Abstract
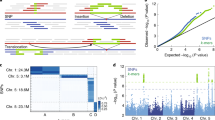
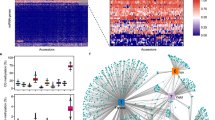
The extent to which sequence variation impacts plant fitness is poorly understood. High-resolution maps detailing the constraint acting on the genome, especially in regulatory sites, would be beneficial as functional annotation of noncoding sequences remains sparse. Here, we present a fitness consequence (fitCons) map for rice (Oryza sativa). We inferred fitCons scores (ρ) for 246 inferred genome classes derived from nine functional genomic and epigenomic datasets, including chromatin accessibility, messenger RNA/small RNA transcription, DNA methylation, histone modifications and engaged RNA polymerase activity. These were integrated with genome-wide polymorphism and divergence data from 1,477 rice accessions and 11 reference genome sequences in the Oryzeae. We found ρ to be multimodal, with ~9% of the rice genome falling into classes where more than half of the bases would probably have a fitness consequence if mutated. Around 2% of the rice genome showed evidence of weak negative selection, frequently at candidate regulatory sites, including a novel set of 1,000 potentially active enhancer elements. This fitCons map provides perspective on the evolutionary forces associated with genome diversity, aids in genome annotation and can guide crop breeding programs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The read data used to generate the ChromHMM model and genomic classes have been deposited at the NCBI SRA (https://www.ncbi.nlm.nih.gov/sra) and can be accessed through BioProject ID PRJNA586887. Genome assemblies of O. officinalis and O. australiensis are available from the CoGe CyVerse website (https://genomevolution.org/coge/) with genome IDs id56031 and id56030, respectively. Access to genomic class annotation and INSIGHT scoring of the rice genome is available via a genome browser linked from the project’s website (http://purugganan-genomebrowser.bio.nyu.edu/insightJuly2018/greenInsight.html). All epigenomic data tracks, genome annotations, multiple alignments, conservation scores, fitCons scores and site classes are available for visualization and download on a local installation on the USCSC Genome Browser at http://purugganan-genomebrowser.bio.nyu.edu/cgi-bin/hgTracks?db=Osaj&position=Osaj.1%3A166356–178595, and are also available for download from the NCBI SRA (PRJNA586887). The greenINSIGHT-specific data used to generate the greenINSIGHT online tool are available in the “Additional information, scripts & data” section at http://purugganan-genomebrowser.bio.nyu.edu/insightJuly2018/greenInsight.html. The greenINSIGHT-specific code used to generate the greenINSIGHT online tool, as well as the code described in the Methods, are available in the “Additional information, scripts & data” section at http://purugganan-genomebrowser.bio.nyu.edu/insightJuly2018/greenInsight.html.
Code availability
The greenINSIGHT-specific data used to generate the greenINSIGHT online tool are available in the “Additional information, scripts & data” section at http://purugganan-genomebrowser.bio.nyu.edu/insightJuly2018/greenInsight.html. The greenINSIGHT-specific code used to generate the greenINSIGHT online tool, as well as the code described in the Methods, are available in the “Additional information, scripts & data” section at http://purugganan-genomebrowser.bio.nyu.edu/insightJuly2018/greenInsight.html.
References
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Schrider, D. R. & Kern, A. D. Inferring selective constraint from population genomic data suggests recent regulatory turnover in the human brain. Genome Biol. Evol. 7, 3511–3528 (2015).
Gronau, I., Arbiza, L., Mohammed, J. & Siepel, A. Inference of natural selection from interspersed genomic elements based on polymorphism and divergence. Mol. Biol. Evol. 30, 1159–1171 (2013).
McDonald, J. H. & Kreitman, M. Adaptive protein evolution at the Adh locus in Drosophila. Nature 351, 652–654 (1991).
Sawyer, S. A. & Hartl, D. L. Population genetics of polymorphism and divergence. Genetics 132, 1161–1176 (1992).
Bustamante, C. D. et al. Natural selection on protein-coding genes in the human genome. Nature 437, 1153–1157 (2005).
Smith, N. G. C. & Eyre-Walker, A. Adaptive protein evolution in Drosophila. Nature 415, 1022–1024 (2002).
Gulko, B., Hubisz, M. J., Gronau, I. & Siepel, A. A method for calculating probabilities of fitness consequences for point mutations across the human genome. Nat. Genet. 47, 276–283 (2015).
Gulko, B. & Siepel, A. An evolutionary framework for measuring epigenomic information and estimating cell-type-specific fitness consequences. Nat. Genet. 51, 335–342 (2019).
Wing, R. A., Purugganan, M. D. & Zhang, Q. The rice genome revolution: from an ancient grain to Green Super Rice. Nat. Rev. Genet. 19, 505–517 (2018).
Wang, W. et al. Genomic variation in 3,010 diverse accessions of Asian cultivated rice. Nature 557, 43–49 (2018).
Stein, J. C. et al. Genomes of 13 domesticated and wild rice relatives highlight genetic conservation, turnover and innovation across the genus Oryza. Nat. Genet. 50, 285–296 (2018).
Cao, J. et al. Whole-genome sequencing of multiple Arabidopsis thaliana populations. Nat. Genet. 43, 956–963 (2011).
Haudry, A. et al. An atlas of over 90,000 conserved noncoding sequences provides insight into crucifer regulatory regions. Nat. Genet. 45, 891–898 (2013).
Huang, X. et al. A map of rice genome variation reveals the origin of cultivated rice. Nature 490, 497–501 (2012).
Gutaker, R. M. et al. Genomic history and ecology of the geographic spread of rice. Preprint at https://www.biorxiv.org/content/10.1101/748178v1 (2019).
Josephs, E. B., Lee, Y. W., Stinchcombe, J. R. & Wright, S. I. Association mapping reveals the role of purifying selection in the maintenance of genomic variation in gene expression. Proc. Natl Acad. Sci. USA 112, 15390–15395 (2015).
Flowers, J. M. et al. Natural selection in gene-dense regions shapes the genomic pattern of polymorphism in wild and domesticated rice. Mol. Biol. Evol. 29, 675–687 (2012).
Caicedo, A. L. et al. Genome-wide patterns of nucleotide polymorphism in domesticated rice. PLoS Genet. 3, 1745–1756 (2007).
Bradnam, K. R. & Korf, I. Longer first introns are a general property of eukaryotic gene structure. PLoS ONE 3, e3093 (2008).
Rigau, M., Juan, D., Valencia, A. & Rico, D. Intronic CNVs and gene expression variation in human populations. PLoS Genet. 15, e1007902 (2019).
Berendzen, K. W. et al. Bioinformatic cis-element analyses performed in Arabidopsis and rice disclose bZIP- and MYB-related binding sites as potential AuxRE-coupling elements in auxin-mediated transcription. BMC Plant Biol. 12, 125 (2012).
Freeling, M., Rapaka, L., Lyons, E., Pedersen, B. & Thomas, B. C. G-boxes, bigfoot genes, and environmental response: characterization of intragenomic conserved noncoding sequences in Arabidopsis. Plant Cell 19, 1441–1457 (2007).
Choi, H. I., Hong, J. H., Ha, J. O., Kang, J. Y. & Kim, S. Y. ABFs, a family of ABA-responsive element binding factors. J. Biol. Chem. 275, 1723–1730 (2000).
Lu, T. et al. Function annotation of the rice transcriptome at single-nucleotide resolution by RNA-Seq. Genome Res. 20, 1238–1249 (2010).
Peng, T. et al. Differentially expressed microRNA cohorts in seed development may contribute to poor grain filling of inferior spikelets in rice. BMC Plant Biol. 14, 196 (2014).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Buenrostro, J. D., Wu, B., Chang, H. Y. & Greenleaf, W. J. ATAC-Seq: a method for assaying chromatin accessibility genome-wide. Curr. Protoc. Mol. Biol. 109, 21.29.1–21.29.9 (2015).
Feng, S. et al. Conservation and divergence of methylation patterning in plants and animals. Proc. Natl Acad. Sci. USA 107, 8689–8694 (2010).
Mahat, D. B. et al. Base-pair-resolution genome-wide mapping of active RNA polymerases using precision nuclear run-on (PRO-Seq). Nat. Protoc. 11, 1455–1476 (2016).
Kwak, H., Fuda, N. J., Core, L. J. & Lis, J. T. Precise maps of RNA polymerase reveal how promoters direct initiation and pausing. Science 339, 950–953 (2013).
Liu, Y. et al. PCSD: a plant chromatin state database. Nucleic Acids Res. 46, D1157–D1167 (2018).
Yan, W. et al. Dynamic control of enhancer activity drives stage-specific gene expression during flower morphogenesis. Nat. Commun. 10, 1705 (2019).
Wen, M. et al. Expression variations of miRNAs and mRNAs in rice (Oryza sativa). Genome Biol. Evol. 8, 3529–3544 (2016).
Zong, W., Zhong, X., You, J. & Xiong, L. Genome-wide profiling of histone H3K4-tri-methylation and gene expression in rice under drought stress. Plant Mol. Biol. 81, 175–188 (2013).
Lozano, R. et al. RNA polymerase mapping in plants identifies enhancers enriched in causal variants. Preprint at https://www.biorxiv.org/content/10.1101/376640v1 (2018).
Xia, J. et al. Detecting and characterizing microRNAs of diverse genomic origins via miRvial. Nucleic Acids Res. 45, e176 (2017).
Wilkins, O. et al. EGRINs (environmental gene regulatory influence networks) in rice that function in the response to water deficit, high temperature, and agricultural environments. Plant Cell 28, 2365–2384 (2016).
Tan, F. et al. Analysis of chromatin regulators reveals specific features of rice DNA methylation pathways. Plant Physiol. 171, 2041–2054 (2016).
Liu, C., Lu, F., Cui, X. & Cao, X. Histone methylation in higher plants. Annu. Rev. Plant Biol. 61, 395–420 (2010).
Liu, N., Fromm, M. & Avramova, Z. H3K27me3 and H3K4me3 chromatin environment at super-induced dehydration stress memory genes of Arabidopsis thaliana. Mol. Plant 7, 502–513 (2014).
Fang, H., Liu, X., Thorn, G., Duan, J. & Tian, L. Expression analysis of histone acetyltransferases in rice under drought stress. Biochem. Biophys. Res. Commun. 443, 400–405 (2014).
Du, Z. et al. Genome-wide analysis of histone modifications: H3K4me2, H3K4me3, H3K9ac, and H3K27ac in Oryza sativa L. Japonica. Mol. Plant 6, 1463–1472 (2013).
Lee, T., Zhai, J. & Meyers, B. C. Conservation and divergence in eukaryotic DNA methylation. Proc. Natl Acad. Sci. USA 107, 9027–9028 (2010).
Ernst, J. & Kellis, M. ChromHMM: automating chromatin-state discovery and characterization. Nat. Methods 9, 215–216 (2012).
Ernst, J. & Kellis, M. Chromatin-state discovery and genome annotation with ChromHMM. Nat. Protoc. 12, 2478–1492 (2017).
Roudier, F. et al. Integrative epigenomic mapping defines four main chromatin states in Arabidopsis. EMBO J. 30, 1928–1938 (2011).
Sequeira-Mendes, J. et al. The functional topography of the Arabidopsis genome is organized in a reduced number of linear motifs of chromatin states. Plant Cell 26, 2351–2366 (2014).
Liu, C. et al. Genome-wide analysis of chromatin packing in Arabidopsis thaliana at single-gene resolution. Genome Res. 26, 1057–1068 (2016).
Guo, H. & Moose, S. P. Conserved noncoding sequences among cultivated cereal genomes identify candidate regulatory sequence elements and patterns of promoter evolution. Plant Cell 15, 1143–1158 (2003).
Liu, L., Xu, W., Hu, X., Liu, H. & Lin, Y. W-box and G-box elements play important roles in early senescence of rice flag leaf. Sci. Rep. 6, 20881 (2016).
Ding, M. et al. Enhancer RNAs (eRNAs): new insights into gene transcription and disease treatment. J. Cancer 9, 2334–2340 (2018).
Wang, Z., Chu, T., Choate, L. A. & Danko, C. G. Identification of regulatory elements from nascent transcription using dREG. Genome Res. 29, 293–303 (2019).
Danko, C. G. et al. Dynamic evolution of regulatory element ensembles in primate CD4+ T cells. Nat. Ecol. Evol. 2, 537–548 (2018).
Savisaar, R. & Hurst, L. D. Exonic splice regulation imposes strong selection at synonymous sites. Genome Res. 28, 1442–1454 (2018).
Cannavò, E. et al. Shadow enhancers are pervasive features of developmental regulatory networks. Curr. Biol. 26, 38–51 (2016).
Prescott, S. L. et al. Enhancer divergence and cis-regulatory evolution in the human and chimp neural crest. Cell 163, 68–83 (2015).
Mezmouk, S. & Ross-Ibarra, J. The pattern and distribution of deleterious mutations in maize. G3 (Bethesda) 4, 163–171 (2014).
Wallace, J. G., Rodgers-Melnick, E. & Buckler, E. S. On the road to breeding 4.0: unraveling the good, the bad, and the boring of crop quantitative genomics. Annu. Rev. Genet. 52, 421–444 (2018).
Moyers, B. T., Morrell, P. L. & McKay, J. K. Genetic costs of domestication and improvement. J. Hered. 109, 103–116 (2018).
Morrell, P. L., Buckler, E. S. & Ross-Ibarra, J. Crop genomics: advances and applications. Nat. Rev. Genet. 13, 85–96 (2012).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10–12 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Kent, W. J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Raurell-Vila, H., Ramos-Rodríguez, M. & Pasquali, L. in CpG Islands. Methods in Molecular Biology Vol. 1766 (eds Vavouri, T. & Peinado, M. A.) 197–208 (Humana Press, 2018).
Hetzel, J., Duttke, S. H., Benner, C. & Chory, J. Nascent RNA sequencing reveals distinct features in plant transcription., Proc. Natl Acad. Sci. USA 113, 12316–12321 (2016).
Boisvert, S., Raymond, F., Godzaridis, É., Laviolette, F. & Corbeil, J. Ray Meta: scalable de novo metagenome assembly and profiling. Genome Biol. 13, R122 (2012).
Butler, J. et al. ALLPATHS: de novo assembly of whole-genome shotgun microreads. Genome Res. 18, 810–820 (2008).
Green, E. D. et al. Aligning multiple genomic sequences with the threaded blockset aligner. Genome Res. 14, 708–715 (2004).
Siepel, A. & Haussler, D. Phylogenetic estimation of context-dependent substitution rates by maximum likelihood. Mol. Biol. Evol. 21, 468–488 (2004).
Acknowledgements
We thank the New York University Center for Genomics and Systems Biology GenCore Facility and the Next Generation Sequencing core at Cold Spring Harbor Laboratory for sequencing support. We thank O. Wilkins and C. Danko for valuable suggestions relating to the ATAC and PRO-Seq protocols, respectively. This work was supported primarily by a grant from the Zegar Family Foundation (no. A16-0051-004), as well as some support from the National Science Foundation Plant Genome Research Program (no. IOS-1546218) and NYU Abu Dhabi Research Institute to M.D.P., the National Science Foundation CAREER award (no. MCB-1552455), the US National Institutes of Health (no. R35GM124806) and US Department of Agriculture Hatch Program (no. 1012915) to X.Z., the US National Institutes of Health (no. R35GM127070) to A.S., and fellowships from the Gordon and Betty Moore Foundation and Life Sciences Research Foundation (no. GBMF2550.06) to S.C.G. and from the Natural Sciences and Engineering Research Council of Canada (no. PDF-502464-2017) to Z.J.-L.
Author information
Authors and Affiliations
Contributions
M.D.P. conceived of the study idea. M.D.P., Z.J.-L., A.E.P. and A.S. designed the study. M.D.P. directed the study. Z.J.-L. and X.Z. collected the data, A.E.P., Z.J.-L., J.Y.C., B.G., S.C.G. and M.D.P. analysed the data. Z.J.-L., A.E.P., A.S. and M.D.P. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Robin Allaby, Peter Civan and Peter Morrell for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12.
Supplementary Tables
Supplementary Tables 1–11.
Rights and permissions
About this article
Cite this article
Joly-Lopez, Z., Platts, A.E., Gulko, B. et al. An inferred fitness consequence map of the rice genome. Nat. Plants 6, 119–130 (2020). https://doi.org/10.1038/s41477-019-0589-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-019-0589-3
This article is cited by
-
The first long-read nuclear genome assembly of Oryza australiensis, a wild rice from northern Australia
Scientific Reports (2022)
-
Rice functional genomics: decades’ efforts and roads ahead
Science China Life Sciences (2022)
-
Advancing crop genomics from lab to field
Nature Genetics (2021)
-
A quantitative genomics map of rice provides genetic insights and guides breeding
Nature Genetics (2021)