Abstract
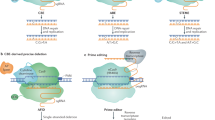
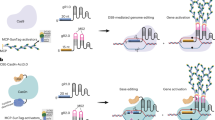
The application of clustered regularly interspaced short palindromic repeats (CRISPR) for genetic manipulation has revolutionized life science over the past few years. CRISPR was first discovered as an adaptive immune system in bacteria and archaea, and then engineered to generate targeted DNA breaks in living cells and organisms. During the cellular DNA repair process, various DNA changes can be introduced. The diverse and expanding CRISPR toolbox allows programmable genome editing, epigenome editing and transcriptome regulation in plants. However, challenges in plant genome editing need to be fully appreciated and solutions explored. This Review intends to provide an informative summary of the latest developments and breakthroughs of CRISPR technology, with a focus on achievements and potential utility in plant biology. Ultimately, CRISPR will not only facilitate basic research, but also accelerate plant breeding and germplasm development. The application of CRISPR to improve germplasm is particularly important in the context of global climate change as well as in the face of current agricultural, environmental and ecological challenges.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bibikova, M., Golic, M., Golic, K. G. & Carroll, D. Targeted chromosomal cleavage and mutagenesis in Drosophila using zinc-finger nucleases. Genetics 161, 1169–1175 (2002).
Christian, M. et al. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186, 757–761 (2010).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Li, J.-F. et al. Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat. Biotechnol. 31, 688–691 (2013).
Nekrasov, V., Staskawicz, B., Weigel, D., Jones, J. D. G. & Kamoun, S. Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat. Biotechnol. 31, 691–693 (2013).
Shan, Q. et al. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotechnol. 31, 686–688 (2013).
Chen, K., Wang, Y., Zhang, R., Zhang, H. & Gao, C. CRISPR/Cas genome editing and precision plant breeding in agriculture. Annu. Rev. Plant Biol. 70, 667–697 (2019).
Zhang, Y., Massel, K., Godwin, I. D. & Gao, C. Applications and potential of genome editing in crop improvement. Genome Biol. 19, 210 (2018).
Ma, X., Zhu, Q., Chen, Y. & Liu, Y.-G. CRISPR/Cas9 platforms for genome editing in plants: developments and applications. Mol. Plant 9, 961–974 (2016).
Schindele, P., Wolter, F. & Puchta, H. Transforming plant biology and breeding with CRISPR/Cas9, Cas12 and Cas13. FEBS Lett. 592, 1954–1967 (2018).
Yin, K., Gao, C. & Qiu, J.-L. Progress and prospects in plant genome editing. Nat. Plants 3, 17107 (2017).
Kim, J.-S. Precision genome engineering through adenine and cytosine base editing. Nat. Plants 4, 148–151 (2018).
Huang, T. K. & Puchta, H. CRISPR/Cas-mediated gene targeting in plants: finally a turn for the better for homologous recombination. Plant Cell Rep. 38, 443–453 (2019).
Mahas, A., Neal Stewart, C. & Mahfouz, M. M. Harnessing CRISPR/Cas systems for programmable transcriptional and post-transcriptional regulation. Biotechnol. Adv. 36, 295–310 (2018).
Makarova, K. S. et al. An updated evolutionary classification of CRISPR–Cas systems. Nat. Rev. Microbiol. 13, 722–736 (2015).
Burstein, D. et al. New CRISPR–Cas systems from uncultivated microbes. Nature 542, 237–241 (2017).
Harrington, L. B. et al. Programmed DNA destruction by miniature CRISPR-Cas14 enzymes. Science 362, 839–842 (2018).
Liu, J.-J. et al. CasX enzymes comprise a distinct family of RNA-guided genome editors. Nature 566, 218 (2019).
Shmakov, S. et al. Discovery and functional characterization of diverse Class 2 CRISPR-Cas systems. Mol. Cell 60, 385–397 (2015).
Shmakov, S. et al. Diversity and evolution of class 2 CRISPR–Cas systems. Nat. Rev. Microbiol. 15, 169–182 (2017).
Yan, W. X. et al. Functionally diverse type V CRISPR-Cas systems. Science 363, 88–91 (2019).
Ma, X. et al. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol. Plant 8, 1274–1284 (2015).
Fauser, F., Schiml, S. & Puchta, H. Both CRISPR/Cas-based nucleases and nickases can be used efficiently for genome engineering in Arabidopsis thaliana. Plant J. 79, 348–359 (2014).
Wang, Z.-P. et al. Egg cell-specific promoter-controlled CRISPR/Cas9 efficiently generates homozygous mutants for multiple target genes in Arabidopsis in a single generation. Genome Biol. 16, 144 (2015).
Lee, K. et al. Activities and specificities of CRISPR/Cas9 and Cas12a nucleases for targeted mutagenesis in maize. Plant Biotechnol. J. 17, 839–842 (2019).
Michno, J.-M. et al. CRISPR/Cas mutagenesis of soybean and Medicago truncatula using a new web-tool and a modified Cas9 enzyme. GM Crops Food 6, 243–252 (2015).
Mali, P. et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 31, 833–838 (2013).
Ran, F. A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Kleinstiver, B. P. et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015).
Anders, C., Bargsten, K. & Jinek, M. Structural plasticity of PAM recognition by engineered variants of the RNA-guided endonuclease Cas9. Mol. Cell 61, 895–902 (2016).
Hu, J. H. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57–63 (2018).
Nishimasu, H. et al. Engineered CRISPR-Cas9 nuclease with expanded targeting space. Science 361, 1259–1262 (2018).
Hu, X. et al. Expanding the range of CRISPR/Cas9 genome editing in rice. Mol. Plant 9, 943–945 (2016).
Hu, X., Meng, X., Liu, Q., Li, J. & Wang, K. Increasing the efficiency of CRISPR-Cas9-VQR precise genome editing in rice. Plant Biotechnol. J. 16, 292–297 (2017).
Wang, J. et al. xCas9 expands the scope of genome editing with reduced efficiency in rice. Plant Biotechnol. J. 17, 709–711 (2019).
Hua, K., Tao, X., Han, P., Wang, R. & Zhu, J.-K. Genome engineering in rice using Cas9 variants that recognize NG PAM sequences. Mol. Plant https://doi.org/10.1016/j.molp.2019.03.009 (2019).
Zhong, Z. et al. Improving plant genome editing with high-fidelity xCas9 and non-canonical PAM-targeting Cas9-NG. Mol. Plant https://doi.org/10.1016/j.molp.2019.03.011 (2019).
Li, J. et al. Plant genome editing using xCas9 with expanded PAM compatibility. J. Genet. Genomics https://doi.org/10.1016/j.jgg.2019.03.004 (2019).
Endo, M. et al. Genome editing in plants by engineered CRISPR–Cas9 recognizing NG PAM. Nat. Plants 5, 14–17 (2018).
Ren, B. et al. Cas9-NG greatly expands the targeting scope of genome-editing toolkit by recognizing NG and other atypical PAMs in rice. Mol. Plant https://doi.org/10.1016/j.molp.2019.03.010 (2019).
Negishi, K. et al. An adenine base editor with expanded targeting scope using SpCas9-NGv1 in rice. Plant Biotechnol. J. https://doi.org/10.1111/pbi.13120 (2019).
Wang, M. et al. Optimizing baseeditors for improved efficiency and expanded editing scope in rice. Plant Biotechnol. J. https://doi.org/10.1111/pbi.13124 (2019).
Ge, Z. et al. Engineered xCas9 and SpCas9-NG variants broaden PAM recognition sites to generate mutations in Arabidopsis plants. Plant Biotechnol. J. https://doi.org/10.1111/pbi.13148 (2019).
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015).
Friedland, A. E. et al. Characterization of Staphylococcus aureus Cas9: a smaller Cas9 for all-in-one adeno-associated virus delivery and paired nickase applications. Genome Biol. 16, 257 (2015).
Kleinstiver, B. P. et al. Broadening the targeting range of Staphylococcus aureus CRISPR-Cas9 by modifying PAM recognition. Nat. Biotechnol. 33, 1293–1298 (2015).
Kaya, H., Mikami, M., Endo, A., Endo, M. & Toki, S. Highly specific targeted mutagenesis in plants using Staphylococcus aureus Cas9. Sci. Rep. 6, 26871 (2016).
Steinert, J., Schiml, S., Fauser, F. & Puchta, H. Highly efficient heritable plant genome engineering using Cas9 orthologues from Streptococcus thermophilus and Staphylococcus aureus. Plant J. 84, 1295–1305 (2015).
Jia, H., Xu, J., Orbović, V., Zhang, Y. & Wang, N. Editing Citrus genome via SaCas9/sgRNA system. Front. Plant Sci. 8, 2135 (2017).
Müller, M. et al. Streptococcus thermophilus CRISPR-Cas9 systems enable specific editing of the human genome. Mol. Ther. 24, 636–644 (2016).
Lee, C. M., Cradick, T. J. & Bao, G. The Neisseria meningitidis CRISPR-Cas9 system enables specific genome editing in mammalian cells. Mol. Ther. 24, 645–654 (2016).
Hirano, H. et al. Structure and engineering of Francisella novicida Cas9. Cell 164, 950–961 (2016).
Esvelt, K. M. et al. Orthogonal Cas9 proteins for RNA-guided gene regulation and editing. Nat. Methods 10, 1116–21 (2013).
Kim, E. et al. In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni. Nat. Commun. 8, 14500 (2017).
Chatterjee, P., Jakimo, N. & Jacobson, J. M. Minimal PAM specificity of a highly similar SpCas9 ortholog. Sci. Adv. 4, eaau0766 (2018).
Karvelis, T. et al. Rapid characterization of CRISPR-Cas9 protospacer adjacent motif sequence elements. Genome Biol. 16, 253 (2015).
Rousseau, B. A., Hou, Z., Gramelspacher, M. J. & Zhang, Y. Programmable RNA cleavage and recognition by a natural CRISPR-Cas9 system from Neisseria meningitidis. Mol. Cell 69, 906–914 (2018).
Jakimo, N., Chatterjee, P., Nip, L. & Jacobson, J. M. A Cas9 with complete PAM recognition for adenine dinucleotides. Preprint at https://www.biorxiv.org/node/129129.abstract (2018).
Zetsche, B. et al. Cpf1 Is a single RNA-guided endonuclease of a Class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Yamano, T. et al. Crystal structure of Cpf1 in complex with guide RNA and target DNA. Cell 165, 949–962 (2016).
Kim, D. et al. Genome-wide analysis reveals specificities of Cpf1 endonucleases in human cells. Nat. Biotechnol. 34, 863–868 (2016).
Kleinstiver, B. P. et al. Genome-wide specificities of CRISPR-Cas Cpf1 nucleases in human cells. Nat. Biotechnol. 34, 869–874 (2016).
Tang, X. et al. A CRISPR–Cpf1 system for efficient genome editing and transcriptional repression in plants. Nat. Plants 3, 17103 (2017).
Zhong, Z. et al. Plant genome editing using FnCpf1 and LbCpf1 nucleases at redefined and altered PAM sites. Mol. Plant 11, 999–1002 (2018).
Endo, A., Masafumi, M., Kaya, H. & Toki, S. Efficient targeted mutagenesis of rice and tobacco genomes using Cpf1 from Francisella novicida. Sci. Rep. 6, 38169 (2016).
Hu, X., Wang, C., Liu, Q., Fu, Y. & Wang, K. Targeted mutagenesis in rice using CRISPR-Cpf1 system. J. Genet. Genom. 44, 71–73 (2017).
Wang, M., Mao, Y., Lu, Y., Tao, X. & Zhu, J. Multiplex gene editing in rice using the CRISPR-Cpf1 system. Mol. Plant 10, 1011–1013 (2017).
Yin, X. et al. CRISPR-Cas9 and CRISPR-Cpf1 mediated targeting of a stomatal developmental gene EPFL9 in rice. Plant Cell Rep. 36, 745–757 (2017).
Begemann, M. B. et al. Precise insertion and guided editing of higher plant genomes using Cpf1 CRISPR nucleases. Sci. Rep. 7, 11606 (2017).
Bernabé‐Orts, J. M. et al. Assessment of Cas12a-mediated gene editing efficiency in plants. Plant Biotechnol. J. https://doi.org/10.1111/pbi.13113 (2019).
Kim, H. et al. CRISPR/Cpf1-mediated DNA-free plant genome editing. Nat. Commun. 8, 14406 (2017).
Li, B. et al. Robust CRISPR/Cpf1(Cas12a) mediated genome editing in allotetraploid cotton (G. hirsutum). Plant Biotechnol. J. https://doi.org/10.1111/pbi.13147 (2019).
Jia, H., Orbović, V. & Wang, N. CRISPR-LbCas12a-mediated modification of citrus. Plant Biotechnol. J. https://doi.org/10.1111/pbi.13109 (2019).
Zhang, X. et al. Multiplex gene regulation by CRISPR-ddCpf1. Cell Discov. 3, 17018 (2017).
Gao, L. et al. Engineered Cpf1 variants with altered PAM specificities. Nat. Biotechnol. 35, 789 (2017).
Tóth, E. et al. Mb- and FnCpf1 nucleases are active in mammalian cells: activities and PAM preferences of four wild-type Cpf1 nucleases and of their altered PAM specificity variants. Nucleic Acids Res. 46, 10272–10285 (2018).
Li, S. et al. Expanding the scope of CRISPR/Cpf1-mediated genome editing in rice. Mol. Plant 11, 995–998 (2018).
Kleinstiver, B. P. et al. Engineered CRISPR–Cas12a variants with increased activities and improved targeting ranges for gene, epigenetic and base editing. Nat. Biotechnol. 37, 276 (2019).
Zetsche, B. et al. A survey of genome editing activity for 16 Cpf1 orthologs. Preprint at https://www.biorxiv.org/content/10.1101/134015v1.article-info (2017).
Teng, F. et al. Enhanced mammalian genome editing by new Cas12a orthologs with optimized crRNA scaffolds. Genome Biol. 20, 15 (2019).
Liu, L. et al. C2c1-sgRNA complex structure reveals RNA-guided DNA cleavage mechanism. Mol. Cell 65, 310–322 (2017).
Yang, H., Gao, P., Rajashankar, K. R. & Patel, D. J. PAM-dependent target DNA recognition and cleavage by C2c1 CRISPR-Cas endonuclease. Cell 167, 1814–1828 (2016).
Teng, F. et al. Repurposing CRISPR-Cas12b for mammalian genome engineering. Cell Discov. 4, 63 (2018).
Strecker, J. et al. Engineering of CRISPR-Cas12b for human genome editing. Nat. Commun. 10, 212 (2019).
Dolan, A. E. et al. Introducing a spectrum of long-range genomic deletions in human embryonic stem cells using Type I CRISPR-Cas. Mol. Cell 74, 1–15 (2019).
Tanenbaum, M. E., Gilbert, L. A., Qi, L. S., Weissman, J. S. & Vale, R. D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159, 635–646 (2014).
Leonetti, M. D., Sekine, S., Kamiyama, D., Weissman, J. S. & Huang, B. A scalable strategy for high-throughput GFP tagging of endogenous human proteins. Proc. Natl Acad. Sci. USA 113, E3501–E3508 (2016).
Konermann, S. et al. Genome-scale transcriptional activation by an engineered CRISPR-Cas9 complex. Nature 517, 583–588 (2015).
Lowder, L. G. et al. Robust transcriptional activation in plants using multiplexed CRISPR-Act2.0 and mTALE-Act systems. Mol. Plant 11, 245–256 (2018).
Shao, S. et al. Long-term dual-color tracking of genomic loci by modified sgRNAs of the CRISPR/Cas9 system. Nucleic Acids Res. 44, e86 (2016).
Shechner, D. M., Hacisuleyman, E., Younger, S. T. & Rinn, J. L. Multiplexable, locus-specific targeting of long RNAs with CRISPR-Display. Nat. Methods 12, 664–670 (2015).
Zalatan, J. G. et al. Engineering complex synthetic transcriptional programs with CRISPR RNA scaffolds. Cell 160, 339–350 (2015).
Ma, H. et al. Multiplexed labeling of genomic loci with dCas9 and engineered sgRNAs using CRISPRainbow. Nat. Biotechnol. 34, 528–530 (2016).
Carlson-Stevermer, J. et al. Assembly of CRISPR ribonucleoproteins with biotinylated oligonucleotides via an RNA aptamer for precise gene editing. Nat. Commun. 8, 1711 (2017).
Qin, P. et al. Live cell imaging of low- and non-repetitive chromosome loci using CRISPR-Cas9. Nat. Commun. 8, 14725 (2017).
Ma, H. et al. CRISPR-Sirius: RNA scaffolds for signal amplification in genome imaging. Nat. Methods 15, 928 (2018).
Kunii, A. et al. Three-component repurposed technology for enhanced expression: highly accumulable transcriptional activators via branched tag arrays. CRISPR J. 1, 337–347 (2018).
Abudayyeh, O. O. et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 353, aaf5573 (2016).
Abudayyeh, O. O. et al. RNA targeting with CRISPR–Cas13. Nature 550, 280–284 (2017).
Cox, D. B. T. et al. RNA editing with CRISPR-Cas13. Science 358, 1019–1027 (2017).
Gootenberg, J. S. et al. Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 356, 438–442 (2017).
Aman, R. et al. RNA virus interference via CRISPR/Cas13a system in plants. Genome Biol. 19, 1 (2018).
Aman, R. et al. Engineering RNA virus interference via the CRISPR/Cas13 machinery in Arabidopsis. Viruses 10, 732 (2018).
Lowder, L., Malzahn, A. & Qi, Y. Rapid evolution of manifold CRISPR systems for plant genome editing. Front. Plant Sci. 7, 1683 (2016).
Gao, Y. & Zhao, Y. Self-processing of ribozyme-flanked RNAs into guide RNAs in vitro and in vivo for CRISPR-mediated genome editing. J. Integr. Plant Biol. 56, 343–349 (2014).
Gao, Y. et al. Auxin binding protein 1 (ABP1) is not required for either auxin signaling or Arabidopsis development. Proc. Natl Acad. Sci. USA 112, 2275–2280 (2015).
Tang, X. et al. A single transcript CRISPR-Cas9 system for efficient genome editing in plants. Mol. Plant 9, 1088–1091 (2016).
Tang, X. et al. Single transcript unit CRISPR 2.0 systems for robust Cas9 and Cas12a mediated plant genome editing. Plant Biotechnol. J. https://doi.org/10.1111/pbi.13068 (2018).
Wang, M. et al. Multiplex gene editing in rice with simplified CRISPR-Cpf1 and CRISPR-Cas9 systems: simplified single transcriptional unit CRISPR systems. J. Integr. Plant Biol. 60, 626–631 (2018).
Yoshioka, S., Fujii, W., Ogawa, T., Sugiura, K. & Naito, K. Development of a mono-promoter-driven CRISPR/Cas9 system in mammalian cells. Sci. Rep. 5, 18341 (2015).
Ding, D., Chen, K., Chen, Y., Li, H. & Xie, K. Engineering introns to express RNA guides for Cas9- and Cpf1-mediated multiplex genome editing. Mol. Plant 11, 542–552 (2018).
Lowder, L. G. et al. A CRISPR/Cas9 toolbox for multiplexed plant genome editing and transcriptional regulation. Plant Physiol. 169, 971–985 (2015).
Vazquez-Vilar, M. et al. A modular toolbox for gRNA–Cas9 genome engineering in plants based on the GoldenBraid standard. Plant Methods 12, 10 (2016).
Xing, H.-L. et al. A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol. 14, 327 (2014).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mikami, M., Toki, S. & Endo, M. In Planta processing of the SpCas9–gRNA complex. Plant Cell Physiol. 58, 1857–1867 (2017).
Li, S. et al. Synthesis-dependent repair of Cpf1-induced double strand DNA breaks enables targeted gene replacement in rice. J. Exp. Bot. 69, 4715–4721 (2018).
Xie, K., Minkenberg, B. & Yang, Y. Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc. Natl Acad. Sci. USA 112, 3570–3575 (2015).
Wang, W., Akhunova, A., Chao, S. & Akhunov, E. Optimizing multiplex CRISPR/Cas9-based genome editing for wheat. Preprint at https://www.biorxiv.org/content/10.1101/051342v1 (2016).
Qi, W. et al. High-efficiency CRISPR/Cas9 multiplex gene editing using the glycine tRNA-processing system-based strategy in maize. BMC Biotechnol. 16, 58 (2016).
Tsai, S. Q. et al. Dimeric CRISPR RNA-guided FokI nucleases for highly specific genome editing. Nat. Biotechnol. 32, 569–576 (2014).
Yan, Q. et al. Multiplex CRISPR/Cas9-based genome engineering enhanced by Drosha-mediated sgRNA-shRNA structure. Sci. Rep. 6, 38970 (2016).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Nishida, K. et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science 353, aaf8729 (2016).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464 (2017).
Conticello, S. G. The AID/APOBEC family of nucleic acid mutators. Genome Biol. 9, 229 (2008).
Kunz, C., Saito, Y. & Schär, P. DNA Repair in mammalian cells: Mismatched repair: variations on a theme. Cell. Mol. Life Sci. CMLS 66, 1021–1038 (2009).
Mol, C. D. et al. Crystal structure of human uracil-DNA glycosylase in complex with a protein inhibitor: Protein mimicry of DNA. Cell 82, 701–708 (1995).
Komor, A. C. et al. Improved base excision repair inhibition and bacteriophage Mu Gam protein yields C:G-to-T: Abase editors with higher efficiency and product purity. Sci. Adv. 3, eaao4774 (2017).
Wang, L. et al. Enhanced base editing by co-expression of free uracil DNA glycosylase inhibitor. Cell Res. 27, 1289–1292 (2017).
Zafra, M. P. et al. Optimized base editors enable efficient editing in cells, organoids and mice. Nat. Biotechnol. 36, 888–893 (2018).
Kuscu, C. & Adli, M. CRISPR-Cas9-AID base editor is a powerful gain-of-function screening tool. Nat. Methods 13, 983–984 (2016).
Gehrke, J. M. et al. An APOBEC3A-Cas9 base editor with minimized bystander and off-target activities. Nat. Biotechnol. 36, 977–982 (2018).
Wang, X. et al. Efficient base editing in methylated regions with a human APOBEC3A-Cas9 fusion. Nat. Biotechnol. 36, 946–949 (2018).
Rees, H. A. & Liu, D. R. Base editing: precision chemistry on the genome and transcriptome of living cells. Nat. Rev. Genet. 19, 770 (2018).
Molla, K. A. & Yang, Y. CRISPR/Cas-mediated base editing: technical considerations and practical applications. Trends Biotechnol. https://doi.org/10.1016/j.tibtech.2019.03.008 (2019).
Hua, K., Tao, X. & Zhu, J.-K. Expanding the base editing scope in rice by using Cas9 variants. Plant Biotechnol. J. 17, 499–504 (2019).
Li, J., Sun, Y., Du, J., Zhao, Y. & Xia, L. Generation of targeted point mutations in rice by a modified CRISPR/Cas9 system. Mol. Plant 10, 526–529 (2017).
Lu, Y. & Zhu, J.-K. Precise editing of a target base in the rice genome using a modified CRISPR/Cas9 system. Mol. Plant 10, 523–525 (2017).
Ren, B. et al. A CRISPR/Cas9 toolkit for efficient targeted base editing to induce genetic variations in rice. Sci. China Life Sci. 60, 516–519 (2017).
Zong, Y. et al. Precise base editing in rice, wheat and maize with a Cas9-cytidine deaminase fusion. Nat. Biotechnol. 35, 438–440 (2017).
Zong, Y. et al. Efficient C-to-T base editing in plants using a fusion of nCas9 and human APOBEC3A. Nat. Biotechnol. 36, 950–953 (2018).
Ren, B. et al. Improved base editor for efficiently inducing genetic variations in rice with CRISPR/Cas9-guided hyperactive hAID mutant. Mol. Plant 11, 623–626 (2018).
Shimatani, Z. et al. Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion. Nat. Biotechnol. 35, 441–443 (2017).
Yan, F. et al. Highly efficient A·T to G·C base editing by Cas9n-guided tRNA adenosine deaminase in rice. Mol. Plant 11, 631–634 (2018).
Kang, B.-C. et al. Precision genome engineering through adenine base editing in plants. Nat. Plants 4, 427–431 (2018).
Matthews, M. M. et al. Structures of human ADAR2 bound to dsRNA reveal base-flipping mechanism and basis for site selectivity. Nat. Struct. Mol. Biol. 23, 426–433 (2016).
Montiel-Gonzalez, M. F., Vallecillo-Viejo, I., Yudowski, G. A. & Rosenthal, J. J. C. Correction of mutations within the cystic fibrosis transmembrane conductance regulator by site-directed RNA editing. Proc. Natl Acad. Sci. USA 110, 18285–18290 (2013).
Jin, S. et al. Cytosine, but not adenine, base editors induce genome-wide off-target mutations in rice. Science 364, 292–295 (2019).
Gajula, K. S. Designing an elusive C•G→G•C CRISPR base editor. Trends Biochem. Sci. 44, 91–94 (2019).
Čermák, T., Baltes, N. J., Čegan, R., Zhang, Y. & Voytas, D. F. High-frequency, precise modification of the tomato genome. Genome Biol. 16, 232 (2015).
Sun, Y. et al. Engineering herbicide-resistant rice plants through CRISPR/Cas9-mediated homologous recombination of Acetolactate Synthase. Mol. Plant 9, 628–631 (2016).
Malzahn, A., Lowder, L. & Qi, Y. Plant genome editing with TALEN and CRISPR. Cell Biosci. 7, 21 (2017).
Li, H. et al. Design and specificity of long ssDNA donors for CRISPR-based knock-in. Preprint at https://www.biorxiv.org/content/10.1101/178905v1 (2017).
Richardson, C. D., Ray, G. J., DeWitt, M. A., Curie, G. L. & Corn, J. E. Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-Cas9 using asymmetric donor DNA. Nat. Biotechnol. 34, 339–344 (2016).
Baker, O. et al. The contribution of homology arms to nuclease-assisted genome engineering. Nucleic Acids Res. 45, 8105–8115 (2017).
Fauser, F. et al. In planta gene targeting. Proc. Natl Acad. Sci. USA 109, 7535–7540 (2012).
Schiml, S., Fauser, F. & Puchta, H. The CRISPR/Cas system can be used as nuclease for in planta gene targeting and as paired nickases for directed mutagenesis in Arabidopsis resulting in heritable progeny. Plant J. 80, 1139–1150 (2014).
Zhang, J.-P. et al. Efficient precise knockin with a double cut HDR donor after CRISPR/Cas9-mediated double-stranded DNA cleavage. Genome Biol. 18, 35 (2017).
Li, J. et al. Efficient allelic replacement in rice by gene editing: A case study of the NRT1.1B gene. J. Integr. Plant Biol. 60, 536–540 (2018).
Baltes, N. J., Gil-Humanes, J., Cermak, T., Atkins, P. A. & Voytas, D. F. DNA replicons for plant genome engineering. Plant Cell 26, 151–163 (2014).
Dahan‐Meir, T. et al. Efficient in planta gene targeting in tomato using geminiviral replicons and the CRISPR/Cas9 system. Plant J. 95, 5–16 (2018).
Wang, M. et al. Gene targeting by homology-directed repair in rice using a geminivirus-based CRISPR/Cas9 system. Mol. Plant 10, 1007–1010 (2017).
Cunningham, F. J., Goh, N. S., Demirer, G. S., Matos, J. L. & Landry, M. P. Nanoparticle-mediated delivery towards advancing plant genetic engineering. Trends Biotechnol. 36, 882–897 (2018).
Demirer, G. S. et al. High aspect ratio nanomaterials enable delivery of functional genetic material without DNA integration in mature plants. Nat. Nanotechnol. 14, 456–464 (2019).
Ghanta, K. S. et al. 5′ Modifications improve potency and efficacy of DNA donors for precision genome editing. Preprint at https://www.biorxiv.org/content/10.1101/354480v1.full (2018).
Ma, M. et al. Efficient generation of mice carrying homozygous double-floxp alleles using the Cas9-Avidin/Biotin-donor DNA system. Cell Res. 27, 578–581 (2017).
Aird, E. J., Lovendahl, K. N., Martin, A. S., Harris, R. S. & Gordon, W. R. Increasing Cas9-mediated homology-directed repair efficiency through covalent tethering of DNA repair template. Commun. Biol. 1, 54 (2018).
Keskin, H. et al. Transcript-RNA-templated DNA recombination and repair. Nature 515, 436–439 (2014).
Li, S. et al. Precise gene replacement in rice by RNA transcript-templated homologous recombination. Nat. Biotechnol. 37, 445–450 (2019).
Jayathilaka, K. et al. A chemical compound that stimulates the human homologous recombination protein RAD51. Proc. Natl Acad. Sci. USA 105, 15848–15853 (2008).
Song, J. et al. RS-1 enhances CRISPR/Cas9- and TALEN-mediated knock-in efficiency. Nat. Commun. 7, 10548 (2016).
Schuermann, D., Molinier, J., Fritsch, O. & Hohn, B. The dual nature of homologous recombination in plants. Trends Genet. 21, 172–181 (2005).
Shaked, H., Melamed-Bessudo, C. & Levy, A. A. High-frequency gene targeting in Arabidopsis plants expressing the yeast RAD54 gene. Proc. Natl Acad. Sci. USA 102, 12265–12269 (2005).
Chen, X. et al. In trans paired nicking triggers seamless genome editing without double-stranded DNA cutting. Nat. Commun. 8, 657 (2017).
Chu, V. T. et al. Increasing the efficiency of homology-directed repair for CRISPR-Cas9-induced precise gene editing in mammalian cells. Nat. Biotechnol. 33, 543–548 (2015).
Maruyama, T. et al. Increasing the efficiency of precise genome editing with CRISPR-Cas9 by inhibition of nonhomologous end joining. Nat. Biotechnol. 33, 538–542 (2015).
Robert, F., Barbeau, M., Éthier, S., Dostie, J. & Pelletier, J. Pharmacological inhibition of DNA-PK stimulates Cas9-mediated genome editing. Genome Med. 7, 93 (2015).
Qi, Y. et al. Increasing frequencies of site-specific mutagenesis and gene targeting in Arabidopsis by manipulating DNA repair pathways. Genome Res. 23, 547–554 (2013).
Endo, M., Mikami, M. & Toki, S. Biallelic gene targeting in rice. Plant Physiol. 170, 667–677 (2016).
Rodríguez-Leal, D., Lemmon, Z. H., Man, J., Bartlett, M. E. & Lippman, Z. B. Engineering quantitative trait variation for crop improvement by genome editing. Cell 171, 470–480 (2017).
Zhang, H. et al. Genome editing of upstream open reading frames enables translational control in plants. Nat. Biotechnol. 36, 894–898 (2018).
Li, Z., Xiong, X., Wang, F., Liang, J. & Li, J.-F. Gene disruption through base editing-induced messenger RNA missplicing in plants. New Phytol. 222, 1139–1148 (2019).
Xue, C., Zhang, H., Lin, Q., Fan, R. & Gao, C. Manipulating mRNA splicing by base editing in plants. Sci. China Life Sci. 61, 1293–1300 (2018).
Piatek, A. et al. RNA-guided transcriptional regulation in planta via synthetic dCas9-based transcription factors. Plant Biotechnol. J. 13, 578–589 (2015).
Gilbert, L. A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015).
Zhang, X. et al. Gene activation in human cells using CRISPR/Cpf1-p300 and CRISPR/Cpf1-SunTag systems. Protein Cell 9, 380–383 (2018).
Li, Z. et al. A potent Cas9-derived gene activator for plant and mammalian cells. Nat. Plants 3, 930–936 (2017).
Papikian, A., Liu, W., Gallego-Bartolomé, J. & Jacobsen, S. E. Site-specific manipulation of Arabidopsis loci using CRISPR-Cas9 SunTag systems. Nat. Commun. 10, 729 (2019).
Gilbert, L. A. et al. Genome-scale CRISPR-mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Cheng, A. W. et al. Multiplexed activation of endogenous genes by CRISPR-on, an RNA-guided transcriptional activator system. Cell Res. 23, 1163–1171 (2013).
Farzadfard, F., Perli, S. D. & Lu, T. K. Tunable and multifunctional eukaryotic transcription factors based on CRISPR/Cas. ACS Synth. Biol. 2, 604–613 (2013).
Deaner, M., Mejia, J. & Alper, H. S. Enabling graded and large-scale multiplex of desired genes using a dual-mode dCas9 activator in Saccharomyces cerevisiae. ACS Synth. Biol. 6, 1931–1943 (2017).
Kiani, S. et al. Cas9 gRNA engineering for genome editing, activation and repression. Nat. Methods 12, 1051–1054 (2015).
Breinig, M. et al. Multiplexed orthogonal genome editing and transcriptional activation by Cas12a. Nat. Methods 16, 51–54 (2019).
Braun, S. M. G. et al. Rapid and reversible epigenome editing by endogenous chromatin regulators. Nat. Commun. 8, 560 (2017).
Huang, Y.-H. et al. DNA epigenome editing using CRISPR-Cas SunTag-directed DNMT3A. Genome Biol. 18, 176 (2017).
Liao, H.-K. et al. In vivo target gene activation via CRISPR/Cas9-mediated trans-epigenetic modulation. Cell 171, 1495–1507 (2017).
Morita, S. et al. Targeted DNA demethylation in vivo using dCas9–peptide repeat and scFv–TET1 catalytic domain fusions. Nat. Biotechnol. 34, 1060–1065 (2016).
Gallego-Bartolomé, J. et al. Targeted DNA demethylation of the Arabidopsis genome using the human TET1 catalytic domain. Proc. Natl Acad. Sci. USA 115, E2125–E2134 (2018).
Galonska, C. et al. Genome-wide tracking of dCas9-methyltransferase footprints. Nat. Commun. 9, 597 (2018).
O’Connell, M. R. et al. Programmable RNA recognition and cleavage by CRISPR/Cas9. Nature 516, 263–266 (2014).
Batra, R. et al. Elimination of toxic microsatellite repeat expansion RNA by RNA-targeting Cas9. Cell 170, 899–912 (2017).
Xiang, G., Zhang, X., An, C., Cheng, C. & Wang, H. Temperature effect on CRISPR-Cas9 mediated genome editing. J. Genet. Genom. 44, 199–205 (2017).
Moreno-Mateos, M. A. et al. CRISPR-Cpf1 mediates efficient homology-directed repair and temperature-controlled genome editing. Nat. Commun. 8, 2024 (2017).
LeBlanc Chantal. et al. Increased efficiency of targeted mutagenesis by CRISPR/Cas9 in plants using heat stress. Plant J. 93, 377–386 (2018).
Malzahn, A. A. et al. Application of CRISPR-Cas12a temperature sensitivity for improved genome editing in rice, maize, and Arabidopsis. BMC Biol. 17, 9 (2019).
LeBlanc, C. et al. Increased efficiency of targeted mutagenesis by CRISPR/Cas9 in plants using heat stress. Plant J. 93, 377–386 (2018).
Gao, X., Chen, J., Dai, X., Zhang, D. & Zhao, Y. An effective strategy for reliably isolating heritable and Cas9-free Arabidopsis mutants generated by CRISPR/Cas9-mediated genome editing. Plant Physiol. 171, 1794–1800 (2016).
Lu, H.-P. et al. CRISPR-S: an active interference element for a rapid and inexpensive selection of genome-edited, transgene-free rice plants. Plant Biotechnol. J. 15, 1371–1373 (2017).
Iaffaldano, B., Zhang, Y. & Cornish, K. CRISPR/Cas9 genome editing of rubber producing dandelion Taraxacum kok-saghyz using Agrobacterium rhizogenes without selection. Ind. Crops Prod. 89, 356–362 (2016).
Zhang, Y. et al. Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA. Nat. Commun. 7, 12617 (2016).
Woo, J. W. et al. DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat. Biotechnol. 33, 1162–1164 (2015).
Malnoy, M. et al. DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front. Plant Sci. 7, 1904 (2016).
Subburaj, S. et al. Site-directed mutagenesis in Petunia × hybrida protoplast system using direct delivery of purified recombinant Cas9 ribonucleoproteins. Plant Cell Rep. 35, 1535–1544 (2016).
Li, J. et al. Whole genome sequencing reveals rare off-target mutations and considerable inherent genetic or/and somaclonal variations in CRISPR/Cas9-edited cotton plants. Plant Biotechnol. J. 17, 858–868 (2019).
Liang, Z. et al. Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes. Nat. Commun. 8, 14261 (2017).
Svitashev, S., Schwartz, C., Lenderts, B., Young, J. K. & Cigan, A. M. Genome editing in maize directed by CRISPR–Cas9 ribonucleoprotein complexes. Nat. Commun. 7, 13274 (2016).
Toda, E. et al. An efficient DNA- and selectable-marker-free genome-editing system using zygotes in rice. Nat. Plants 5, 363–368 (2019).
He, Y. et al. Programmed self-elimination of the CRISPR/Cas9 construct greatly accelerates the isolation of edited and transgene-free rice plants. Mol. Plant 11, 1210–1213 (2018).
Ryder, P., McHale, M., Fort, A. & Spillane, C. Generation of stable nulliplex autopolyploid lines of Arabidopsis thaliana using CRISPR/Cas9 genome editing. Plant Cell Rep. 36, 1005–1008 (2017).
Shan, S. et al. Application of CRISPR/Cas9 to Tragopogon (Asteraceae), an evolutionary model for the study of polyploidy. Mol. Ecol. Resour. 18, 1427–1443 (2018).
Andersson, M. et al. Efficient targeted multiallelic mutagenesis in tetraploid potato (Solanum tuberosum) by transient CRISPR-Cas9 expression in protoplasts. Plant Cell Rep. 36, 117–128 (2017).
Braatz, J. et al. CRISPR-Cas9 targeted mutagenesis leads to simultaneous modification of different homoeologous gene copies in polyploid oilseed Rape (Brassica napus). Plant Physiol. 174, 935–942 (2017).
Gao, W. et al. Genome editing in cotton with the CRISPR/Cas9 system. Front. Plant Sci. 8, 1364 (2017).
Jiang, W. Z. et al. Significant enhancement of fatty acid composition in seeds of the allohexaploid, Camelina sativa, using CRISPR/Cas9 gene editing. Plant Biotechnol. J. 15, 648–657 (2017).
Liu, Y. et al. Targeted mutagenesis in tetraploid switchgrass (Panicum virgatum L.) using CRISPR/Cas9. Plant Biotechnol. J. 16, 381–393 (2018).
Morineau, C. et al. Selective gene dosage by CRISPR-Cas9 genome editing in hexaploid Camelina sativa. Plant Biotechnol. J. 15, 729–739 (2017).
Wang, Y. et al. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 32, 947–951 (2014).
Zhang, Y. et al. Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA. Nat. Commun. 7, 12617 (2016).
Wang, W. et al. Transgenerational CRISPR-Cas9 activity facilitates multiplex gene editing in allopolyploid wheat. CRISPR J. 1, 65–74 (2018).
Zhang, Z. et al. Development of an Agrobacterium-delivered CRISPR/Cas9 system for wheat genome editing. Plant Biotechnol. J. https://doi.org/10.1111/pbi.13088 (2019).
Jia, H. et al. Genome editing of the disease susceptibility gene CsLOB1 in citrus confers resistance to citrus canker. Plant Biotechnol. J. 15, 817–823 (2017).
Fauser, F., Schiml, S. & Puchta, H. Both CRISPR/Cas-based nucleases and nickases can be used efficiently for genome engineering in Arabidopsis thaliana. Plant J. 79, 348–359 (2014).
Feng, Z. et al. Efficient genome editing in plants using a CRISPR/Cas system. Cell Res. 23, 1229–1232 (2013).
Shan, Q. et al. ZFN, TALEN and CRISPR-Cas9 mediated homology directed gene insertion in Arabidopsis: a disconnect between somatic and germinal cells. J. Genet. Genom. 45, 681–684 (2018).
Mao, Y. et al. Development of germ-line-specific CRISPR-Cas9 systems to improve the production of heritable gene modifications in Arabidopsis. Plant Biotechnol. J. 14, 519–532 (2016).
Eid, A., Ali, Z. & Mahfouz, M. M. High efficiency of targeted mutagenesis in Arabidopsis via meiotic promoter-driven expression of Cas9 endonuclease. Plant Cell Rep. 35, 1555–1558 (2016).
Feng, Z. et al. A highly efficient cell division-specific CRISPR/Cas9 system generates homozygous mutants for multiple genes in Arabidopsis. Int. J. Mol. Sci. 19, 3925 (2018).
Li, H.-J., Liu, N.-Y., Shi, D.-Q., Liu, J. & Yang, W.-C. YAO is a nucleolar WD40-repeat protein critical for embryogenesis and gametogenesis in Arabidopsis. BMC Plant Biol. 10, 169 (2010).
Yan, L. et al. High-efficiency genome editing in Arabidopsis using YAO promoter-driven CRISPR/Cas9 system. Mol. Plant 8, 1820–1823 (2015).
Tsutsui, H. & Higashiyama, T. pKAMA-ITACHI vectors for highly efficient CRISPR/Cas9-mediated gene knockout in Arabidopsis thaliana. Plant Cell Physiol. 58, 46–56 (2017).
Miki, D., Zhang, W., Zeng, W., Feng, Z. & Zhu, J.-K. CRISPR/Cas9-mediated gene targeting in Arabidopsis using sequential transformation. Nat. Commun. 9, 1967 (2018).
Kosicki, M., Tomberg, K. & Bradley, A. Repair of double-strand breaks induced by CRISPR–Cas9 leads to large deletions and complex rearrangements. Nat. Biotechnol. 36, 765–771 (2018).
Feng, Z. et al. Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in Arabidopsis. Proc. Natl Acad. Sci. USA 111, 4632–4637 (2014).
Tang, X. et al. A large-scale whole-genome sequencing analysis reveals highly specific genome editing by both Cas9 and Cpf1 (Cas12a) nucleases in rice. Genome Biol. 19, 84 (2018).
Rees, H. A., Wilson, C., Doman, J. L. & Liu, D. R. Analysis and minimization of cellular RNA editing by DNA adenine base editors. Sci. Adv. 5, eaax5717 (2019).
Grünewald, J. et al. Transcriptome-wide off-target RNA editing induced byCRISPR-guided DNA base editors. Nature 569, 433 (2019).
Kleinstiver, B. P. et al. High-fidelity CRISPR-Cas9 variants with undetectable genome-wide off-targets. Nature 529, 490–495 (2016).
Slaymaker, I. M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016).
Chen, J. S. et al. Enhanced proofreading governs CRISPR–Cas9 targeting accuracy. Nature 550, 407–410 (2017).
Casini, A. et al. A highly specific SpCas9 variant is identified by in vivo screening in yeast. Nat. Biotechnol. 36, 265–271 (2018).
Lee, J. K. et al. Directed evolution of CRISPR-Cas9 to increase its specificity. Nat. Commun. 9, 3048 (2018).
Vakulskas, C. A. et al. A high-fidelity Cas9 mutant delivered as a ribonucleoprotein complex enables efficient gene editing in human hematopoietic stem and progenitor cells. Nat. Med. 24, 1216 (2018).
Zhang, D. et al. Perfectly matched 20-nucleotide guide RNA sequences enable robust genome editing using high-fidelity SpCas9 nucleases. Genome Biol. 18, 191 (2017).
Liang, Z., Chen, K., Yan, Y., Zhang, Y. & Gao, C. Genotyping genome-edited mutations in plants using CRISPR ribonucleoprotein complexes. Plant Biotechnol. J. 16, 2053–2062 (2018).
Zhang, Q. et al. Potential high-frequency off-target mutagenesis induced by CRISPR/Cas9 in Arabidopsis and its prevention. Plant Mol. Biol. 96, 445–456 (2018).
Lowe, K. et al. Morphogenic regulators Baby boom and Wuschel improve monocot transformation. Plant Cell 28, 1998–2015 (2016).
Mookkan, M., Nelson-Vasilchik, K., Hague, J., Zhang, Z. J. & Kausch, A. P. Selectable marker independent transformation of recalcitrant maize inbred B73 and sorghum P898012 mediated by morphogenic regulators BABY BOOM and WUSCHEL2. Plant Cell Rep. 36, 1477–1491 (2017).
Khanday, I., Skinner, D., Yang, B., Mercier, R. & Sundaresan, V. A male-expressed rice embryogenic trigger redirected for asexual propagation through seeds. Nature 565, 91 (2019).
Wang, C. et al. Clonal seeds from hybrid rice by simultaneous genome engineering of meiosis and fertilization genes. Nat. Biotechnol. 37, 283–286 (2019).
d’Erfurth, I. et al. Turning meiosis into mitosis. PLoS Biol. 7, e1000124 (2009).
Mieulet, D. et al. Turning rice meiosis into mitosis. Cell Res. 26, 1242–1254 (2016).
Kelliher, T. et al. One-step genome editing of elite crop germplasm during haploid induction. Nat. Biotechnol. 37, 287–292 (2019).
Wang, B. et al. Development of a haploid-inducer mediated genome editing (IMGE) system for accelerating maize breeding. Mol. Plant 12, 597–602 (2019).
Lemmon, Z. H. et al. Rapid improvement of domestication traits in an orphan crop by genome editing. Nat. Plants 4, 766 (2018).
Li, T. et al. Domestication of wild tomato is accelerated by genome editing. Nat. Biotechnol. 36, 1160–1163 (2018).
Zsögön, A. et al. De novo domestication of wild tomato using genome editing. Nat. Biotechnol. 36, 1211–1216 (2018).
Sarno, R. et al. Programming sites of meiotic crossovers using Spo11 fusion proteins. Nucleic Acids Res. 45, e164 (2017).
Jacobs, T. B., Zhang, N., Patel, D. & Martin, G. B. Generation of a collection of mutant tomato lines using pooled CRISPR libraries. Plant Physiol. 174, 2023–2037 (2017).
Lu, Y. et al. Genome-wide targeted mutagenesis in rice using the CRISPR/Cas9 system. Mol. Plant 10, 1242–1245 (2017).
Meng, X. et al. Construction of a genome-wide mutant library in rice using CRISPR/Cas9. Mol. Plant 10, 1238–1241 (2017).
Butt, H. et al. CRISPR directed evolution of the spliceosome for resistance to splicing inhibitors. Genome Biol. 20, 73 (2019).
Boesch, P. et al. DNA repair in organelles: Pathways, organization, regulation, relevance in disease and aging. Biochim. Biophys. Acta BBA - Mol. Cell Res. 1813, 186–200 (2011).
Srivastava, S. & Moraes, C. T. Manipulating mitochondrial DNA heteroplasmy by a mitochondrially targeted restriction endonuclease. Hum. Mol. Genet. 10, 3093–3099 (2001).
Tanaka, M. et al. Gene therapy for mitochondrial disease by delivering restriction endonuclease SmaI into mitochondria. J. Biomed. Sci. 9, 534–541 (2002).
Bacman, S. R., Williams, S. L., Pinto, M., Peralta, S. & Moraes, C. T. Specific elimination of mutant mitochondrial genomes in patient-derived cells by mitoTALENs. Nat. Med. 19, 1111–1113 (2013).
Gammage, P. A., Rorbach, J., Vincent, A. I., Rebar, E. J. & Minczuk, M. Mitochondrially targeted ZFNs for selective degradation of pathogenic mitochondrial genomes bearing large‐scale deletions or point mutations. EMBO Mol. Med. 6, 458–466 (2014).
Gammage, P. A., Moraes, C. T. & Minczuk, M. Mitochondrial genome engineering: the revolution may not be CRISPR-Ized. Trends Genet. 34, 101–110 (2018).
Jo, A. et al. Efficient mitochondrial genome editing by CRISPR/Cas9. BioMed. Res. Int. 2015, 305716 (2015).
Piatek, A. A., Lenaghan, S. C. & Neal Stewart, C. Advanced editing of the nuclear and plastid genomes in plants. Plant Sci. 273, 42–49 (2018).
Ruf, S. et al. High-efficiency generation of fertile transplastomic Arabidopsis plants. Nat. Plants 5, 282 (2019).
Bae, S., Kweon, J., Kim, H. S. & Kim, J.-S. Microhomology-based choice of Cas9 nuclease target sites. Nat. Methods 11, 705–706 (2014).
van Overbeek, M. et al. DNA repair profiling reveals nonrandom outcomes at Cas9-mediated breaks. Mol. Cell 63, 633–646 (2016).
Bothmer, A. et al. Characterization of the interplay between DNA repair and CRISPR/Cas9-induced DNA lesions at an endogenous locus. Nat. Commun. 8, 13905 (2017).
Chang, H. H. Y. et al. Different DNA end configurations dictate which NHEJ components are most important for joining efficiency. J. Biol. Chem. 291, 24377–24389 (2016).
Allen, F. et al. Predicting the mutations generated by repair of Cas9-induced double-strand breaks. Nat. Biotechnol. 37, 64–72 (2018).
Chakrabarti, A. M. et al. Target-specific precision of CRISPR-mediated genome editing. Mol. Cell 73, 699–713 (2019).
Dreissig, S. et al. Live-cell CRISPR imaging in plants reveals dynamic telomere movements. Plant J. Cell Mol. Biol. 91, 565–573 (2017).
Ma, H. et al. Multiplexed labeling of genomic loci with dCas9 and engineered sgRNAs using CRISPRainbow. Nat. Biotechnol. 34, 528–530 (2016).
Tanenbaum, M. E., Gilbert, L. A., Qi, L. S., Weissman, J. S. & Vale, R. D. A protein-tagging system for signal amplification in gene expression and fluorescence imaging. Cell 159, 635–646 (2014).
Fujita, T. & Fujii, H. Efficient isolation of specific genomic regions and identification of associated proteins by engineered DNA-binding molecule-mediated chromatin immunoprecipitation (enChIP) using CRISPR. Biochem. Biophys. Res. Commun. 439, 132–136 (2013).
Nishimasu, H. et al. Crystal Structure of Staphylococcus aureus Cas9. Cell 162, 1113–1126 (2015).
Kaya, H., Ishibashi, K. & Toki, S. A split Staphylococcus aureus Cas9 as a compact genome-editing tool in plants. Plant Cell Physiol. 58, 643–649 (2017).
Acknowledgements
Due to limited space, we could not cite all the related literature. We apologize to the authors whose work was not cited in this Review. Our plant genome engineering research is supported by the National Science Foundation Plant Genome Research Program (grant no. IOS-1758745), Biotechnology Risk Assessment Grant Program (grant no. 2018-33522-28789) from the US Department of Agriculture, the Foundation for Food and Agriculture Research (grant no. 593603) and Syngenta Biotechnology.
Author information
Authors and Affiliations
Contributions
Y.Z., A.A.M., S.S. and Y.Q. wrote the manuscript. Y.Z. prepared the figures. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Plants thanks S. Toki, L. Xia and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhang, Y., Malzahn, A.A., Sretenovic, S. et al. The emerging and uncultivated potential of CRISPR technology in plant science. Nat. Plants 5, 778–794 (2019). https://doi.org/10.1038/s41477-019-0461-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-019-0461-5
This article is cited by
-
Optimized protoplast isolation and transfection with a breakpoint: accelerating Cas9/sgRNA cleavage efficiency validation in monocot and dicot
aBIOTECH (2024)
-
Engineered plant extracellular vesicles for autoimmune diseases therapy
Nano Research (2024)
-
CRISPR/Cas9-mediated knockout of NYC1 gene enhances chlorophyll retention and reduces tillering in Zoysia matrella (L.) Merrill
Plant Cell Reports (2024)
-
Understanding the biochemical, physiological, molecular, and synthetic biology approaches towards the development of C4 rice (Oryza sativa L.)
Cereal Research Communications (2024)
-
Applications of CRISPR/Cas genome editing in economically important fruit crops: recent advances and future directions
Molecular Horticulture (2023)