Abstract
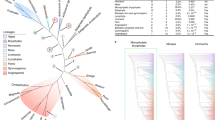
Angiosperms are by far the most species-rich clade of land plants, but their origin and early evolutionary history remain poorly understood. We reconstructed angiosperm phylogeny based on 80 genes from 2,881 plastid genomes representing 85% of extant families and all orders. With a well-resolved plastid tree and 62 fossil calibrations, we dated the origin of the crown angiosperms to the Upper Triassic, with major angiosperm radiations occurring in the Jurassic and Lower Cretaceous. This estimated crown age is substantially earlier than that of unequivocal angiosperm fossils, and the difference is here termed the ‘Jurassic angiosperm gap’. Our time-calibrated plastid phylogenomic tree provides a highly relevant framework for future comparative studies of flowering plant evolution.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Sequence alignments underlying analyses and all trees are available from the Dryad Digital Repository: https://doi.org/10.5061/dryad.bq091cg.
References
Friis, E. M., Crane, P. R. & Pedersen, K. R. Early Flowers and Angiosperm Evolution (Cambridge Univ. Press, 2011).
Benton, M. J. The origins of modern biodiversity on land. Phil. Trans. R. Soc. Lond. B 365, 3667–3679 (2010).
Darwin, C. in More Letters of Charles Darwin Vol. 12 (eds Darwin, F. & Seward, A. C.) Vol. 2, 12–13 (John Murray, 1903).
Misof, B. et al. Phylogenomics resolves the timing and pattern of insect evolution. Science 346, 763–767 (2014).
Peters, R. S. et al. Evolutionary history of the Hymenoptera. Curr. Biol. 27, 1013–1018 (2017).
Roelants, K. et al. Global patterns of diversification in the history of modern amphibians. Proc. Natl Acad. Sci. USA 104, 887–892 (2007).
Bininda-Emonds, O. R. P. et al. The delayed rise of present-day mammals. Nature 446, 507–512 (2007).
Schneider, H. et al. Ferns diversified in the shadow of angiosperms. Nature 428, 553–557 (2004).
Davis, C. C., Xi, Z. & Mathews, S. Plastid phylogenomics and green plant phylogeny: almost full circle but not quite there. BMC Biol. 12, 11 (2014).
Angiosperm Phylogeny Group IV. An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot. J. Linn. Soc. 181, 1–20 (2016).
Qiu, Y.-L. et al. The earliest angiosperms: evidence from mitochondrial, plastid and nuclear genomes. Nature 402, 404–407 (1999).
Soltis, D. E. et al. Angiosperms phylogeny: 17 genes, 640 taxa. Am. J. Bot. 98, 704–730 (2011).
Moore, M. J., Bell, C. D., Soltis, P. S. & Soltis, D. E. Using plastid genome-scale data to resolve enigmatic relationships among basal angiosperms. Proc. Natl Acad. Sci. USA 104, 19363–19368 (2007).
Cantino, P. D. et al. Towards a phylogenetic nomenclature of Tracheophyta. Taxon 56, 822–846 (2007).
Zeng, L. et al. Resolution of deep angiosperm phylogeny using conserved nuclear genes and estimates of early divergence times. Nat. Comm. 5, 4956 (2014).
Wickett, N. J. et al. Phylotranscriptomic analysis of the origin and early diversification land plants. Proc. Natl Acad. Sci. USA 111, E4859–E4868 (2014).
Gitzendanner, M. A., Soltis, P. S., Wong, G. K.-S., Ruhfel, B. R. & Solits, D. E. Plastid phylogenomic analysis of green plants: a billion years of evolutionary history. Am. J. Bot. 105, 291–301 (2018).
Drew, B. T. et al. Another look at the root of the angiosperms reveals a familiar tale. Syst. Biol. 63, 368–382 (2014).
Coiro, M., Doyle, J. A. & Hilton, J. How deep is the conflict between molecular and fossil evidence on the age of angiosperms? New Phytol. https://doi.org/10.1111/nph.15708 (2019).
Barba-Montoya, J., Dos Reis, M., Schneider, H., Donoghue, P. C. J. & Yang, Z. Constraining uncertainty in the timescale of angiosperm evolution and the veracity of Cretaceous terrestrial revolution. New Phytol. 218, 819–834 (2018).
Foster, C. S. P. et al. Evaluating the impact of genomic data and priors on Bayesian estimates of the angiosperm evolutionary timescale. Syst. Biol. 66, 338–351 (2017).
Magallón, S., Hilu, K. W. & Quandt, D. Land plant evolutionary timeline: gene effects are secondary to fossil constraints in relaxed clock estimation of age and substitution rates. Am. J. Bot. 100, 556–573 (2013).
Murat, F., Armero, A., Pont, C., Klopp, C. & Salse, J. Reconstructing the genome of the most recent common ancestor of flowering plants. Nat. Genet. 49, 490–496 (2017).
Magallón, S., Gómez-Acevedo, S., Sánchez-Reyes, L. L. & Hernández-Hernández, T. A metacalibrated time-tree documents the early rise of flowering plant phylogenetic diversity. New Phytol. 207, 437–453 (2015).
Bell, C. D., Soltis, D. E. & Soltis, P. S. The age and diversification of the angiosperms re-revisited. Am. J. Bot. 97, 1296–1303 (2010).
Beaulieu, J. M., O’Meara, B. C., Crane, P. & Donoghue, M. J. Heterogeneous rates of molecular evolution and diversification could explain the Triassic age estimate for angiosperms. Syst. Biol. 64, 869–878 (2015).
Zanne, A. E. et al. Three keys to the radiation of angiosperms into freezing enviroments. Nature 506, 89–92 (2014).
Doyle, J. A. Molecular and fossil evidence on the origin of angiosperms. Annu. Rev. Earth Planet. Sci. 40, 301–326 (2012).
Hochuli, P. A. & Feist-Burkhardt, S. Angiosperm-like pollen and Afropollis from the Middle Triassic (Anisian) of the Germanic Basin (northern Switzerland). Front. Plant Sci. 4, 344 (2013).
Herendeen, P. S., Peter, B. G. & Snapp, S. S. Palaeobotanical redux: revisiting the age of the angiosperms. Nat. Plants 3, 17015 (2017).
Friis, E. M., Crane, P. R., Pedersen, K. R., Stampanoni, M. & Marone, F. Exceptional preservation of tiny embryos documents seed dormancy in early angiosperms. Nature 528, 551–554 (2015).
Labandeira, C. C. in Evolutionary Biology: Genome Evolution, Speciation, Coevolution and Origin of Life (ed. Pontarotti, P.) 261–299 (Springer, 2014).
Davies, T. J. et al. Darwin’s abominable mystery: insights from a supertree of the angiosperms. Proc. Natl Acad. Sci. USA 101, 1904–1909 (2004).
Dilcher, D. Toward a new synthesis: major evolutionary trends in the angiosperm fossil record. Proc. Natl Acad. Sci. USA 97, 7030–7036 (2000).
Meredith, R. W. et al. Impacts of the Cretaceous terrestrial revolution and KPg extinction on mammal diversification. Science 334, 521–524 (2011).
Cardinal, S. & Danforth, B. N. Bees diversified in the age of eudicots. P. R. Soc. B 280, 20122686 (2013).
Moreau, C. S., Bell, C. D., Vila, R., Archibald, S. B. & Pierce, N. E. Phylogeny of the ants: diversification in the age of angiosperms. Science 312, 101–104 (2006).
Augusto, L., Davies, T. J., Delzon, S. & De Schrijver, A. The enigma of the rise of angiosperms: can we untie the knot? Ecol. Lett. 17, 1326–1338 (2014).
Wang, H. C. et al. Rosid radiation and the rapid rise of angiosperm-dominated forests. Proc. Natl Acad. Sci. USA 106, 3853–3858 (2009).
Chaboureau, A. C., Sepulchre, P., Donnadieu, Y. & Franc, A. Tectonic-driven climate change and the diversification of angiosperms. Proc. Natl Acad. Sci. USA 111, 14066–14070 (2014).
Deenen, M. H. L. et al. A new chronology for the end-Triassic mass extinction. Earth Planet. Sci. Lett. 291, 113–125 (2010).
Huynh, T. T. & Poulsen, C. J. Rising atmospheric CO2 as a possible trigger for the end-Triassic mass extinction. Palaeogeogr. Palaeoclimatol. Palaeoecol. 217, 223–242 (2005).
McElwain, J. C., Beerling, D. J. & Woodward, F. I. Fossil plants and global warming at the Triassic–Jurassic boundary. Science 285, 1386–1390 (1999).
Ren, D. Flower-associated Brachycera flies as fossil evidence for Jurassic angiosperm origins. Science 280, 85–88 (1998).
Nel, A. et al. The earliest known holometabolous insects. Nature 503, 257–261 (2013).
Blomenkemper, P., Kerp, H., Hamad, A. A., DiMichele, W. A. & Bomfleur, B. A hidden cradle of plant evolution in Permian tropical lowlands. Science 362, 1414–1416 (2018).
Doyle, J. J. & Doyle, J. L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 19, 11–15 (1987).
Yang, J.-B., Li, D.-Z. & Li, H.-T. Highly effective sequencing whole chloroplast genomes of angiosperms by nine novel universal primer pairs. Mol. Ecol. Resour. 14, 1024–1031 (2014).
Zhang, T., Zeng, C.-X., Yang, J.-B., Li, H.-T. & Li, D.-Z. Fifteen novel universal primer pairs for sequencing whole chloroplast genomes and a primer pair for nuclear ribosomal DNAs. J. Syst. Evol. 54, 219–229 (2016).
Zhang, Y.-J., Ma, P.-F. & Li, D.-Z. High-throughput sequencing of six bamboo chloroplast genomes: phylogenetic implications for temperate woody bamboos (Poaceae: Bambusoideae). PLoS ONE 6, e20596 (2011).
Huang, C.-H. et al. Resolution of Brassicaceae phylogeny using nuclear genes uncovers nested radiations and supports convergent morphological evolution. Mol. Biol. Evol. 33, 394–412 (2016).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. GigaScience 1, 18 (2012).
Katoh, K., Kuma, K., Toh, H. & Miyata, T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 33, 511–518 (2005).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Lanfear, R., Calcott, B., Ho, S. Y. W. & Guindon, S. PartitionFinder: combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol. Biol. Evol. 29, 1695–1701 (2012).
Smith, S. A. & O’Meara, B. C. TreePL: divergence time estimation using penalized likelihood for large phylogenies. Bioinformatics 28, 2689–2690 (2012).
Bouckaert, R. et al. BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 10, e1003537 (2014).
Bell, C. D., Soltis, D. E. & Soltis, P. S. The age of the angiosperms: a molecular timescale without a clock. Evolution 59, 1245–1258 (2005).
Magallón, S. & Castillo, A. Angiosperm diversification through time. Am. J. Bot. 96, 349–365 (2009).
Smith, S. A., Beaulieu, J. M. & Donoghue, M. J. An uncorrelated relaxed-clock analysis suggests an earlier origin for flowering plants. Proc. Natl Acad. Sci. USA 107, 5897–5902 (2010).
Clarke, J. T., Warnock, R. C. M. & Donoghue, C. J. Establishing a time-scale for plant evolution. New Phytol. 192, 266–301 (2011).
Acknowledgements
We thank the Germplasm Bank of Wild Species at the Kunming Institute of Botany (KIB) for facilitating this study; the curators and staff of the Beijing Botanical Garden (BG), Blue Mountains BG, Brisbane BG, Kunming BG, Missouri BG, Wuhan BG, Royal BG Edinburgh, RBG Kew, RBG Sydney, RBG Victoria (both Melbourne and Cranbourne), San Francisco BG, Shanghai Chenshan BG, South China BG, UC Berkeley BG, Xianhu BG Shenzhen, Xishuangbanna Tropical BG, Yinchuan BG and O. Maurin (Johannesburg, now Kew), J. R. Shevock (California), Y.-M. Shui (Kunming), and N. Zamora (Costa Rica) for samples; and S. R. Manchester (Florida) for critical discussion on fossil selection and calibration. This work was funded by the Strategic Priority Research Programme of the Chinese Academy of Sciences (CAS) (grant No. XDB31000000 to D.-Z.L.), CAS’ Large-scale Scientific Facilities (grant No. 2017-LSF-GBOWS-02 to D.-Z.L. and J.-B.Y.), KIB’s iFlora initiative (grant No. 2014-4-11 to D.-Z.L.) and the National Natural Science Foundation of China (grant No. 31570333 to H.-T.L.). P.-F.M. was supported by CAS’ Youth Innovation Promotion Association (grant No. 2015321) and P.S.S. was supported by the Ten Thousand Talents Programme of China and the Yunling International High-end Experts Programme of Yunnan Province.
Author information
Authors and Affiliations
Contributions
D.-Z.L, J.-B.Y., H.-T.L., T.-S.Y., D.E.S. and P.S.S. conceived the project and designed the research. T.-S.Y., T.Z., J.C., L.-M.G. and S.-D.Z. designed and carried out field collection work. Q.-F.W., J.W., P.W.F., M.v.d.B., P.M.H. and M.W.C. provided and/or collected samples. J.-B.Y., H.-T.L., Z.-R.Z., C.-N.F. and J.Y. performed DNA laboratory work. M.A.G., D.E.S. and P.S.S. prepared the OneKP dataset. H.-T.L., L.-M.G., T.-S.Y., P.-F.M., D.E.S. and P.S.S. designed and coordinated computational analyses. H.-T.L., T.Z., J.C., Y.L. and H.W. prepared the Figures and Tables. T.-S.Y., Y.L., L.-M.G., P.-F.M., D.-Z.L. and H.W. wrote the supplementary information. D.-Z.L., T.-S.Y., L.-M.G., P.-F.M. and P.W.F. wrote the first manuscript draft with input from all co-authors, particularly P.S.S., M.W.C., D.E.S. and P.M.H.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Journal peer review information: Nature Plants thanks Jennifer Mandel and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Information
Supplementary Text and Supplementary Figures 1–6.
Supplementary Table 1
Species sampled, including 1,659 newly sequenced samples for this study.
Supplementary Table 2
The details of removal of most rapidly evolving sites using Gblocks.
Supplementary Table 3
Ordinal and interordinal node age estimates using treePL based on the phylogenetics trees of 80 plastid genes of 2,881 samples with ML analysis.
Supplementary Table 4
Familial and interfamilial node age estimates using treePL based on the phylogenetics trees of 80 plastid genes of 2,881 samples with ML analysis.
Supplementary Table 5
Families age estimates using treePL based on the phylogenetics trees of 80 plastid genes of 2,881 samples with ML analysis.
Supplementary Data
The configuration file for running the software TreePL.
Rights and permissions
About this article
Cite this article
Li, HT., Yi, TS., Gao, LM. et al. Origin of angiosperms and the puzzle of the Jurassic gap. Nat. Plants 5, 461–470 (2019). https://doi.org/10.1038/s41477-019-0421-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-019-0421-0
This article is cited by
-
Plastid phylogenomics and fossil evidence provide new insights into the evolutionary complexity of the ‘woody clade’ in Saxifragales
BMC Plant Biology (2024)
-
Complete chloroplast genome and comparison of herbicides toxicity on Aeschynomene indica (Leguminosae) in upland direct-seeding paddy field
BMC Genomics (2024)
-
Draft genome assemblies for two species of Escallonia (Escalloniales)
BMC Genomic Data (2024)
-
Phylogenomics resolves the higher-level phylogeny of herbivorous eriophyoid mites (Acariformes: Eriophyoidea)
BMC Biology (2024)
-
Macroevolutionary dynamics of gene family gain and loss along multicellular eukaryotic lineages
Nature Communications (2024)