/oxygenase: a different perspective. Photosynth. Res. 60, 1–28 (1999).
Whitney, S. M. & Andrews, T. J. The gene for the ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) small subunit relocated to the plastid genome of tobacco directs the synthesis of small subunits that assemble into Rubisco. Plant Cell 13, 193–205 (2001).
Bryant, D. A. & Liu, Z. in Advances in Botanical Research (ed. Beatty, J. T.) 99–150 (Academic Press, 2013).
Shih, P. M., Ward, L. M. & Fischer, W. W. Evolution of the 3-hydroxypropionate bicycle and recent transfer of anoxygenic photosynthesis into the Chloroflexi. Proc. Natl Acad. Sci. USA 114, 10749–10754 (2017).
Ward, L. M., Hemp, J., Shih, P. M., McGlynn, S. E. & Fischer, W. W. Evolution of phototrophy in the Chloroflexi phylum driven by horizontal gene transfer. Front. Microbiol. 9, 260 (2018).
Fischer, W. W., Hemp, J. & Johnson, J. E. Evolution of oxygenic photosynthesis. Annu. Rev. Earth Planet. Sci. 44, 647–683 (2016).
Roy, H. Rubisco assembly: a model system for studying the mechanism of chaperonin action. Plant Cell 1, 1035–1042 (1989).
Hayer-Hartl, M. From chaperonins to Rubisco assembly and metabolic repair. Protein Sci. 26, 2324–2333 (2017).
Aigner, H. et al. Plant RuBisCo assembly in E. coli with five chloroplast chaperones including BSD2. Science 358, 1272–1278 (2017).
Wilson, R. H. & Hayer-Hartl, M. Complex chaperone dependence of Rubisco biogenesis. Biochemistry 57, 3210–3216 (2018).
Saschenbrecker, S. et al. Structure and function of RbcX, an assembly chaperone for hexadecameric Rubisco. Cell 129, 1189–1200 (2007).
Gunn, L. H., Valegård, K. & Andersson, I. A unique structural domain in Methanococcoides burtonii ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) acts as a small subunit mimic. J. Biol. Chem. 292, 6838–6850 (2017).
Goloubinoff, P., Christeller, J. T., Gatenby, A. A. & Lorimer, G. H. Reconstitution of active dimeric ribulose bisphosphate carboxylase from an unfolded state depends on two chaperonin proteins and Mg-ATP. Nature 342, 884–889 (1989).
Parry, M. A. J., Keys, A. J. & Gutteridge, S. Variation in the specificity factor of C3 higher plant Rubiscos determined by the total consumption of ribulose-P2. J. Exp. Bot. 40, 317–320 (1989).
Tcherkez, G. G. B., Farquhar, G. D. & Andrews, T. J. Despite slow catalysis and confused substrate specificity, all ribulose bisphosphate carboxylases may be nearly perfectly optimized. Proc. Natl Acad. Sci. USA 103, 7246–7251 (2006).
Flamholz, A. I. et al. Revisiting trade-offs between Rubisco kinetic parameters. Biochemistry 58, 3365–3376 (2019).
Yamada, T. & Sekiguchi, Y. Cultivation of uncultured Chloroflexi subphyla: significance and ecophysiology of formerly uncultured Chloroflexi ‘subphylum i' with natural and biotechnological relevance. Microbes Environ. 24, 205–216 (2009).
Hemp, J., Ward, L. M., Pace, L. A. & Fischer, W. W. Draft genome sequence of Ornatilinea apprima P3M-1, an anaerobic member of the Chloroflexi class Anaerolineae. Genome Announc. 3, e01353-15 (2015).
Ward, L. M., Hemp, J., Pace, L. A. & Fischer, W. W. Draft genome sequence of Leptolinea tardivitalis YMTK-2, a mesophilic anaerobe from the Chloroflexi class Anaerolineae. Genome Announc. 3, e01356-15 (2015).
Alonso, H., Blayney, M. J., Beck, J. L. & Whitney, S. M. Substrate-induced assembly of Methanococcoides burtonii d-ribulose-1,5-bisphosphate carboxylase/oxygenase dimers into decamers. J. Biol. Chem. 284, 33876–33882 (2009).
Knott, G. J. et al. Structural basis for AcrVA4 inhibition of specific CRISPR-Cas12a. eLife 8, e49110 (2019).
Duff, A. P., Andrews, T. J. & Curmi, P. M. The transition between the open and closed states of Rubisco is triggered by the inter-phosphate distance of the bound bisphosphate. J. Mol. Biol. 298, 903–916 (2000).
Newman, J., Branden, C. I. & Jones, T. A. Structure determination and refinement of ribulose 1,5-bisphosphate carboxylase/oxygenase from Synechococcus PCC6301. Acta Crystallogr. D. Biol. Crystallogr. 49, 548–560 (1993).
Lu, Z., Zhao, Z. & Fu, B. Efficient protein alignment algorithm for protein search. BMC Bioinf. 11, S34 (2010).
Cleland, W. W., Andrews, T. J., Gutteridge, S., Hartman, F. C. & Lorimer, G. H. Mechanism of Rubisco: the carbamate as general base. Chem. Rev. 98, 549–562 (1998).
Andersson, I. & Backlund, A. Structure and function of Rubisco. Plant Physiol. Biochem. 46, 275–291 (2008).
van Lun, M., van der Spoel, D. & Andersson, I. Subunit interface dynamics in hexadecameric Rubisco. J. Mol. Biol. 411, 1083–1098 (2011).
Schneider, G. et al. Comparison of the crystal structures of L2 and L8S8 Rubisco suggests a functional role for the small subunit. EMBO J. 9, 2045–2050 (1990).
Huynh, K. & Partch, C. L. Analysis of protein stability and ligand interactions by thermal shift assay. Curr. Protoc. Protein Sci. 79, 28.9.1–28.9.14 (2015).
Greene, D. N., Whitney, S. M. & Matsumura, I. Artificially evolved Synechococcus PCC6301 Rubisco variants exhibit improvements in folding and catalytic efficiency. Biochem. J. 404, 517–524 (2007).
DePristo, M. A., Weinreich, D. M. & Hartl, D. L. Missense meanderings in sequence space: a biophysical view of protein evolution. Nat. Rev. Genet. 6, 678–687 (2005).
Tokuriki, N., Stricher, F., Serrano, L. & Tawfik, D. S. How protein stability and new functions trade off. PLoS Comput. Biol. 4, e1000002 (2008).
Tokuriki, N. & Tawfik, D. S. Protein dynamism and evolvability. Science 324, 203–207 (2009).
Erb, T. J. & Zarzycki, J. A short history of RubisCO: the rise and fall (?) of Nature’s predominant CO2 fixing enzyme. Curr. Opin. Biotechnol. 49, 100–107 (2018).
Badger, M. R., Hanson, D. & Dean Price, G. Evolution and diversity of CO2 concentrating mechanisms in cyanobacteria. Funct. Plant Biol. 29, 161–173 (2002).
Studer, R. A., Christin, P.-A., Williams, M. A. & Orengo, C. A. Stability–activity tradeoffs constrain the adaptive evolution of RubisCO. Proc. Natl Acad. Sci. USA 111, 2223–2228 (2014).
Zhou, Y. & Whitney, S. Directed evolution of an improved Rubisco; in vitro analyses to decipher fact from fiction. Int. J. Mol. Sci. 20, 5019 (2019).
Wilson, R. H., Alonso, H. & Whitney, S. M. Evolving Methanococcoides burtonii archaeal Rubisco for improved photosynthesis and plant growth. Sci. Rep. 6, 22284 (2016).
Frey, S. & Görlich, D. A new set of highly efficient, tag-cleaving proteases for purifying recombinant proteins. J. Chromatogr. A 1337, 95–105 (2014).
Kane, H. J., Wilkin, J. M., Portis, A. R. & John Andrews, T. Potent inhibition of ribulose-bisphosphate carboxylase by an oxidized impurity in ribulose-1,5-bisphosphate. Plant Physiol. 117, 1059–1069 (1998).
Pierce, J., Tolbert, N. E. & Barker, R. Interaction of ribulosebisphosphate carboxylase/oxygenase with transition-state analogues. Biochemistry 19, 934–942 (1980).
Pereira, J. H., McAndrew, R. P., Tomaleri, G. P. & Adams, P. D. Berkeley Screen: a set of 96 solutions for general macromolecular crystallization. J. Appl. Crystallogr. 50, 1352–1358 (2017).
Winter, G., Lobley, C. M. C. & Prince, S. M. Decision making in xia2. Acta Crystallogr. D. Biol. Crystallogr. 69, 1260–1273 (2013).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Dyer, K. N. et al. High-throughput SAXS for the characterization of biomolecules in solution: a practical approach. Methods Mol. Biol. 1091, 245–258 (2014).
Hura, G. L. et al. Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS). Nat. Methods 6, 606–612 (2009).
Rambo, R. P. & Tainer, J. A. Accurate assessment of mass, models and resolution by small-angle scattering. Nature 496, 477–481 (2013).
Sali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Schneidman-Duhovny, D., Hammel, M. & Sali, A. FoXS: a web server for rapid computation and fitting of SAXS profiles. Nucleic Acids Res. 38, W540–W544 (2010).
Schneidman-Duhovny, D., Hammel, M., Tainer, J. A. & Sali, A. Accurate SAXS profile computation and its assessment by contrast variation experiments. Biophys. J. 105, 962–974 (2013).
Prins, A. et al. Rubisco catalytic properties of wild and domesticated relatives provide scope for improving wheat photosynthesis. J. Exp. Bot. 67, 1827–1838 (2016).
Sharwood, R. E., Ghannoum, O. & Whitney, S. M. Prospects for improving CO2 fixation in C3-crops through understanding C4-Rubisco biogenesis and catalytic diversity. Curr. Opin. Plant Biol. 31, 135–142 (2016).
Pei, J., Kim, B.-H. & Grishin, N. V. PROMALS3D: a tool for multiple protein sequence and structure alignments. Nucleic Acids Res. 36, 2295–2300 (2008).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 20, 1160–1166 (2017).
Potterton, E., Briggs, P., Turkenburg, M. & Dodson, E. A graphical user interface to the CCP4 program suite. Acta Crystallogr. D 59, 1131–1137 (2003).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Krissinel, E. Crystal contacts as nature’s docking solutions. J. Comput. Chem. 31, 133–143 (2010).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Diamond, S. et al. Mediterranean grassland soil C-N compound turnover is dependent on rainfall and depth, and is mediated by genomically divergent microorganisms. Nat. Microbiol. 4, 1356–1367 (2019).
Lavy, A. et al. Microbial communities across a hillslope–riparian transect shaped by proximity to the stream, groundwater table, and weathered bedrock. Ecol. Evol. 9, 6869–6900 (2019).
Knight, S., Andersson, I. & Brändén, C. I. Crystallographic analysis of ribulose 1,5-bisphosphate carboxylase from spinach at 2.4 A resolution. Subunit interactions and active site. J. Mol. Biol. 215, 113–160 (1990).
Acknowledgements
D.M.B., A.K.L. and P.M.S. acknowledge support from a Society in Science–Branco Weiss fellowship from ETH Zurich. J.H.P., P.D.A. and P.M.S. acknowledge support from the Joint BioEnergy Institute which is supported by the US Department of Energy, Office of Science, Office of Biological and Environmental Research under contract no. DE-AC02-05CH11231 between LBNL and the US Department of Energy. C.H. and J.F.B. thank A. Lavy and A. Sharrar for providing unpublished Rubisco sequences, J. West-Roberts for assistance, the Rifle IFRC/SFA 2.0 Metagenomics and Proteomics Data Analysis Project, the Allen Foundation, the Chan Zuckerberg Biohub and the Innovative Genomics Institute for support. C.H. acknowledges the Camille and Henry Dreyfus Foundation for a postdoctoral fellowship and the Joint Genome Institute CSP for sequencing. M.H. acknowledges support from the Department of Energy BER Integrated Diffraction Analysis Technologies (IDAT) program, NIGMS grant no. P30 GM124169-01 (ALS-ENABLE) for SAXS data collection at SIBYLS. D.J.O., M.A.J.P. and E.C.S. acknowledge support from the UK Biotechnology and Biological Sciences Research Council grant no. BB/I024488/1. We thank M. Hayer-Hartl (Max Planck Institute of Biochemistry, Martinsried, Germany) for the kind donation of the Syn6301-rbcL-pET11a, Syn6301-rbcLS-pET11a, pG-KJE8 and pBAD33ES/EL plasmids used in this study. Additionally, we thank N. Prywes for the kind donation of the pET28-His14-bdSUMO and pSF1389 plasmids. We also thank F. Guo and the UC Davis BioEM core facility for electron microscope images and the laboratory of J. Siegel (UC Davis Genome Center) for use of their qPCR machine for protein thermal shift experiments. We are grateful to A. Flamholz for collecting publicly available form I Rubisco kinetic data used in this study, and to A. Marinas and R. Vermon Callado for assisting with enzyme purifications. We thank K. Markel for his edits and suggestions on the manuscript.
Author information
Authors and Affiliations
Contributions
D.M.B., A.K.L. and P.M.S. designed experiments. D.M.B. and A.K.L. prepared all protein samples and performed all PAGE analyses and protein thermal shift experiments. M.H. performed all SEC–SAXS–MALS experiments and data analysis. J.H.P. performed X-ray crystallography data acquisition, image processing and structure determination. D.M.B. performed all structural analyses. A.K.L. performed all site-directed mutagenesis experiments. D.J.O. performed all Rubisco activity and kinetic measurements. C. H. and J.F.B performed all metagenomic and phylogenetic analyses. All authors participated in writing and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Martin Hagemann, Spencer Whitney and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Distribution of form I' Chloroflexi genomes.
Maximum-likelihood phylogenetic tree of Chloroflexi using ribosomal protein S3 (rpS3) as a marker gene. To map the distribution of form I' Rubisco genes onto genomes, all MAGs were scanned for presence of both rpS3 and form I' Rubisco. MAGs containing form I' Rubisco are highlighted in orange. The scaffolds that encode RbcL vary in size substantially, ranging up to ~106 kbp in length (available as Supplementary Data). At least partial genomic context could be determined in most cases and the gene for phosphoribulokinase was adjacent. In some cases, additional CBB Cycle pathway genes were present in an operon with Rubisco, strongly supporting the function of Rubisco in this pathway. In a subset of cases, other pentose phosphate pathway genes were co-encoded. In no case was there evidence for RbcS, either on the scaffold or in the draft genome bin (where a bin was available). Gene predictions were established via a standard annotation pipeline73,74 and augmented by HMM-based profiling and domain analysis.
Extended Data Fig. 2 In form I-containing Chloroflexi operons, rbcL and rbcS are always found next to each other, unlike form I'-containing Chloroflexi operons that lack rbcS.
Fragment operons from an example set of 10 form I Rubisco-containing Chloroflexi genomes shows that rbcS is always found next to rbcL, similar to form I Rubisco found in cyanobacteria and proteobacteria11. form I' Rubisco-containing Chloroflexi genomes do not contain small subunit rbcS (Fig. 1b). Scaffold names are shown to the right of their corresponding genome fragments.
Extended Data Fig. 3 PAGE analyses.
a, Non-denaturing PAGE gel with a molecular weight marker (M, lane 1), and purified proteins of all three candidate form I' Rubisco (P. breve, 241187, and 170907) with (+) or without (-) prior activation and incubation with 10-fold molar excess of 2CABP. 241187 and 170907 denotes scaffolds B_1_S1_170907_scaffold_241187_5_Tax=RBG_16_Chloroflexi_63_12 and S_p2_S4_170907_scaffold_85440 Rubisco, respectively. b, SDS–PAGE analysis of crude cell lysate from 1) overexpression of untagged P. breve Rubisco with co-expression of GroEL/ES from pBAD33EL/ES, 2) overexpression of His14-bdSUMO-tagged P. breve Rubisco with co-expression of GroEL/ES from pBAD33EL/ES, and 3) overexpression of His14-bdSUMO-tagged P. breve Rubisco without overexpression of GroEL/ES (background GroEL/ES expression from E. coli). Without GroEL/ES overexpression, untagged RbcL comprises 8 ± 1.0 (n = 3) of the total soluble protein, which improves to 14 ± 0.5 (n = 3) when GroEL/ES overexpression in induced (see Methods). When the His14-bdSUMO tag is included on the N-terminal end of RbcL, soluble expression is 7 ± 0.8 (n = 3) and 14 ± 0.8 (n = 3) of the total soluble protein, without and with GroEL/ES overexpression, respectively. Reported values collected from n separate experiments (separately grown E. coli cultures) reflect the mean ± standard deviation.
Extended Data Fig. 4 Form I Rubisco possess a unique RbcL C-terminal extension that interacts with RbcS, which is not found in form I' Rubisco.
a, Sequence alignment of representative Rubisco RbcL sequences from forms I, I', II, II/III, IIIA and IIIb. Strictly conserved residues have a red background, residues well conserved within a group are indicated by red letters, and the remaining residues are in black letters. Gaps are represented by dots. Residue numbering along the top refers to P. breve RbcL. Symbols above blocks of sequences correspond to the secondary structure of P. breve RbcL: α, α-helix; β, β-strand; η, 310-helix. The secondary structure elements were named according to Knight et al., 199075. The positions of loop 6 (black dotted lined), the form II/III-specific Rubisco assembly domain (cyan line), and the form I-specific C-terminal extension (purple line) are indicated. The RbcX binding domain-specific to form IB Rubisco is boxed in pink. The sequence alignment was created using the UniProt RbcL sequences P22859 (Allochromatium vinosum), O85040 (Halothiobacillus neapolitanus), A0A4D4IZ26 (Zea mays), P00880 (Syn6301), Q1QH22 (Nitrobacter hamburgensis), Q3IYC2 (Rhodobacter sphaeroides), P51226 (Porphyra purpurea), Q9GGQ2 (Vaucheria litorea), E1IGS1 (Oscillochloris trichoides), A0A0P9FAF0 (Kouleothrix aurantiaca), A4WW35 (Rhodobacter sphaeroides), P04718 (Rhodospirillum rubrum), Q12TQ0 (Methanococcoides burtonii), A0A1L3Q3Y6 (Methanohalophilus halophilus), B5IH56 (Aciduliprofundum boonei), O93627 (Thermococcus kodakarensis), J1ANE7 (Methanofollis liminatans), and Q2FSY4 (Methanospirillum hungatei). The sequences for representative form I' homologues are presented in this study (Supplementary Data 1). b, Overlay of amino acid residues 408-458 of Syn6301 Rubisco (tan) with residues 415-453 of P. breve Rubisco (blue) depicting the unique RbcL C-terminal extension found in form I enzymes, but not in Rubisco homologues that do not possess RbcS. Residues R428, N429, and E430 of Syn6301 RbcL contact residues N29 and Y32 at the interface of Syn6301 RbcS (purple).
Extended Data Fig. 5 Negative-staining electron microscopy 2D images of P. breve Rubisco.
Images reflect the highest resolution data collected with activated P. breve Rubisco in phosphate buffer. The experiment was performed once (n = 1).
Extended Data Fig. 6 Extended SEC-SAXS-MALS data.
Experimental SAXS profiles (black) of P. breve Rubisco in the absence (purple) or presence (blue) of bound 2CABP is displayed with the calculated scattering from the atomistic models shown in Fig. 3c. Inset shows the Guinier plot of experimental SAXS profiles with the linear fit in the q×Rg < 1.6 limits.
Extended Data Fig. 7 Amino acid sequence alignment of Syn6301 RbcL and P. breve RbcL.
a, Structure-based sequence alignment was originally made using PROMALS3D67 using 1RBL and 6URA structures, then aligned with the complete RbcL sequences using MAFFT68. Darker shades indicate higher sequence conservation between amino acids. Syn6301 and P. breve RbcL residues involved in dimer–dimer interactions are highlighted in green and blue, respectively. Syn6301 RbcL residues involved in RbcS contacts are annotated with red stars. All contact residues were identified using CCP4 CONTACTS69. b-c, Cross-section depictions of 1RBL, without RbcS, and P. breve Rubisco highlighting dimer–dimer interactions as in panel a. d, Map of Syn6301 RbcL residues involved in RbcS interactions, highlighted in red as in panel a.
Extended Data Fig. 8 Mutating key amino acid residues at the dimer–dimer interface of P. breve Rubisco disrupts octameric oligomeric assembly.
Native PAGE gel of recombinant WT, K150A, D161A, W165A, D220A, and Y224A P. breve Rubisco. Native Mark protein ladder denoted by ‘M’. Site-directed mutants destabilize the interface between RbcL dimers leading to break down of higher-order (that is, L8) oligomers into Rubisco species with variable oligomeric state and conformations, which results in a variety of lower molecular weight migration patterns within the Native PAGE gel. Experiment was performed once (n = 1).
Extended Data Fig. 9 Site-directed mutagenesis of Syn6301 dimer–dimer interface residues imparts marginal stability in the absence of RbcS.
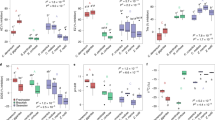
a, Protein thermal shift data displaying the mean fluorescent signal collected from four separate trials for WT Syn6301 RbcL, three separate mutant proteins, L158W, V154D, D349R and a combined four mutant protein, 4SDM (L158W, V154D, F217Y, and D349R). Mutations were designed to reflect homologous dimer–dimer interface residues present in P. breve Rubisco. The peaks corresponding to thermal denaturation of L8 quaternary structure are boxed, and analysis statistics are presented in the below table. Tm values represent the mean and standard deviation of n number of experiments conducted with the same protein sample. Two-tailed P-values for unpaired t test with Welch’s corrections are reported in the last column using WT Syn6301 RbcL as the reference comparison. n = number of technical replicates conducted in experiment. ns = not significant. ** P < 0.005, *** P < 0.0005. b, Native gel of purified recombinant WT and mutant Syn6301 proteins used in experiment.
Supplementary information
Supplementary Information
Supplementary note and Tables 1–3.
Supplementary Data 1
Fasta file containing protein amino acid sequences for form I' enzymes identified from MAGs.
Supplementary Data 2
Representative MAG genbank scaffolds.
Supplementary Data 3
Site-directed mutagenesis primers and synthesized candidate form I' rbcL gene sequences.
Rights and permissions
About this article
Cite this article
Banda, D.M., Pereira, J.H., Liu, A.K. et al. Novel bacterial clade reveals origin of form I Rubisco. Nat. Plants 6, 1158–1166 (2020). https://doi.org/10.1038/s41477-020-00762-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-00762-4
This article is cited by
-
Anoxygenic phototroph of the Chloroflexota uses a type I reaction centre
Nature (2024)
-
Discovery of a readily heterologously expressed Rubisco from the deep sea with potential for CO2 capture
Bioresources and Bioprocessing (2021)