Abstract
Diel and seasonal oscillations are two major environmental changes in nature. While organisms cope with the former by the well-characterized mechanism of the circadian clock1,2, there is limited information on the molecular mechanisms underlying long-term responses to the latter3,4,5. Histone H3 lysine 27 trimethylation (H3K27me3), a repressive histone modification, imparts stability and plasticity to gene regulation during developmental transitions6,7,8,9. Here we studied the seasonal and diel dynamics of H3K27me3 at the genome-wide level in a natural population of perennial Arabidopsis halleri and compared these dynamics with those of histone H3 lysine 4 trimethylation (H3K4me3), an active histone modification. Chromatin immunoprecipitation sequencing revealed that H3K27me3 exhibits seasonal plasticity and diel stability. Furthermore, we found that the seasonal H3K27me3 oscillation is delayed in phase relative to the H3K4me3 oscillation, particularly for genes associated with environmental memory. Our findings suggest that H3K27me3 monitors past transcriptional activity to create long-term gene expression trends during organismal responses over weeks in natural fluctuating environments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The ChIP-seq dataset has been deposited in the DNA Data Bank of Japan (DDBJ) under accession numbers DRA007511 (diel) and DRA007520 (seasonal). The ChIP-seq graph files are available at http://sohi.ecology.kyoto-u.ac.jp/AhgRNAseq/AhgChIPseq_bigwig.zip. The genome browser session is available at http://sohi.ecology.kyoto-u.ac.jp/jbrowse-ahg-chip/. All other data in this study are available at http://sohi.ecology.kyoto-u.ac.jp/AhgRNAseq/Nishio_Nat.Plants_script_data.zip.
Code availability
The R code used in this study is available at https://github.com/hnishio/Nishio_Nat.Plants_script.
Change history
23 November 2020
A Correction to this paper has been published: https://doi.org/10.1038/s41477-020-00757-1.
References
Millar, A. J. The intracellular dynamics of circadian clocks reach for the light of ecology and evolution. Annu. Rev. Plant Biol. 67, 595–618 (2016).
Webb, A. A. R., Seki, M., Satake, A. & Caldana, C. Continuous dynamic adjustment of the plant circadian oscillator. Nat. Commun. 10, 550 (2019).
Kudoh, H. Molecular phenology in plants: in natura systems biology for the comprehensive understanding of seasonal responses under natural environments. New Phytol. 210, 399–412 (2016).
Nishio, H. et al. From the laboratory to the field: assaying histone methylation at FLOWERING LOCUS C in naturally growing Arabidopsis halleri. Genes Genet. Syst. 91, 15–26 (2016).
Nishio, H. et al. Repressive chromatin modification underpins the long-term expression trend of a perennial flowering gene in nature. Nat. Commun. 11, 2065 (2020).
Dambacher, S., Hahn, M. & Schotta, G. Epigenetic regulation of development by histone lysine methylation. Heredity 105, 24–37 (2010).
Song, J., Irwin, J. & Dean, C. Remembering the prolonged cold of winter. Curr. Biol. 23, R807–R811 (2013).
Zheng, H. et al. Resetting epigenetic memory by reprogramming of histone modifications in mammals. Mol. Cell 63, 1066–1079 (2016).
Inoue, A., Jiang, L., Lu, F., Suzuki, T. & Zhang, Y. Maternal H3K27me3 controls DNA methylation-independent imprinting. Nature 547, 419–424 (2017).
Nagano, A. J. et al. Annual transcriptome dynamics in natural environments reveals plant seasonal adaptation. Nat. Plants 5, 74–83 (2019).
Aikawa, S., Kobayashi, M. J., Satake, A., Shimizu, K. K. & Kudoh, H. Robust control of the seasonal expression of the Arabidopsis FLC gene in a fluctuating environment. Proc. Natl Acad. Sci. USA 107, 11632–11637 (2010).
Satake, A. et al. Forecasting flowering phenology under climate warming by modelling the regulatory dynamics of flowering-time genes. Nat. Commun. 4, 2303 (2013).
Malapeira, J., Khaitova, L. C. & Mas, P. Ordered changes in histone modifications at the core of the Arabidopsis circadian clock. Proc. Natl Acad. Sci. USA 109, 21540–21545 (2012).
Black, J. C., Van Rechem, C. & Whetstine, J. R. Histone lysine methylation dynamics: establishment, regulation, and biological impact. Mol. Cell 48, 491–507 (2012).
Steffen, P. A. & Ringrose, L. What are memories made of? How Polycomb and Trithorax proteins mediate epigenetic memory. Nat. Rev. Mol. Cell Biol. 15, 340–356 (2014).
Zhang, T., Cooper, S. & Brockdorff, N. The interplay of histone modifications—writers that read. EMBO Rep. 16, 1467–1481 (2015).
O’Shea, P. Phase Measurement. in The Measurement, Instrumentation, and Sensors Handbook (ed. Webster, J. G.) Ch. 41 (CRC Press, 1999).
Sung, S. & Amasino, R. M. Vernalization in Arabidopsis thaliana is mediated by the PHD finger protein VIN3. Nature 427, 159–164 (2004).
Bastow, R. et al. Vernalization requires epigenetic silencing of FLC by histone methylation. Nature 427, 164–167 (2004).
Sheldon, C. C., Jean Finnegan, E., James Peacock, W. & Dennis, E. S. Mechanisms of gene repression by vernalization in Arabidopsis. Plant J. 59, 488–498 (2009).
Kim, D.-H. & Sung, S. Coordination of the vernalization response through a VIN3 and FLC gene family regulatory network in Arabidopsis. Plant Cell 25, 454–469 (2013).
Ding, Y., Fromm, M. & Avramova, Z. Multiple exposures to drought ‘train’ transcriptional responses in Arabidopsis. Nat. Commun. 3, 740 (2012).
Liu, N., Fromm, M. & Avramova, Z. H3K27me3 and H3K4me3 chromatin environment at super-induced dehydration stress memory genes of Arabidopsis thaliana. Mol. Plant 7, 502–513 (2014).
Liu, X. et al. Distinct features of H3K4me3 and H3K27me3 chromatin domains in pre-implantation embryos. Nature 537, 558–562 (2016).
Engelhorn, J. et al. Dynamics of H3K4me3 chromatin marks prevails over H3K27me3 for gene regulation during flower morphogenesis in Arabidopsis thaliana. Epigenomes 1, 8 (2017).
Buzas, D. M., Robertson, M., Finnegan, E. J. & Helliwell, C. A. Transcription-dependence of histone H3 lysine 27 trimethylation at the Arabidopsis polycomb target gene FLC. Plant J. 65, 872–881 (2011).
Hosogane, M., Funayama, R., Shirota, M. & Nakayama, K. Lack of transcription triggers H3K27me3 accumulation in the gene body. Cell Rep. 16, 696–706 (2016).
Finnegan, E. J. et al. Polycomb proteins regulate the quantitative induction of VERNALIZATION INSENSITIVE 3 in response to low temperatures. Plant J. 65, 382–391 (2011).
Bellegarde, F. et al. Polycomb Repressive Complex 2 attenuates the very high expression of the Arabidopsis gene NRT2.1. Sci. Rep. 8, 7905 (2018).
Park, E. Y., Tsuyuki, K. M., Hu, F., Lee, J. & Jeong, J. PRC2-mediated H3K27me3 contributes to transcriptional regulation of FIT-dependent iron deficiency response. Front. Plant Sci. 10, 627 (2019).
Gendrel, A.-V., Lippman, Z., Martienssen, R. & Colot, V. Profiling histone modification patterns in plants using genomic tiling microarrays. Nat. Methods 2, 213–218 (2005).
HiSeq Control Software (v.2.0.12.0) (Illumina, Inc., 2013).
Real Time Analysis (v.1.17.21.3) (Illumina, Inc., 2013).
BCL2FASTQ (v.1.8.4) (Illumina, Inc., 2013).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Briskine, R. V. et al. Genome assembly and annotation of Arabidopsis halleri, a model for heavy metal hyperaccumulation and evolutionary ecology. Mol. Ecol. Resour. 17, 1025–1036 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Cheng, C.-Y. et al. Araport11: a complete reannotation of the Arabidopsis thaliana reference genome. Plant J. 89, 789–804 (2017).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Barnett, A. G. & Dobson, A. J. Cosinor. in Analysing Seasonal Health Data (eds Barnett, A. G. & Dobson, A. J.) 75–92 (Springer Berlin Heidelberg, 2010).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Nagano, A. J. et al. Deciphering and prediction of transcriptome dynamics under fluctuating field conditions. Cell 151, 1358–1369 (2012).
Hui, W., Gel, Y. R. & Gastwirth, J. L. lawstat: an R package for law, public policy and biostatistics. J. Stat. Softw. 28, 3 (2008).
Nychka, D., Furrer, R., Paige, J. & Sain, S. fields: Tools for spatial data. R package version 10.3 (2015).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Pollard, K. S., Dudoit, S. & van der Laan, M. J. in Bioinformatics and Computational Biology Solutions Using R and Bioconductor (eds. Gentleman, R. et al.) 249–271 (Springer, 2005).
Asaf, S. et al. Chloroplast genomes of Arabidopsis halleri ssp. gemmifera and Arabidopsis lyrata ssp. petraea: structures and comparative analysis. Sci. Rep. 7, 7556 (2017).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Acknowledgements
We thank T. Kawagoe and J. Sugisaka for support during fieldwork; S. Sugano, M. Tosaka, T. Horiuchi, T. Kikuchi and K. Imamura for support in genome sequencing; and D. M. Buzas, K. Iwayama and A. Dodd for comments on the manuscript. The computations were performed partially on the NIG supercomputer at the ROIS National Institute of Genetics. This study was supported by JST CREST grant no. JPMJCR15O1, JSPS Grant-in-Aid for Scientific Research (S) no. 26221106 and (A) no. 19H01001, and MEXT KAKENHI grant no. 221S0002.
Author information
Authors and Affiliations
Contributions
H.N., A.J.N. and H.K. designed the study. H.N. and Y.S. performed the ChIP-seq experiments. H.N. and T.I. analysed the ChIP-seq data. H.N. and H.K. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genome-wide enrichment patterns of H3K27me3 and H3K4me3 in seasonal and diel ChIP-seq.
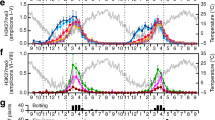
a, Genome browser view of the H3K27me3 and H3K4me3 density at AhgCCA1 (left) and AhgTOC1 (right) from diel ChIP-seq, shown in rpkm on the y axis. Black arrows indicate gene orientation and represent 1 kbp. b, c, Metagene plots of the H3K27me3 and H3K4me3 accumulation in the genes (b) and at the TSS (c) and their ±2-kbp flanking regions. Rpm was calculated for 50 bp sliding windows along each region. Annual means of rpm in the seasonal ChIP-seq are shown. The grey shading in b indicates standardised gene lengths. d–f, Seasonal and diel changes in H3K27me3 and H3K4me3 at genes with seasonal oscillations of both modifications (d), at genes with seasonal oscillations of either H3K27me3 or H3K4me3 (e), and at genes with diel oscillations of H3K4me3 (f). The values represent log2 rpkm at gene regions (H3K27me3) and 1 kbp-downstream regions from TSS (H3K4me3). Circles represent observations of seasonal and diel changes, and solid lines represent spline curves. The spline curves are shown irrespective of presence/absence of oscillations. The grey shading indicates time between sunset and sunrise. g, The seasonal and diel amplitudes were compared for H3K27me3- and H3K4me3-enriched genes. The boxes span from the first to the third quartiles, the lines inside the boxes represent the medians, and the whiskers above and below the boxes represent 1.5 × the interquartile range from the quartiles; Brunner-Munzel tests (two-sided).
Extended Data Fig. 2 Comparison between seasonal and diel oscillation amplitudes.
a, Scatter plots comparing seasonal and diel amplitudes of H3K27me3 (left) and H3K4me3 (right) for H3K27me3- and H3K4me3-enriched genes, respectively. b, The ratio of seasonal amplitude to diel amplitude for H3K27me3 and H3K4me3. c–h, Four out of 12 time points [three months intervals starting from Nov. (c, d), Dec. (e, f), and Jan. (g, h)] were used to calculate seasonal amplitudes of H3K27me3 and H3K4me3 for all genes, otherwise the same as a and b. We compared seasonal and diel data with the same number of time points per cycle by reducing data points of seasonal data. In b, d, f, h, The boxes span from the first to the third quartiles, the bands inside the boxes represent the medians, and the whiskers above and below the boxes represent 1.5 × the interquartile range from the quartiles; Brunner-Munzel tests (two-sided).
Extended Data Fig. 3 Definition of the genes with H3K27me3 and H3K4me3 seasonal oscillations.
a, b, The distribution of H3K27me3 and H3K4me3 enrichment for all genes in seasonal (a) and diel (b) ChIP-seq. Enriched genes were defined by the threshold of log2 (maximum rpkm) > 2 (red lines). c, d, The distribution of seasonal (c) and diel (d) amplitudes of H3K27me3 and H3K4me3 for the enriched genes. As one of the criteria of the genes with the seasonal and diel oscillations of H3K27me3 or H3K4me3, the threshold of log2 (amplitude) > 1 (red lines) was set. e, Seasonal and diel changes in H3K27me3 and H3K4me3 at AhgFLC, AhgLTI30, AhgCCA1, and AhgTOC1. The values represent the standardised levels at gene regions (H3K27me3) and 1 kbp-downstream regions from the TSS (H3K4me3). Circles represent observations of seasonal and diel changes, and solid lines represent significant seasonal patterns estimated by cosinor models (P < 0.05 for the coefficients of sine and cosine terms; n.s., not significant). The grey shading indicates time between sunset and sunrise. f, g, The genes with seasonal oscillations of H3K27me3 and H3K4me3 were defined as the overlap of the three categories: maximum/minimum rpkm > 2, FDR < 0.05 in the generalised linear model likelihood ratio test using edgeR, and P < 0.05 for the coefficients of sine and cosine terms using cosinor models. The statistical tests were two sided. h, i, The genes with diel oscillations of H3K27me3 and H3K4me3 were defined in the same way as f and g.
Extended Data Fig. 4 H3K27me3 and H3K4me3 accumulation patterns revealed by the seasonal ChIP-seq.
a, The enriched gene ontology terms of the genes that exhibited winter accumulation of H3K27me3. b, The enriched gene ontology terms of the genes that exhibited winter (upper) and summer (lower) accumulation of H3K4me3. c, Monthly patterns of histone modification levels for the genes with dual seasonal oscillations. The genes were aligned according to the H3K27me3 peak date. d, The genes were aligned according to the H3K4me3 peak date, otherwise the same as b.
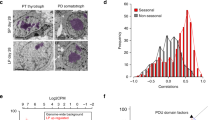
Extended Data Fig. 5 Correlation between seasonal patterns of histone modifications and environmental factors.
a, Simple moving averages (SMAs) of environmental factors (temperature, precipitation, day length, and sunlight hours) for past 1 week (black) and 1 month (red). b, The number of genes which showed the highest correlation between histone modifications (left, H3K27me3; right, H3K4me3) and SMAs of one of the environmental factors for past 1 month. Genes are shown that have significant correlations with one of the environmental factors (P < 0.001; Spearman’s rank correlation, two-sided; corrected by the Benjamini-Hochberg method): 89% and 83% of the genes with seasonal oscillations of H3K27me3 and H3K4me3, respectively.
Extended Data Fig. 6 The effects of H3K27me3 and H3K4me3 seasonal oscillations on gene expression.
a, Venn diagram displaying all categories in which genes are classified by enrichment and seasonality of histone modifications. b, c, The mean of mRNA accumulation for the gene categories with different combinations of enrichment and seasonality of histone modifications. The presence and absence of H3K27me3 (b) and H3K4me3 (c) seasonality was compared. Brunner-Munzel tests (one-sided; corrected by the Benjamini-Hochberg method). d, e, The seasonal amplitude (d) and mean (e) of transcript accumulation for all gene categories with different combinations of enrichment and seasonality of histone modifications. Steel-Dwass tests (two-sided), P < 0.05. In b–e, the boxes span from the first to the third quartiles, the bands inside the boxes represent the medians, and the whiskers above and below the boxes represent 1.5 × the interquartile range from the quartiles.
Extended Data Fig. 7 Phase difference between the H3K27me3 and H3K4me3 dynamics.
a, Frequency distribution of the temporal differences between the trough of H3K27me3 and the peak of H3K4me3 estimated using cosinor models, illustrating genes with advanced (left) and delayed (right) H3K27me3 relative to no difference (vertical line). Because the trough appears six months later after the peak in the cosinor model, the temporal differences between the peak of H3K27me3 and the trough of H3K4me3 shows the identical distribution. b, Conceptual diagrams illustrating the relationship between the degree of phase difference and Lissajous curves. c–e, The Lissajous curve (red line) drawn as a spline of observed values (black dots) of H3K27me3 against H3K4me3 for AhgVIN3 (c), AhgMAF1 (d), and AhgRAB18 (e). The area inside the curve represents the degree of phase difference. Grey dots: all values for the genes with dual seasonal oscillations.
Extended Data Fig. 8 Direction of phase difference between the H3K27me3 and H3K4me3 dynamics for the top 20 genes with the largest areas of the Lissajous curves.
The values at the 12 sampling time points were estimated using smoothing splines and were connected by lines. The numbers next to the data points are chronological ordinals (1: November 6, 2012, 12: September 24, 2013). Open circles, filled circles, open squares, and filled squares represent the data points in autumn, winter, spring, and summer, respectively.
Extended Data Fig. 9 Reproducibility of the ChIP-seq experiments.
a, b, Correlation of log2 rpkm between replicates 1 and 2 in the seasonal ChIP-seq for H3K27me3 (a) and H3K4me3 (b). c, d, Correlation of log2 rpkm between replicates 1 and 2, 1 and 3, and 1 and 4 in the diel ChIP-seq for H3K27me3 (c) and H3K4me3 (d). In a–d, log2 rpkm was calculated from the number of reads overlapping the transcribed region (H3K27me3) and the region from TSS to 1,000 bp downstream (H3K4me3) for each gene.
Supplementary information
Supplementary Data 1
The list of the genes with the seasonal oscillations of H3K27me3.
Supplementary Data 2
The list of the genes with the seasonal oscillations of H3K4me3.
Supplementary Data 3
The list of the genes with the dual seasonal oscillations.
Rights and permissions
About this article
Cite this article
Nishio, H., Nagano, A.J., Ito, T. et al. Seasonal plasticity and diel stability of H3K27me3 in natural fluctuating environments. Nat. Plants 6, 1091–1097 (2020). https://doi.org/10.1038/s41477-020-00757-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-00757-1
This article is cited by
-
Diurnal oscillations of epigenetic modifications are associated with variation in rhythmic expression of homoeologous genes in Brassica napus
BMC Biology (2023)
-
Duration of cold exposure defines the rate of reactivation of a perennial FLC orthologue via H3K27me3 accumulation
Scientific Reports (2020)