Abstract
Photosystem I (PSI) is present as trimeric complexes in most characterized cyanobacteria and as monomers in plants and algae. Recent reports of tetrameric PSI have raised questions regarding its structural basis, physiological role, phylogenetic distribution and evolutionary significance. Here, we examined PSI in 61 cyanobacteria, showing that tetrameric PSI, which correlates with the psaL gene and a distinct genomic structure, is widespread among heterocyst-forming cyanobacteria and their close relatives. Physiological studies revealed that expression of tetrameric PSI is favoured under high light, with an increased content of novel PSI-bound carotenoids (myxoxanthophyll, canthaxanthan and echinenone). In sum, this work suggests that tetrameric PSI is an adaptation to high light intensity, and that change in PsaL leads to monomerization of trimeric PSI, supporting the hypothesis of tetrameric PSI being the evolutionary intermediate in the transition from cyanobacterial trimeric PSI to monomeric PSI in plants and algae.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The cloned cyanobacterial psaL sequences have been deposited in GenBank with the accession numbers KY575410, KY575411, KY575412, KY575413, KY575414, KY575415, KY575416, KY575417, KY575418, KY575419, KY575420, KY575421, KY575422, KY575423 and KY575424. The TS-821 whole-genome shotgun project has been deposited at DNA Data Bank of Japan/European Nucleotide Archive/GenBank under the accession MVDI00000000. The version described in this paper is version MVDI01000000.
References
Raymond, J. & Blankenship, R. E. in Advances in Photosynthesis and Respiration Vol. 24 (Ed. Goldbeck, J. H.) 669–681 (Springer, 2007).
Gardian, Z. et al. Organisation of photosystem i and photosystem iI in red alga Cyanidium caldarium: encounter of cyanobacterial and higher plant concepts. Biochim. Biophys. Acta 1767, 725–731 (2007).
Watanabe, M., Kubota, H., Wada, H., Narikawa, R. & Ikeuchi, M. Novel supercomplex organization of photosystem I in Anabaena and Cyanophora paradoxa. Plant Cell Physiol. 52, 162–168 (2011).
Ben-Shem, A., Frolow, F. & Nelson, N. Crystal structure of plant photosystem I. Nature 426, 630–635 (2003).
Kouril, R., van Oosterwijk, N., Yakushevska, A. E. & Boekema, E. J. Photosystem I: a search for green plant trimers. Photochem. Photobiol. Sci. 4, 1091–1094 (2005).
Veith, T. & Buchel, C. The monomeric photosystem I-complex of the diatom Phaeodactylum tricornutum binds specific fucoxanthin chlorophyll proteins (FCPs) as light-harvesting complexes. Biochim. Biophys. Acta 1767, 1428–1435 (2007).
Wood, W. H. J. et al. Dynamic thylakoid stacking regulates the balance between linear and cyclic photosynthetic electron transfer. Nat. Plants 4, 116–12 (2018).
Boekema, E. J. et al. Evidence for a trimeric organization of the photosystem I complex from the thermophilic cyanobacterium Synechococcus sp. FEBS Lett. 217, 283–286 (1987).
Shubin, V. V., Bezsmertnaya, I. N. & Karapetyan, N. V. Isolation from Spirulina membranes of 2 photosystem i-type complexes, one of which contains chlorophyll responsible for the 77-K fluorescence band at 760 Nm. FEBS Lett. 309, 340–342 (1992).
Shubin, V. V., Tsuprun, V. L., Bezsmertnaya, I. N. & Karapetyan, N. V. Trimeric forms of the photosystem I reaction center complex pre‐exist in the membranes of the cyanobacterium Spirulina platensis. FEBS Lett. 334, 79–82 (1993).
Tsiotis, G., Haase, W., Engel, A. & Michel, H. Isolation and structural characterization of trimeric cyanobacterial photosystem I complex with the help of recombinant antibody fragments. Eur. J. Biochem. 231, 823–830 (1995).
Garczarek, L., van der Staay, G. W. M., Thomas, J. C. & Partensky, F. Isolation and characterization of photosystem I from two strains of the marine oxychlorobacterium Prochlorococcus. Photosynth. Res. 56, 131–141 (1998).
Bibby, T. S., Mary, I., Nield, J., Partensky, F. & Barber, J. Low-light-adapted Prochlorococcus species possess specific antennae for each photosystem. Nature 424, 1051–1054 (2003).
Boekema, E. J. et al. A giant chlorophyll–protein complex induced by iron defciency in cyanobacteria. Nature 412, 745–748 (2001).
Tucker, D. L. & Sherman, L. A. Analysis of chlorophyll–protein complexes from the cyanobacterium Cyanothece sp. ATCC 51142 by non-denaturing gel electrophoresis. Biochim. et Biophys. 1468, 150–160 (2000).
Casella, S. et al. Dissecting the native architecture and dynamics of cyanobacterial photosynthetic machinery. Mol. Plant 10, 1434–1448 (2017).
MacGregor-Chatwin, C. et al. Lateral segregation of photosystem I in cyanobacterial thylakoids. Plant Cell 29, 1119–1136 (2017).
Mangels, D. et al. Photosystem I from the unusual cyanobacterium Gloeobacter violaceus. Photosynth. Res. 72, 307–319 (2002).
Almog, O., Shoham, G., Michaeli, D. & Nechushtai, R. Monomeric and trimeric forms of photosystem I reaction center of Mastigocladus laminosus: crystallization and preliminary characterization. Proc. Natl Acad. Sci. USA 88, 5312–5316 (1991).
Jordan, P. et al. Three-dimensional structure of cyanobacterial photosystem I at 2.5 A resolution. Nature 411, 909–917 (2001).
Watanabe, M. et al. Attachment of phycobilisomes in an antenna–photosystem I supercomplex of cyanobacteria. Proc. Natl Acad. Sci. USA 111, 2512–2517 (2014).
Li, M., Semchonok, D. A., Boekema, E. J. & Bruce, B. D. Characterization and evolution of tetrameric photosystem I from the thermophilic cyanobacterium Chroococcidiopsis sp. TS-821. Plant Cell 26, 1230–1245 (2014).
Semchonok, D. A., Li, M., Bruce, B. D., Oostergetel, G. T. & Boekema, E. J. Cryo-EM structure of a tetrameric cyanobacterial photosystem I complex reveals novel subunit interactions. Biochim. Biophys. Acta 1857, 1619–1626 (2016).
Malavath, T., Caspy, I., Netzer-El, S. Y., Klaiman, D. & Nelson, N. Structure and function of wild-type and subunit-depleted photosystem I in Synechocystis. Biochim. Biophys. Acta 1859, 645–654 (2018).
Gan, F. et al. Extensive remodeling of a cyanobacterial photosynthetic apparatus in far-red light. Science 345, 1312–1317 (2014).
Gan, F. & Bryant, D. A. Adaptive and acclimative responses of cyanobacteria to far-red light. Environ. Microbiol. 17, 3450–3465 (2015).
Qin, X., Suga, M., Kuang, T. & Shen, J. R. Structural basis for energy transfer pathways in the plant PSI–LHCI supercomplex. Science 348, 989–995 (2015).
Mazor, Y., Borovikova, A. & Nelson, N. The structure of plant photosystem I super-complex at 2.8 A resolution. eLife 4, e07433 (2015).
Cardona, T. et al. Electron transfer protein complexes in the thylakoid membranes of heterocysts from the cyanobacterium Nostoc punctiforme. Biochim. Biophys. Acta 1787, 252–263 (2009).
Vintila, S. & El-Shehawy, R. Ammonium ions inhibit nitrogen fixation but do not affect heterocyst frequency in the bloom-forming cyanobacterium Nodularia spumigena strain AV1. Microbiology 153, 3704–3712 (2007).
Vintila, S. & El-Shehawy, R. Variability in the response of the cyanobacterium Nodularia spumigena to nitrogen supplementation. J. Environ. Monit. 12, 1885–1890 (2010).
Rippka, R., Deruelles, J., Waterbury, J. B., Herdman, M. & Stanier, R. Y. Generic assignments, strain histories and properties of pure cultures of cyanobacteria. J. Gen. Microbiol. 111, 1–61 (1979).
Fewer, D., Friedl, T. & Budel, B. Chroococcidiopsis and heterocyst-differentiating cyanobacteria are each other’s closest living relatives. Mol. Phylogenet. Evol. 23, 82–90 (2002).
Shih, P. M. et al. Improving the coverage of the cyanobacterial phylum using diversity-driven genome sequencing. Proc. Natl Acad. Sci. USA 110, 1053–1058 (2013).
Iwuchukwu, I. J. et al. Self-organized photosynthetic nanoparticle for cell-free hydrogen production. Nat. Nanotechnol. 5, 73–79 (2010).
Baker, D. R. et al. Comparative photoactivity and stability of isolated cyanobacterial monomeric and trimeric photosystem I. J. Phys. Chem. B 118, 2703–2711 (2014).
Wang, Q. et al. The high light-inducible polypeptides stabilize trimeric photosystem I complex under high light conditions in Synechocystis PCC 6803. Plant Physiol. 147, 1239–1250 (2008).
Dagan, T. et al. Genomes of Stigonematalean cyanobacteria (subsection V) and the evolution of oxygenic photosynthesis from prokaryotes to plastids. Genome Biol. Evol. 5, 31–44 (2013).
Sánchez-Baracaldo, P., Raven, J. A., Pisani, D. & Knoll, A. H. Early photosynthetic eukaryotes inhabited low-salinity habitats. Proc. Natl Acad. Sci. USA 114, E7737–E7745 (2017).
Ponce-Toledo, R. I. et al. An early-branching freshwater cyanobacterium at the origin of plastids. Curr. Biol. 27, 386–391 (2017).
Amunts, A. & Nelson, N. Plant photosystem I design in the light of evolution. Structure 17, 637–650 (2009).
Alboresi, A. et al. Conservation of core complex subunits shaped the structure and function of photosystem I in the secondary endosymbiont alga Nannochloropsis gaditana. New Phytol. 213, 714–726 (2017).
Watanabe, M., Iwai, M., Narikawa, R. & Ikeuchi, M. Is the photosystem II complex a monomer or a dimer? Plant Cell Physiol. 50, 1674–1680 (2009).
Iwamura, T., Nagai, H. & Ichimura, S.-E. Improved methods for determining contents of chlorophyll, protein, bibonucleic acid, and deoxyribonucleic acid in planktonic populations. Int. Rev. ges. Hydrobiol. Hydrogr. 55, 131–147 (1970).
Schägger, H. & von Jagow, G. Blue native electrophoresis for isolation of membrane protein complexes in enzymatically active form. Anal. Biochem. 199, 223–231 (1991).
Wittig, I., Braun, H.-P. & Schagger, H. Blue native PAGE. Nat. Protoc. 1, 418–428 (2006).
Kolaskar, A. S. & Tongaonkar, P. C. A semi-empirical method for prediction of antigenic determinants on protein antigens. FEBS Lett. 276, 172–174 (1990).
Saha, S. & Raghava, G. P. S. Prediction of continuous B-cell epitopes in an antigen using recurrent neural network. Proteins 65, 40–48 (2006).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Wattam, A. R. et al. PATRIC, the bacterial bioinformatics database and analysis resource. Nucleic Acids Res. 42, D581–D591 (2014).
Overbeek, R. et al. The SEED and the Rapid Annotation of microbial genomes using Subsystems Technology (RAST). Nucleic Acids Res. 42, D206–D214 (2014).
Calteau, A. et al. Phylum-wide comparative genomics unravel the diversity of secondary metabolism in cyanobacteria. BMC Genomics 15, 977 (2014).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Criscuolo, A. & Gribaldo, S. BMGE (Block Mapping and Gathering with Entropy): a new software for selection of phylogenetic informative regions from multiple sequence alignments. BMC Evol. Biol. 10, 210 (2010).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. ProtTest 3: Fast selection of best-fit models of protein evolution. Bioinformatics 27, 1164–1165 (2011).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v3: An online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 44, W242–W245 (2016).
Vallenet, D. et al. MicroScope in 2017: An expanding and evolving integrated resource for community expertise of microbial genomes. Nucleic Acids Res. 45, D517–D528 (2017).
Schirrmeister, B. E., Gugger, M. & Donoghue, P. C. Cyanobacteria and the Great Oxidation Event: evidence from genes and fossils. Palaeontology 58, 769–785 (2015).
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: A sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Acknowledgements
Support to B.D.B., M.L. and J.T.N. was provided by the Gibson Family Foundation, the Bredesen Center for Interdisciplinary Research and Education, the Tennessee Plant Research Center, a UTK Professional Development Award, the Dr. Donald L. Akers Faculty Enrichment Fellowship to B.D.B. and National Science Foundation support to B.D.B. (DGE-0801470 and EPS-1004083). M.L. has been supported as a CIRE Fellow at University of Tennessee, Knoxville. A Professional Development Award from the Graduate School at UTK supported travel of B.D.B. to the Netherlands and to the Institut Pasteur. NWO Chemical Sciences supported work at University of Groningen. J.P.W. has been supported by NIH P30 DK063491. The Institut Pasteur supported Pasteur Culture Collection of cyanobacteria. We thank Y. I. Park for the use of the cyanobacterial genome of PCC 7124, and N.G. Brady and T. Cardona for helpful comments on the manuscript.
Author information
Authors and Affiliations
Contributions
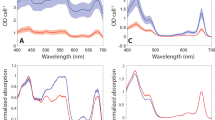
M.L., M.G. and B.D.B. designed the research. M.L. carried out most of the biochemistry and molecular biology experiments. A.C. performed phylogenetic and bioinformatics analyses. D.A.S. and E.J.B. did the electron microscopy imaging and single-particle analyses. T.A.W. did most of the psaL cloning. N.S. and J.T.N. prepared most of the cell materials. J.T.N. carried out the spectral comparison of PCC 7414 PSI oligomers J.W. performed the proteomic analyses. M.L. and B.D.B. wrote the article and all other authors contributed in editing and revising the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Conrad Mullineaux, Ann Magnuson and the other, anonymous, reviewer for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Supplementary Tables 1 and 2.
Supplementary Data 1
PSI oligomeric states in cyanobacteria studied, Cyanobacteria culture conditions and genome sequences used in this study.
Rights and permissions
About this article
Cite this article
Li, M., Calteau, A., Semchonok, D.A. et al. Physiological and evolutionary implications of tetrameric photosystem I in cyanobacteria. Nat. Plants 5, 1309–1319 (2019). https://doi.org/10.1038/s41477-019-0566-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-019-0566-x
This article is cited by
-
Structure of a monomeric photosystem I core associated with iron-stress-induced-A proteins from Anabaena sp. PCC 7120
Nature Communications (2023)
-
Isolation and characterization of trimeric and monomeric PSI cores from Acaryochloris marina MBIC11017
Photosynthesis Research (2023)
-
Structure of a tetrameric photosystem I from a glaucophyte alga Cyanophora paradoxa
Nature Communications (2022)
-
Cyclophilin anaCyp40 regulates photosystem assembly and phycobilisome association in a cyanobacterium
Nature Communications (2022)
-
Over-expression of an electron transport protein OmcS provides sufficient NADH for d-lactate production in cyanobacterium
Biotechnology for Biofuels (2021)